Why be satisfied with coding variant gene impairments? Check out Eva’s poster to learn how we scale DeepRVAT to the WGS UKBiobank. It’s been great fun being a part of this project!
#ASHG25
Why be satisfied with coding variant gene impairments? Check out Eva’s poster to learn how we scale DeepRVAT to the WGS UKBiobank. It’s been great fun being a part of this project!
#ASHG25

Excited to share UKBBGym at #ASHG25, a new benchmark for variant effect predictors using WGS, proteomics and phenotypes from 500K UKBiobank participants. Stop by for insights on the impact of non-coding variants and how computational scores stack up against exp assays.
Poster 5022W, Wed 2:30-4:30.

The Solvathons have been one of our most exciting research community experiences: Hands-on, effective – solving real cases during the events, and multidisciplinary – from clinicians to bioinformaticians. Thumbs up to the SolveRD community. Looking for more now with @erdera.bsky.social
rdcu.be/eFaqO
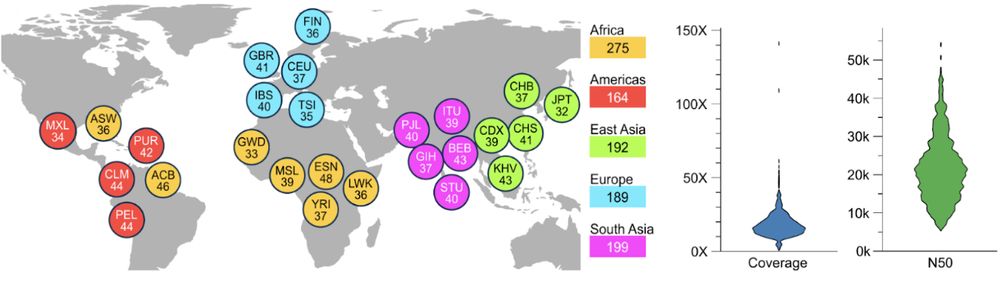
[1/8] *New Open-Access Long Read Resource*. We sequenced 1,019 genomes from the 1000 Genomes Project sample cohort using @nanoporetech.com long-read sequencing (LRS) to median 17x coverage. Publication at go.nature.com/4ffPb8f.
@hhu.de @crg.eu @embl.org @impvienna.bsky.social

Today at 12:20 in the HIT-Seq COSI, @lauradmartens.bsky.social Laura Martens will present her results on spatial chromatin accessibility data deconvolution #ISMBECCB2025. Great collab with Sarah Ouologuem and @fabiantheis.bsky.social
Proceedings paper:
doi.org/10.1093/bioi...
@thbec.bsky.social is going to share preliminary results on Meta-DeepRVAT, a new approach for deep learning based meta-analysis improving the power of rare variant association studies using population scale external control cohorts. #MLCSB
📅 July 21 |📍 Poster A-312
Join us for our next Kipoi Seminar with Katherine Pollard, Gladstone Institute of Data Science & Biotechnology,UCSF, Biohub
@gladstoneinst.bsky.social @czbiohub.bsky.social
👉Human variant interpretation with sequence-to-activity models
📅Wed June 4,5:30pm CET🧬 kipoi.org/seminar/🦋@kipoizoo.bsky.social
Excited to present at #eshg2025. Catch my talk on improving rare variant association studies using functional gene embeddings, on Monday at 11:45am (C26.06).
25.05.2025 10:33 — 👍 2 🔁 0 💬 0 📌 0Excited to be back at #eshg2025! Come by my poster today to check out fresh results on how rare high impact variants influence gene expression across immune cells—analyzed in 5,000 UK Biobank participants
25.05.2025 07:40 — 👍 10 🔁 3 💬 0 📌 0
Tomorrow Johannes Hingerl @johahi.bsky.social gives a talk on scooby at #probgen25. Enjoy learning in the legendary CSHL auditorium how to model RNA-seq and ATAC-seq profiles in individual cells from half a megabase of genomic sequence. Preprint:
doi.org/10.1101/2024...

Excited to share that PROTRIDER, our method to call outliers on mass spectrometry-based proteomics data, is out now!! #proteomics #massspectrometry #raredisease doi.org/10.1101/2025...
19.02.2025 08:15 — 👍 14 🔁 5 💬 1 📌 1
Join us for our next Kipoi Seminar with with Pedro Tomaz da Silva @pedrotomazdasilva.bsky.social @gagneurlab.bsky.social @TU_Muenchen!
👉Nucleotide dependency analysis of DNA language models reveals genomic functional elements
📅Wed Feb 5, 5:30pm CET
🧬https://kipoi.org/seminar/
🦋kipoizoo.bsky
Hey reg genomics folks, here is our little x-mas present: Flashzoi. Borzoi. Just as good. 3x faster. Thumbs up to @johahi.bsky.social for the great initiative, conception & implementation. Big thanks to Johannes Linder, David Kelley and colleagues to have created Borzoi and shared it freely.
23.12.2024 09:15 — 👍 18 🔁 7 💬 0 📌 0