RawBench: A Comprehensive Benchmarking Framework for Raw Nanopore Signal Analysis Techniques https://www.biorxiv.org/content/10.1101/2025.10.04.680405v1
06.10.2025 18:33 — 👍 4 🔁 2 💬 0 📌 0RawBench: A Comprehensive Benchmarking Framework for Raw Nanopore Signal Analysis Techniques https://www.biorxiv.org/content/10.1101/2025.10.04.680405v1
06.10.2025 18:33 — 👍 4 🔁 2 💬 0 📌 0
BREAKING: Our client Mario Guevara, an Emmy-winning journalist detained by ICE in retaliation for livestreaming law enforcement activity, will be deported tomorrow to El Salvador.
Mario and his family are being punished for his reporting. This cruelty is meant to stifle our free press.
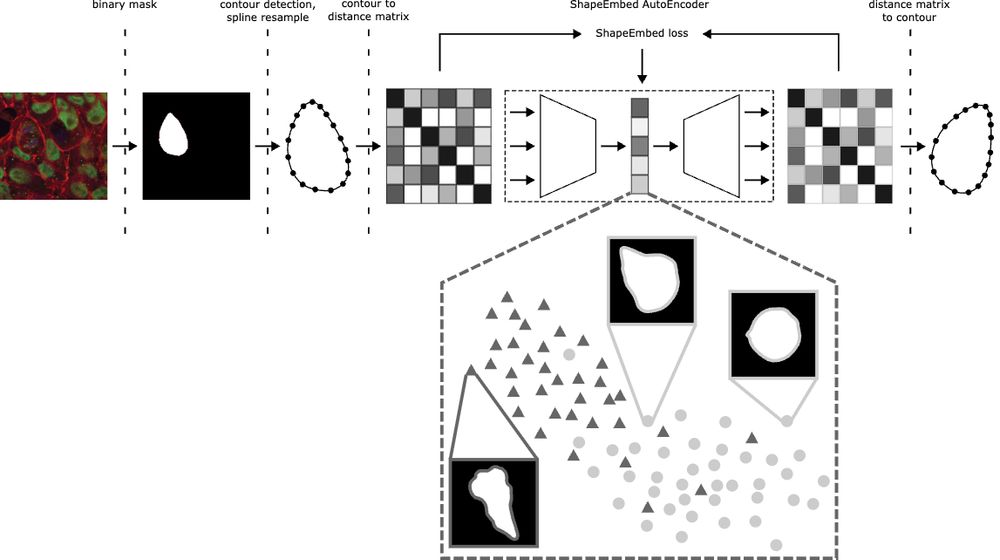
Happy to share that ShapeEmbed has been accepted at @neuripsconf.bsky.social 🎉 SE is self-supervised framework to encode 2D contours from microscopy & natural images into a latent representation invariant to translation, scaling, rotation, reflection & point indexing
📄 arxiv.org/pdf/2507.01009 (1/N)

Some personal news:
I've been fired from the Washington Post in the aftermath of the Charlie Kirk shooting.
Thread incoming.
substack.com/@karenattiah...

We present our new preprint titled "Large Language Model Hacking: Quantifying the Hidden Risks of Using LLMs for Text Annotation". We quantify LLM hacking risk through systematic replication of 37 diverse computational social science annotation tasks. For these tasks, we use a combined set of 2,361 realistic hypotheses that researchers might test using these annotations. Then, we collect 13 million LLM annotations across plausible LLM configurations. These annotations feed into 1.4 million regressions testing the hypotheses. For a hypothesis with no true effect (ground truth $p > 0.05$), different LLM configurations yield conflicting conclusions. Checkmarks indicate correct statistical conclusions matching ground truth; crosses indicate LLM hacking -- incorrect conclusions due to annotation errors. Across all experiments, LLM hacking occurs in 31-50\% of cases even with highly capable models. Since minor configuration changes can flip scientific conclusions, from correct to incorrect, LLM hacking can be exploited to present anything as statistically significant.
🚨 New paper alert 🚨 Using LLMs as data annotators, you can produce any scientific result you want. We call this **LLM Hacking**.
Paper: arxiv.org/pdf/2509.08825
 03.08.2025 00:48 —
👍 18405
🔁 5557
💬 296
📌 157
03.08.2025 00:48 —
👍 18405
🔁 5557
💬 296
📌 157
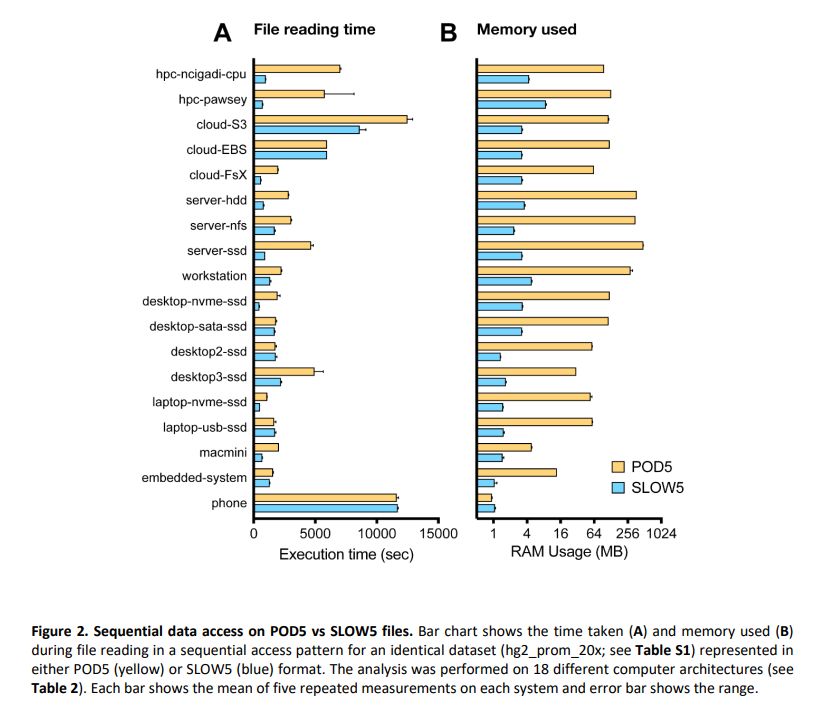
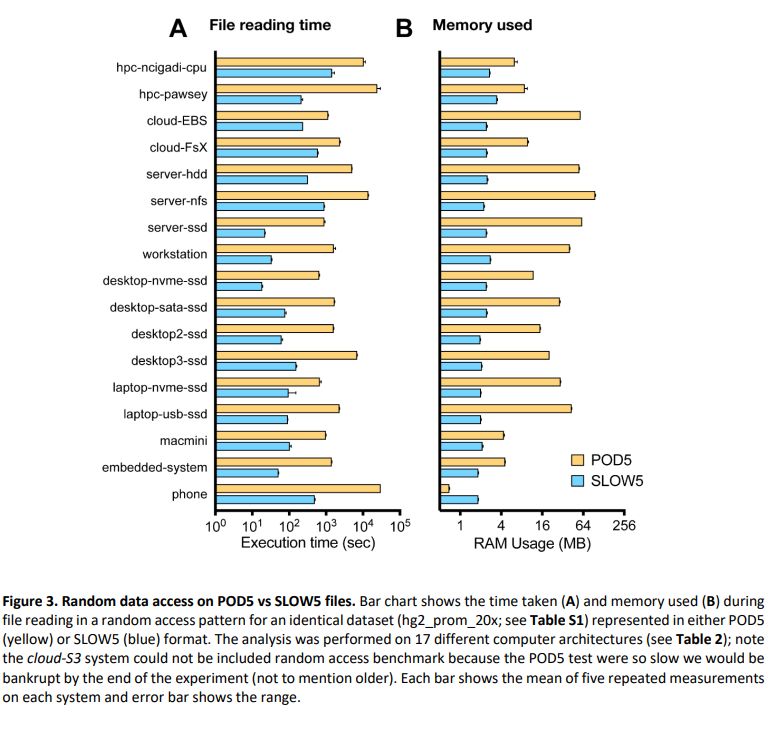
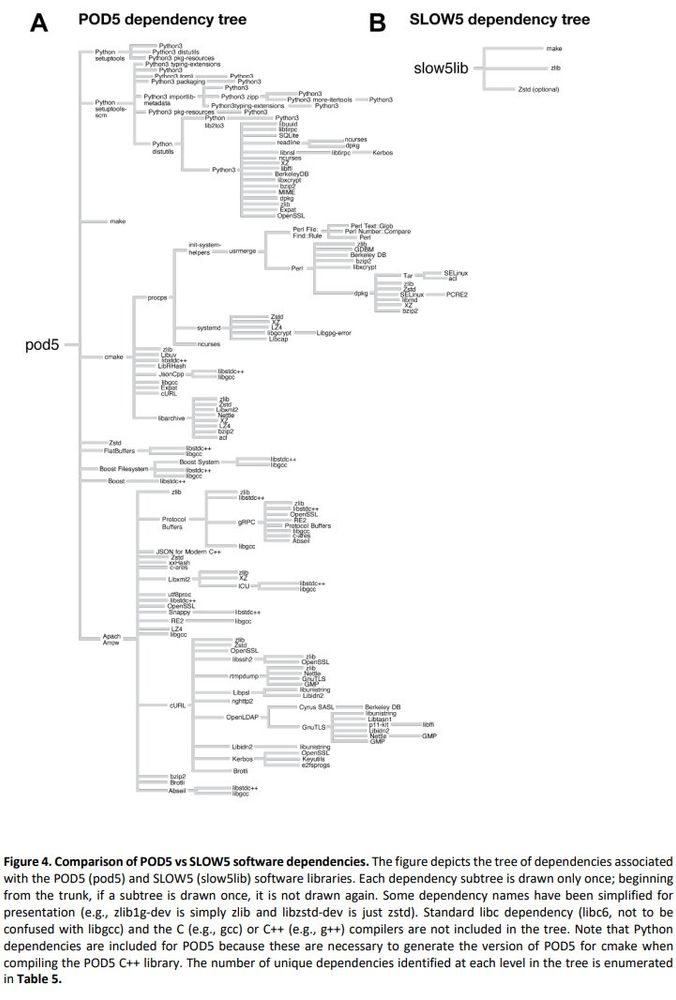
For many of those who were asking on BLOW5 vs POD5 for nanopore signal data, here is a finally detailed benchmark we did:
biorxiv.org/content/10.1...
Summary: performance of BLOW5 is >= POD5 (from ~= to 100X, see below), with benefit of having ~3 dependencies instead of >50.

"LMCompress shatters all previous lossless compression records on four media types: text, images, video and audio."
www.nature.com/articles/s42...

Analysis by altmetric shows increasing posting of research content on Bluesky but more sharing (reposting) on X.
We need to increase Bluesky connectivity and share more.

Toward single-molecule protein sequencing using nanopores
www.nature.com/articles/s41...

Roche SBX preprint out
www.biorxiv.org/content/10.1...
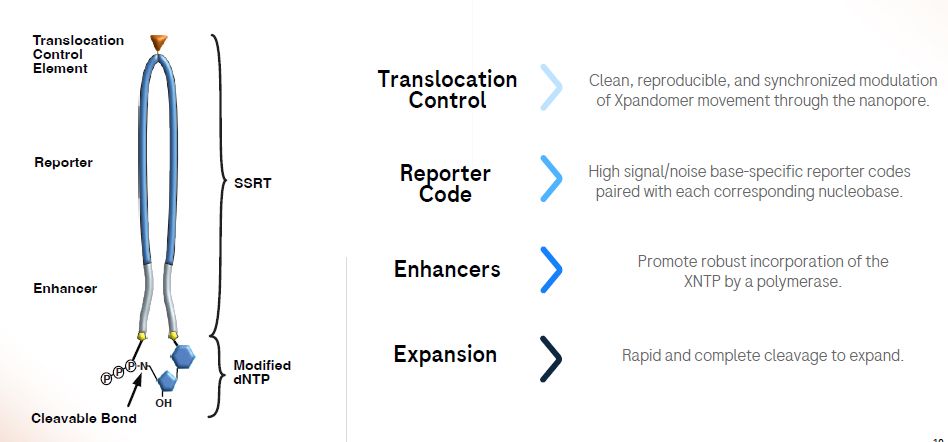
Roche Xpounds on New Sequencing Technology
My deep dive on this exciting new entrant
omicsomics.blogspot.com/2025/02/roch...
🧬🖥️



Verena Rukes telling us about: Charge-based fingerprinting of unlabeled full-length proteins using an
aerolysin nanopore
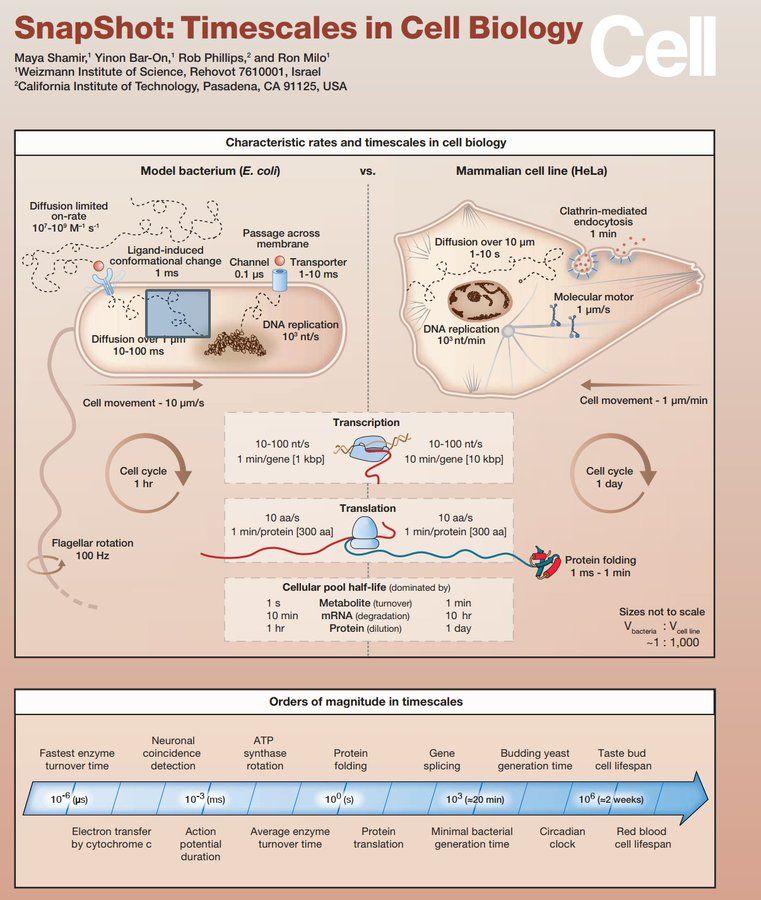
Timescales in Cell Biology
01.12.2024 13:35 — 👍 367 🔁 114 💬 8 📌 17
I took biochem in 2001, and for nearly 20 years read amino acid sequences daily… and I never knew Dayhoff named them or even the logic behind things like Q until last Friday (h/t Mike Janech). Also, this is another big Dayhoff moment for me. She was incredible!
#proteomics #bioinformatics

This worked like a charm to import accounts following on Twitter chromewebstore.google.com/detail/sky-f...
21.11.2024 05:37 — 👍 3 🔁 0 💬 1 📌 0