Spatial Proteomics Workshop for Protists
(see more in thread below)
Registration
Early Bird Monday 22nd June 2026.
Regular Monday 10th August 2026.
There will be a conference dinner on day 1. To register visit protistology.org.uk/autumn-meeti...
06.02.2026 18:11 —
👍 25
🔁 17
💬 1
📌 0
Whoops, I didn't realize he was on Bluesky...sorry Gáspár
23.02.2026 18:10 —
👍 0
🔁 0
💬 0
📌 0
Have a novel protist's genome or transcriptome and want to predict it's mitochondrial proteome? Look no further than CoMR, this handy bioinformatic workflow to help you: www.biorxiv.org/content/10.6... #phylogenetics
23.02.2026 17:06 —
👍 18
🔁 11
💬 0
📌 0
Before he passed away in 2021, Tom Cavalier-Smith had drafted parts of his autobiography. It's now 'published' because Gáspár Jékely put a lot of effort in! Please enjoy and share: doi.org/10.5281/zeno...
23.02.2026 14:37 —
👍 28
🔁 13
💬 1
📌 1
Ok. Well perhaps I need to reconsider what I thought I had learned after three decades of a career in deep-time phylogenetics. I like cooking, so perhaps I'll pivot and put my energies into a pop-up restaurant or a catering business.
20.02.2026 12:01 —
👍 3
🔁 0
💬 0
📌 0
That looks really useful!
19.02.2026 16:33 —
👍 1
🔁 0
💬 0
📌 0
BTW, the reason that our approach didn't work well for anaerobic protists is because they lack the majority of proteins in our data matrix (Anaeramoeba has ~18% site occupancy...all other anaerobic protists had much less). There is just not enough information in the data.
19.02.2026 16:27 —
👍 0
🔁 0
💬 0
📌 0
And is the reason that "long branches" appear "long" relative to shorter branches in trees.
19.02.2026 16:25 —
👍 0
🔁 0
💬 1
📌 0
This is why there is no point on any of our trees which is equidistant to all of the tips of the tree.
19.02.2026 16:22 —
👍 0
🔁 0
💬 1
📌 0
Likelihood-based tree estimation allows for different rates in different lineages. Branchlengths are rate x time, and they are adjustable parameters, so they accommodate different rates in different lineages.
19.02.2026 16:20 —
👍 0
🔁 0
💬 1
📌 0
I think there is a misunderstanding somewhere. At the risk of getting stuck in an infinite loop, I'll give it another go: 1. We can't trust archaeal origin proteins to root euks because they are too divergent; 2. Bacterial origin proteins are NOT too divergent...therefore we can trust them more.
19.02.2026 16:18 —
👍 1
🔁 0
💬 1
📌 0
What you are suggesting is that the relative 'preference' (likelihoods) of rooting topologies would radically change if we included them. If we do include them and nothing changes...will you then accept the results of our analyses?
17.02.2026 23:24 —
👍 0
🔁 0
💬 1
📌 0
Diplonemids or kinetoplastids weren't in our analyses because of computational intensity considerations and the fact that nobody in deep euk. phylogenetics field doubts they are monophyletic.
17.02.2026 23:24 —
👍 0
🔁 0
💬 1
📌 0
Second how specifically does the "evolutionary trajectory" of mitochondrial proteins in different organisms compromise phylogenetic inference? What do you mean by evolutionary trajectory? How does that relate to phylogenetic models? Some precision is needed here otherwise nobody can address it.
17.02.2026 23:20 —
👍 0
🔁 0
💬 2
📌 0
I don't understand the logical point you are making: "if archaeal-origin proteins cannot be used"...."the validity of using bacterial-origin proteins is unclear". The point you "understand" is the rationale, no?
17.02.2026 23:18 —
👍 0
🔁 0
💬 2
📌 0

You 'understand this' yet you quote the following argument:
17.02.2026 23:16 —
👍 0
🔁 0
💬 1
📌 0
It wouldn't be appropriate to claim: "both cell biology studies showed strong conflicting results so I can't trust either (any?) of them" and so I have created an ad hoc scenario (based on no cell biological evidence) that protein X is secreted
16.02.2026 12:05 —
👍 3
🔁 0
💬 0
📌 0
However, a transfection study that expresses a GFP-tagged X might find that X does localize to the mitochondrion. Which one is right? You have to be familiar with methods to understand that the second study may be compromised by the over-expression of X leading to artefactual localization results.
16.02.2026 11:51 —
👍 4
🔁 0
💬 1
📌 0
It's not really that different to cases where different cell biology studies come to different conclusions. For example a study using a monoclonal Ab raised against protein X, might find that X is not localized to the mitochondrion by immunolocalization.
16.02.2026 11:51 —
👍 4
🔁 0
💬 1
📌 0
I note that it is a common misconception that different phylogenetic studies are all 'the same' when it comes to whether or not to trust the result. But you have to dig deeper to investigate the datasets used and the phylogenetic methods to understand why they come to different conclusions.
16.02.2026 11:51 —
👍 7
🔁 1
💬 1
📌 0
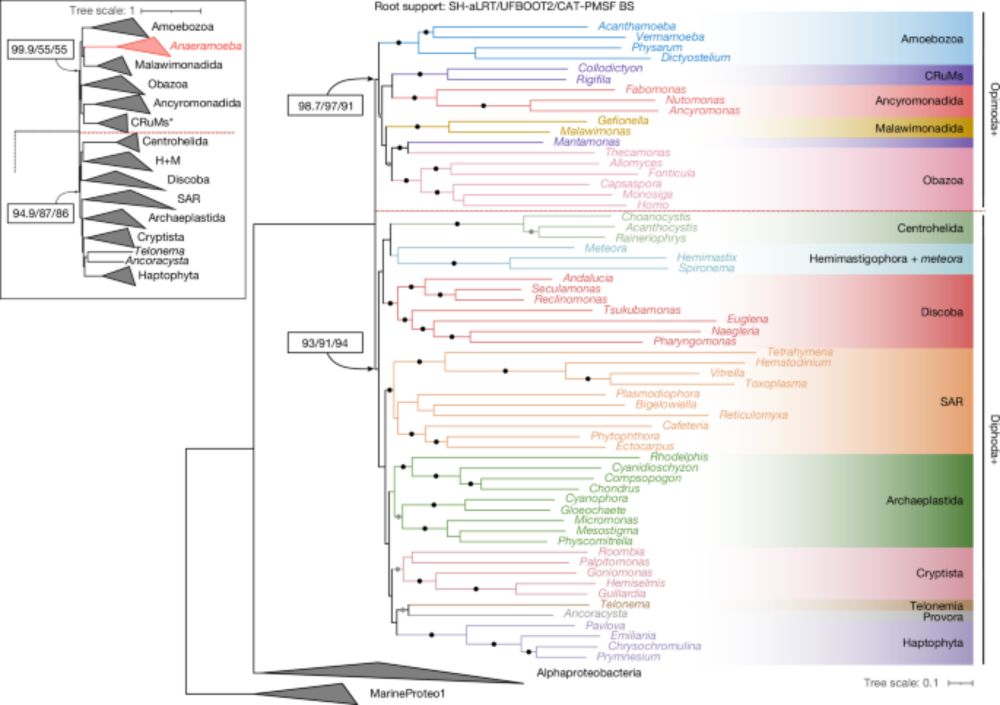
We also showed that the Opimoda+/Diphoda+ root was optimal under all of these models and all alternative root position (including a Euglenozoa root) were statistically rejected under all of these models (except for one -- the Anaeramoeba root).
16.02.2026 11:51 —
👍 1
🔁 0
💬 1
📌 0
In Williamson et al. we 'stress tested' our optimal root by using phylogenetic models that account for different features of deep-time sequence evolution including site-heterogeneity, compositional heterogeneity over the tree, site rate changes over the tree (heterotachy) and functional divergence
16.02.2026 11:51 —
👍 2
🔁 0
💬 1
📌 0
Contrast this with the dataset of Williamson et al. 2025. We used mitochondrial markers with an alphaproteobacterial outgroup. The length of the branch connecting the euk. subtree to the outgroup in this case is much shorter than in the archaeal outgroup analyses; there is less potential for LBA
16.02.2026 11:51 —
👍 3
🔁 0
💬 1
📌 1

An excavate root for the eukaryote tree of life
A dataset of 183 eukaryotic proteins of archaeal descent indicates an anaerobic, amitochondriate ancestry for modern eukaryotes.
And this is exactly what is seen in Al Jewari and Baldauf 2023 doi.org/10.1126/scia.... The longest internal branch in the eukaryote tree is the one leading to Parabasalia...and this is the branch where the archaeal subtree attaches (i.e. the 'root' of the euk. subtree).
16.02.2026 11:51 —
👍 4
🔁 0
💬 1
📌 0
The datasets that use archaeal orthologs as outgroups are problematic because the archaeal sequences are very distant to the eukaryotic ingroup sequences. This creates the potential for highly divergent ingroup seqs. to be attracted to the outgroup by LBA.
16.02.2026 11:51 —
👍 3
🔁 0
💬 1
📌 0
Briefly, the main reasons that recent 'root' analyses get different answers comes down to: i) different datasets which are differentially prone to long branch attraction artefacts (LBA) and ii) different phylogenetic models which are also differentially prone to LBA.
16.02.2026 11:51 —
👍 3
🔁 0
💬 1
📌 0
The 'devil is in the details'. You have to look beyond author claims and try to understand why the various studies have come to different conclusions and read the explanations given by the authors themselves.
16.02.2026 11:17 —
👍 7
🔁 3
💬 1
📌 0
Yes I read what you wrote. With respect, as someone who has worked on deep phylogenetics and deep-time sequence evolution for >2 decades, I don't think you made very clear or convincing arguments against the methods in our paper.
16.02.2026 03:08 —
👍 3
🔁 0
💬 1
📌 0