Very 🔥 science on DNA acessibility and TF footprinting from the @lucapinello.bsky.social lab!
23.12.2024 18:32 — 👍 3 🔁 0 💬 0 📌 0Very 🔥 science on DNA acessibility and TF footprinting from the @lucapinello.bsky.social lab!
23.12.2024 18:32 — 👍 3 🔁 0 💬 0 📌 0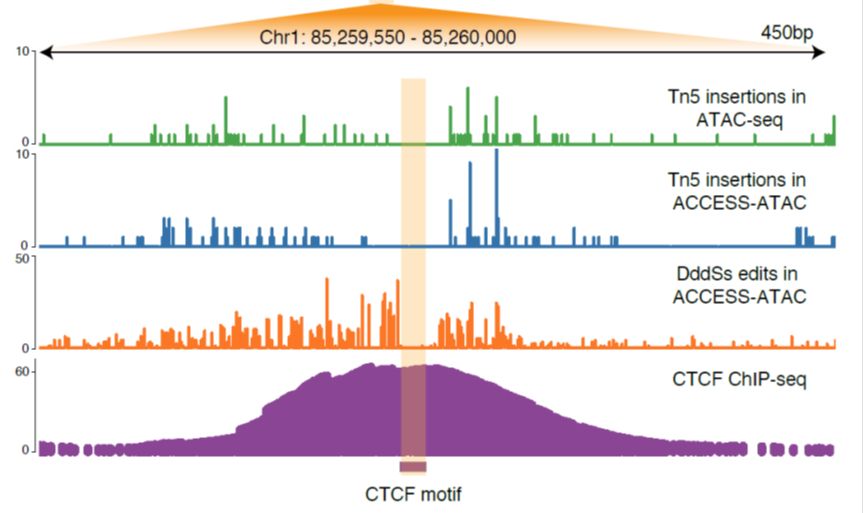

4/ ACCESS-ATAC significantly improves TF binding site prediction, and in single-cell ACCESS-ATAC experiments, we resolve TF binding profiles at equal sensitivity to ATAC-seq with 2-3x fewer cells.
23.12.2024 15:18 — 👍 6 🔁 3 💬 1 📌 1
Our new paper out today: A bioprinted and scalable model of human tubulointerstitial kidney fibrosis. Led by Daphne Bouwens, Nazanin Kabgani and Jitske Jansen www.sciencedirect.com/science/arti...
13.12.2024 06:34 — 👍 14 🔁 4 💬 1 📌 0
I would trust that BPNet does better than linear models, but I am not sure about the LM models, which are often published by only compare among themselves. See the study on DL models in insilico pertubation: www.biorxiv.org/content/10.1...
Maybe a 1-layer BPnet would do the job here?
Very nice benchmarking Anshul! I just missed a weak baseline (linear model?). Experience shown that they might still be better or as good as a lot of DL/LM models.
11.12.2024 13:45 — 👍 1 🔁 0 💬 1 📌 0 10.12.2024 13:07 —
👍 107
🔁 16
💬 2
📌 1
10.12.2024 13:07 —
👍 107
🔁 16
💬 2
📌 1

We are recruiting a Tenure-track Assistant Professor "Machine Learning in the Life Sciences" at the Medical University of Vienna. We will nominate the top candidate for a € 1.8 million starting grant by a Viennese foundation. Deadline: Dec 31st. Details: medical-epigenomics.org/files/Tenure...
02.12.2024 09:35 — 👍 13 🔁 9 💬 0 📌 0Have AI or ML skills? Have applied them to biological problems or high dimensional data problems? Want to get to a mechanistic understanding of animal behaviour and environmental influences on life? Want to live in Rome, work in an international, english speaking institute? Apply below!
01.12.2024 18:14 — 👍 7 🔁 2 💬 0 📌 0
SIGURD is implemented as an R package with associated pre-processing scripts. It is available on github.com/CostaLab/sig.... Amazing collaborative work with M. Kalmer, S. Koschmieder, @brummendorf within the #CRU344 #genotyping #singlecell #MPN 5/5
25.11.2024 15:43 — 👍 2 🔁 0 💬 0 📌 0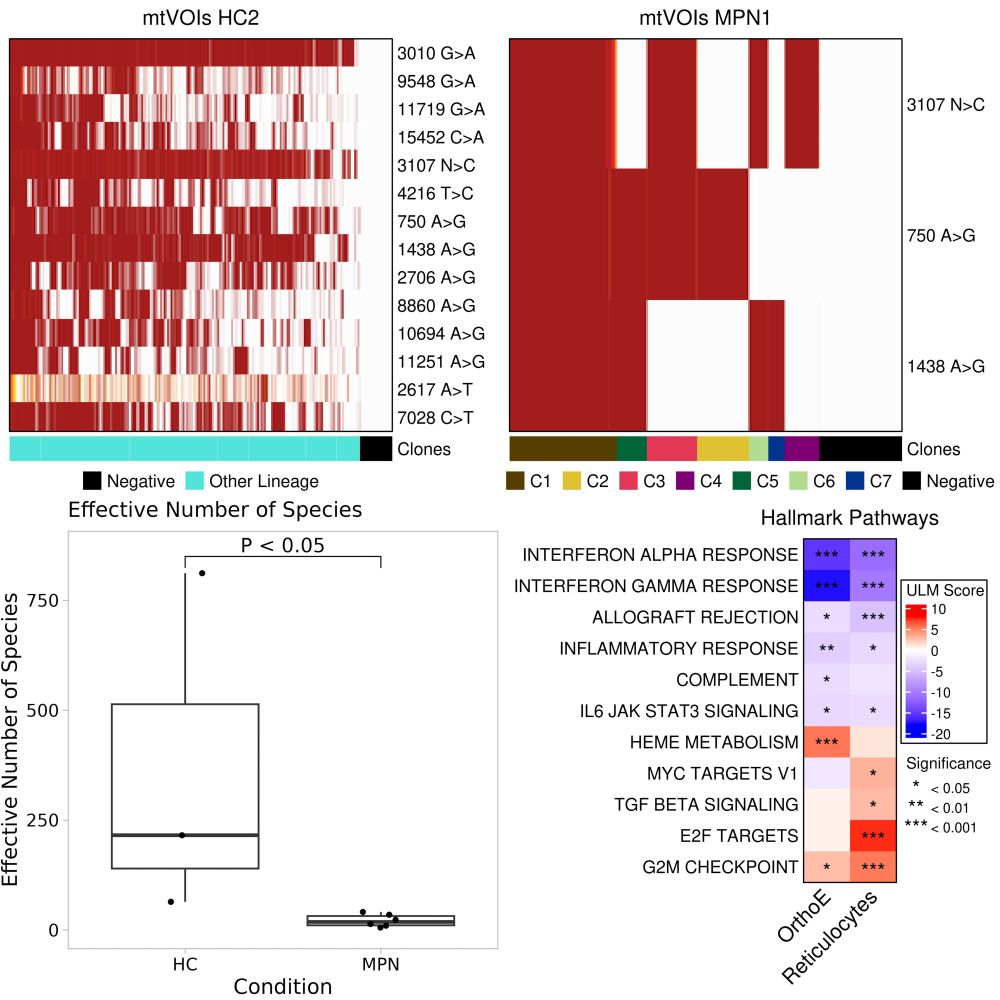
We build on MAEGATK for delineation of mitochondria derived clones. We observed an increase in clonality in MPN compared to HC, as supported by diversity measures. As before, we can functionally characterize clones with GSEA and enrichment analysis 4/5
25.11.2024 15:43 — 👍 2 🔁 0 💬 1 📌 0
We demonstrated SIGURD in scRNA-seq and JAK2V617F amplicon data from Myeloproliferative Neoplasms. Combined analysis with scRNA-seq allowed us to find an enrichment of JAK2V617F cells in late erythroid cells and an increase in heme metabolism 3/5
25.11.2024 15:42 — 👍 1 🔁 0 💬 1 📌 0SIGURD combines distinct genotyping tools (cellSNP-lite, VarTrix, MAEGATK) into a single reproducible framework. Genotyping results are stored in a data container and can be used for integrative analysis with single cell RNA #Seurat 2/5
25.11.2024 15:42 — 👍 0 🔁 0 💬 1 📌 0
We are happy to present SIGURD, a framework for single cell level analysis of somatic and mitochondrial variants from scRNA-seq data. This end-to-end framework allows genotyping, QC, functional analysis and data integration academic.oup.com/bib/article/... 1/5
25.11.2024 15:41 — 👍 4 🔁 2 💬 1 📌 0✋
25.11.2024 07:45 — 👍 0 🔁 0 💬 0 📌 0
And we get no break ... Saturday morning at home and I found another low dimensional embbeding (HATCH-MAP) in the floor.
24.11.2024 09:26 — 👍 2 🔁 0 💬 0 📌 0