Latest from the lab! Led by @xuening-he.bsky.social, establishing a framework for interpretation of the motif syntax of regulatory elements and a quantitative model for transcription factor cooperativity. Plenty of cool stuff revealed, including TF redundancy and synergy, and their regulatory roles!
26.06.2025 17:32 —
👍 26
🔁 5
💬 0
📌 0

Symposium on ‘Enhancer Sequences’ at beautiful @collegedefrance.bsky.social in Paris, by April 11th. Open and free, no registration. Come to relax listening to facts #TherapeuticEffects #VillageGauloix Final programme below. Tell your friends! 🙏🤘
17.03.2025 14:28 —
👍 81
🔁 45
💬 2
📌 1
Job alert!
We are hiring a postdoc or PhD student focusing on
* developing computational methods to resolve gene regulation in single cells
* investigating the mechanisms and cell-type specific effects of genetic variants
PhD student: bsky.app/profile/jobr...
Postdoc: bsky.app/profile/jobr...
06.02.2025 20:36 —
👍 12
🔁 7
💬 0
📌 0
Job alert!
We are hiring a postdoc or PhD student focusing on
* developing computational methods to resolve gene regulation in single cells
* investigating the mechanisms and cell-type specific effects of genetic variants
PhD student: bsky.app/profile/jobr...
Postdoc: bsky.app/profile/jobr...
06.02.2025 20:36 —
👍 12
🔁 7
💬 0
📌 0

Enhancer Mechanics and Enhanceropathies
Join us for the highly anticipated third EMBO workshop on Enhancer Mechanics and Enhanceropathies! Building on the success of our previous meetings in Santander (2021) and Marseilles (2023), this eve…
Together with @smandrup.bsky.social and Minna Kaikkonen we are happy to announce the 3rd Edition of the EMBO Workshop on #Enhancers and #Enhanceropathies. This time we will meet during the beautiful Danish summer (June 16-21).
Book the dates and register soon!!!
meetings.embo.org/event/25-enh...
31.01.2025 14:35 —
👍 57
🔁 33
💬 0
📌 7

My book, An Intuitive Primer on Effective Functional Genomics Study Design, is published! I’d really appreciate it if you could help spread the word, and I’d love to hear your thoughts and feedback. I hope people will find it useful.
It’s available on Amazon: tinyurl.com/mx2hewen
17.01.2025 14:30 —
👍 111
🔁 68
💬 8
📌 3
This looks great! Just ordered.
17.01.2025 20:46 —
👍 1
🔁 0
💬 1
📌 0
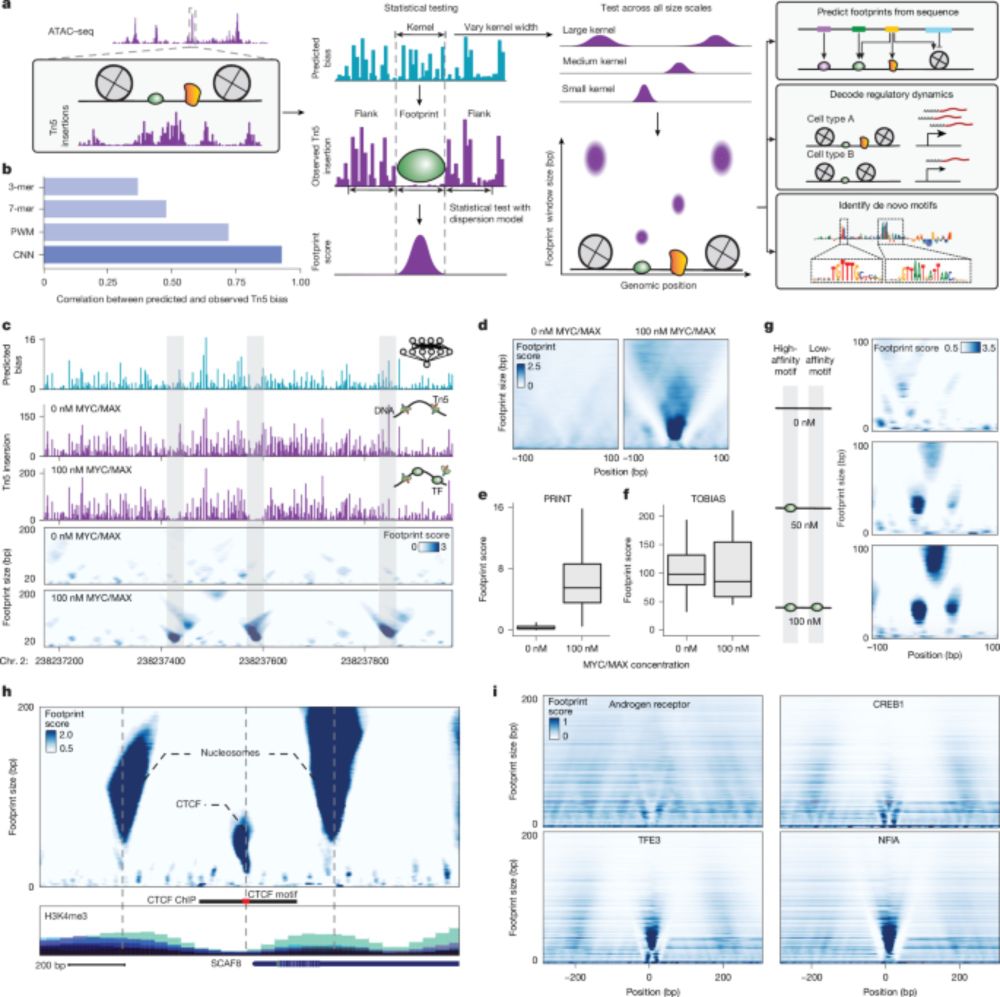
Our ChromBPNet preprint out!
www.biorxiv.org/content/10.1...
Huge congrats to Anusri! This was quite a slog (for both of us) but we r very proud of this one! It is a long read but worth it IMHO. Methods r in the supp. materials. Bluetorial coming soon below 1/
25.12.2024 23:48 —
👍 231
🔁 89
💬 8
📌 5

We are pursuing the wrong applications for AI.
22.12.2024 14:36 —
👍 708
🔁 167
💬 17
📌 16
Scientists, academics, researchers: We’re excited to share that @altmetric.com is now tracking mentions of your research on Bluesky! 🧪
03.12.2024 14:10 —
👍 29674
🔁 5028
💬 458
📌 279
I'd suggest this starter pack for #epigenetics and #generegulation:
go.bsky.app/TKp9YAn
plus see the description for another 3!
26.11.2024 10:25 —
👍 4
🔁 1
💬 1
📌 0
Introducing scE2G: a new model to link enhancers to target genes using single-cell data.
Excited that scE2G will enable building enhancer maps in hundreds of cell types in the human body!
Wonderful collaboration with @randersson.bsky.social @613weilin.bsky.social @mayayayas.bsky.social
Thread👇
25.11.2024 23:22 —
👍 29
🔁 8
💬 0
📌 0
Our preprint:
Molecular convergence of risk variants for congenital heart defects leveraging a regulatory map of the human fetal heart
Multiomic atlas + human genetics = new cell types, genes, and pathways that influence heart development and disease
www.medrxiv.org/content/10.1...
Thread 👇
25.11.2024 23:29 —
👍 25
🔁 11
💬 0
📌 0
What cell types drive congenital heart defects (CHD)?
Some new answers in our latest preprint, where we explored:
1). Key cell types contributing to CHD genetics
2). Impact of noncoding variants on CHD risk
https://www.medrxiv.org/content/10.1101/2024.11.20.24317557v1
25.11.2024 20:08 —
👍 58
🔁 20
💬 1
📌 2
Check out some highlights of the work here bsky.app/profile/613w...
25.11.2024 09:28 —
👍 2
🔁 0
💬 0
📌 0
Check out our latest work, scE2G, for mapping the target genes of enhancers from single cell data! www.biorxiv.org/content/10.1...
Amazing work led by co-first authors @mayayayas.bsky.social and @613weilin.bsky.social in a great collaboration with @jengreitz.bsky.social's lab
See 🧵 by Wei-Lin ⬇️
25.11.2024 09:22 —
👍 34
🔁 12
💬 1
📌 0
Thanks! Glad you liked the talk. Feel free to share as you wish.
16.11.2024 15:14 —
👍 1
🔁 0
💬 1
📌 0
Saturday morning: first session, second talk
14.11.2024 02:40 —
👍 1
🔁 0
💬 0
📌 0
Looking forward to #cshldata24 meeting starting tonight!
I will present #scE2G, a highly accurate model for mapping enhancer-gene regulatory interactions from single cell data. The model was developed in close collaboration with @jengreitz.bsky.social lab.
Who else is attending the meeting?
13.11.2024 13:46 —
👍 18
🔁 3
💬 2
📌 1
Seeing all the excitement from fellow scientists moving from X to here reminds me of academic Twitter 10 years ago. Let’s try out this platform!
I’m a computational genomicist focused on modeling gene regulation.
#firstpost #helloworld #introduction
13.11.2024 13:39 —
👍 42
🔁 2
💬 3
📌 0