New publication from our lab! It was really fun writing this review @jcb.org with my postdoc Dr. Chris Mellor and graduate student Elisabeth Larson. We explore how folate deficiency trigger DNA damage that disrupt genome stability.
21.08.2025 13:21 —
👍 14
🔁 6
💬 0
📌 0
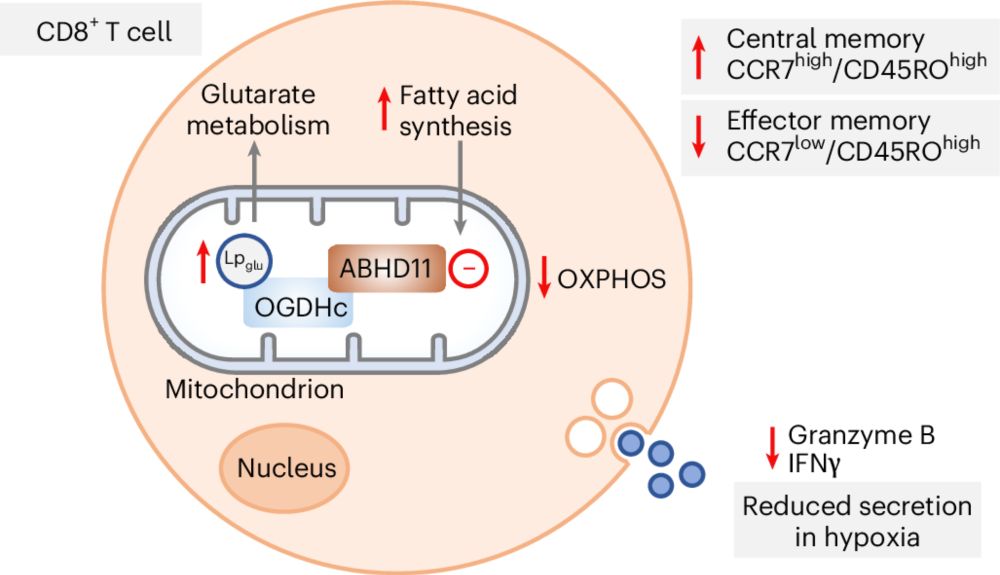
Lipoyl deglutarylation by ABHD11 regulates mitochondrial and T cell metabolism
Nature Chemical Biology - Glutarate rewires cell metabolism through conjugation to proteins or fatty acids, but how glutarylation is regulated is unclear. ABHD11 was identified as a...
Delighted to share our latest work on ABHD11 as a novel deglutarylating enzyme, and its role in T cell metabolism @natchembio.nature.com rdcu.be/ewiYZ. Many congratulations to co-first authors Guine Grice, Eleanor Minogue and @hudsonwilco.bsky.social! Great team effort with Randall Johnson's lab.
15.07.2025 16:12 —
👍 21
🔁 6
💬 1
📌 0

Mapping the genetic landscape of iron metabolism uncovers the SETD2 methyltransferase as a modulator of iron flux
Genetic screens using endogenous reporters map key modulators of cellular iron, including the transcriptional regulator SETD2.
Happy to share latest work from the lab. Led by talented first author @awmartinelli.bsky.social. Mapping the genetic landscape of iron metabolism uncovers the SETD2 methyltransferase as a modulator of iron flux | Science Advances www.science.org/doi/10.1126/... Congratulations to all the team.
18.09.2025 19:02 —
👍 10
🔁 2
💬 0
📌 3

Delighted that Ziqi Dong's PhD paper is out for all to read! Hypoxia is fundamental to normal development, and fascinating! Thanks to all of our co-authors including @jamesnathanlab.bsky.social @jellevda.bsky.social authors.elsevier.com/sd/article/S...
11.10.2025 08:22 —
👍 82
🔁 27
💬 4
📌 3
Mapping SET1B chromatin interactions with DamID using DamMapper, a comprehensive Snakemake workflow
Excited to share our first bioinformatics contribution! 🎉 Led by Niek Wit in collaboration with the van den Ameele lab @jellevda.bsky.social , we developed DamMapper damid-seq.readthedocs.io/en/latest/, a workflow for DamID-seq analysis, to map SET1B and HIF-1 chromatin occupancy. rdcu.be/eK2TS
15.10.2025 10:10 —
👍 9
🔁 3
💬 0
📌 0





