Gotta love a peppered moth story (and a soft selective sweep to boot)!
28.02.2026 15:38 — 👍 1 🔁 0 💬 0 📌 0Gotta love a peppered moth story (and a soft selective sweep to boot)!
28.02.2026 15:38 — 👍 1 🔁 0 💬 0 📌 0
In new work by @jahn0.bsky.social and I in @jbloomlab.bsky.social, we investigate how sequence constraints differ across influenza HA subtypes.
We find ~50% of sites in HA display substantially different amino-acid preferences across H3, H5, and H7.
doi.org/10.64898/202...

SpaceBar: a cellular barcoding approach for simultaneous analysis of cell clonal and spatial identities.
www.nature.com/articles/s41...
We designed SpaceBar to be as easy to use as possible:
Cells are labeled via lentivirus transduction, and barcode detection works with any standard imaging-based spatial transcriptomics workflow.
w/ Yael Heyman and @arjunraj.bsky.social
Check out the thread from the preprint for more info!

Excited that SpaceBar is now out in Nature Methods!🥳
We combined clone tracing with spatial transcriptomics to untangle what drives gene expression in tumors: a cell's identity or its neighborhood?
Most genes were driven by location, but some showed strong clonal patterns.
rdcu.be/eVhpc

🗞️ New preprint from the lab, led by our postdoc Ana Garoña (not on here) in collab with @andreagiometto.bsky.social: “Experimental evolution of cellular miniaturization reveals a mechanism for cell size evolution”, aka: “honey, we shrank the yeasts!” 🎥
www.biorxiv.org/content/10.6...
So excited to share this work led by @alexrob.bsky.social with Ben Kerr!
We investigated a poliovirus capsid inhibitor that exploits a breakdown in the genotype-phenotype map to prevent drug resistance evolution. Or does it?
See Alex's thread, but a few extras:
#socialviruses #evosky #virosky 🧪

Thrilled to finally share the magnum opus of my PhD that focuses on the genetic basis of evolutionary change! Specifically, we know we can map the genetic basis of a trait, but can we tell which genes will underlie the trait shift when it evolves? doi.org/10.1101/2025...
18.11.2025 00:14 — 👍 64 🔁 30 💬 2 📌 3
I am so excited to share new work on a TE insertion that regulates iridescence in swordtails, led by fantastic grad student @nadiahaghani.bsky.social and with help from many coauthors! In a time that has been so difficult to navigate, this & other projects have kept my spirits up: shorturl.at/NE65A
12.11.2025 20:34 — 👍 186 🔁 69 💬 3 📌 15
Glass beads of all sorts balancing on wavy threads.
How is functional variation at large-effect loci maintained in natural populations, even as environments change? In a paper led by @mkarag.bsky.social, we tracked known pesticide resistant alleles in outdoor 𝘋. 𝘮𝘦𝘭𝘢𝘯𝘰𝘨𝘢𝘴𝘵𝘦𝘳 cages & inferred selection and dominance from temporal sequencing data.
06.11.2025 21:51 — 👍 35 🔁 17 💬 1 📌 0Excited to share our recent work from @lowelab.bsky.social on the intersection of life history and cell type evolution: www.biorxiv.org/content/10.1...
06.11.2025 22:03 — 👍 15 🔁 4 💬 1 📌 1
One of the most exciting works of my career, years in the making. We used high-throughput precision genome editing to test the fitness effects of thousands of natural variants. Our findings challenge the long-held assumption that common variants are inconsequential.
www.biorxiv.org/content/10.1...

Our latest work is out in Nature today. In this paper, we introduce an improved version of NanoSeq, a duplex sequencing protocol with <5 errors per billion bp in single DNA molecules, and use it to study the somatic mutation landscape of oral epithelium in >1000 people www.nature.com/articles/s41...
08.10.2025 16:30 — 👍 89 🔁 47 💬 5 📌 1
Cancer is an evolutionary disease, but does knowing a cancer’s evolutionary past help predict its future? Out today in @nature, we learnt the evolution of 2000 lymphoid cancers and found it was highly correlated with clinical outcomes! (1/7)
rdcu.be/eFrrc
The constant barrage of terrible news on bluesky has made me feel weird about promoting papers, but people in the lab have been doing so much amazing work over the past few months that I want to share a few brief teasers/links:
10.09.2025 16:46 — 👍 67 🔁 22 💬 2 📌 1Super cool work, Joao! This matches some hints we had in some of our work that this likely happens after just 1 or 2 adaptive steps in our yeast system! Glad to see it studied in detail!
22.08.2025 14:38 — 👍 1 🔁 0 💬 1 📌 0
How common are frequency dependent fitness effects?
New preprint out today 👇
doi.org/10.1101/2025...
Congratulations, Roshni! Excited to see all the great things from your lab!
06.08.2025 19:19 — 👍 1 🔁 0 💬 1 📌 0Bittersweet to be leaving @docedge.bsky.social after a wonderful postdoc, but excited to share that I'm joining @uoregon.bsky.social next month as an Assistant Professor in the Department of Data Science.
06.08.2025 17:18 — 👍 146 🔁 28 💬 21 📌 4
I'm excited to announce our new biorxiv preprint, wherein we investigate the evolution of the weirdest genetic locus I've ever seen! Behold the tgr genes of the social amoeba, which mediate self/non-self discrimination during facultative multicellularity 🐅 🧵 1/
www.biorxiv.org/content/10.1...
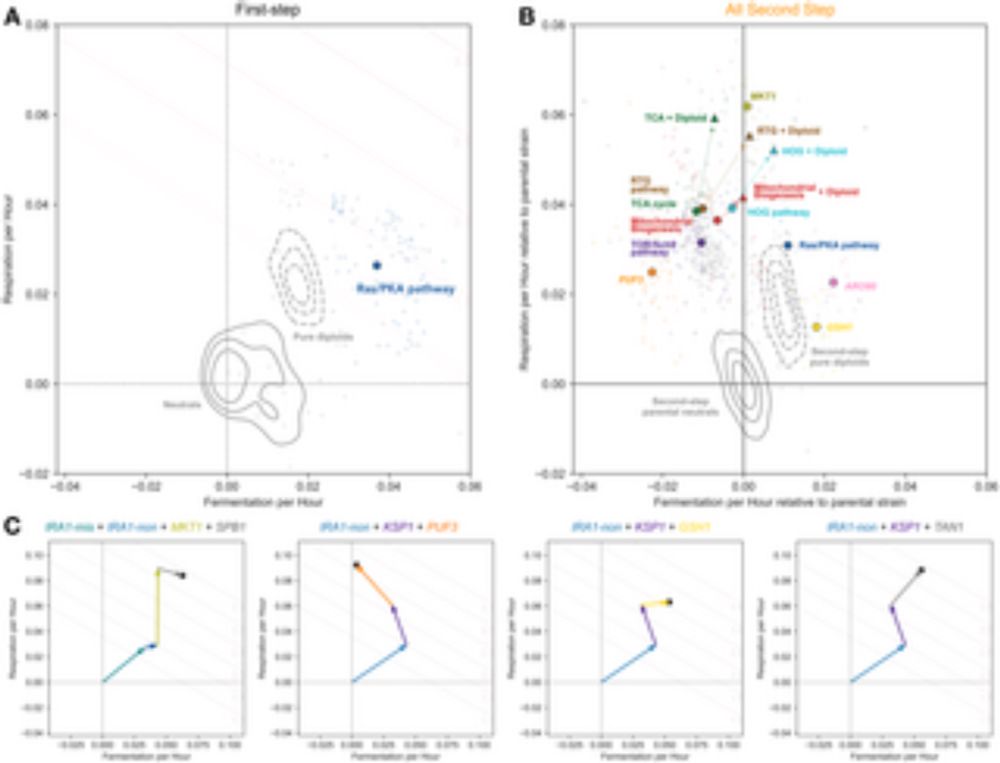
Having a great time at #smbe2025! So many great talks and conversations. Thank you again to @iamphioxus.bsky.social for the opportunity to present our work! Much of it is from the paper by @grantkinsler.bsky.social and @yuping-li.bsky.social journals.plos.org/plosbiology/... 1/n
22.07.2025 22:56 — 👍 28 🔁 9 💬 1 📌 0Excited to read it!
21.07.2025 20:26 — 👍 1 🔁 0 💬 0 📌 0New review article with @mmdesai.bsky.social is out today! Grateful for the opportunity to contribute something we hope will serve the community well
21.07.2025 17:30 — 👍 47 🔁 15 💬 3 📌 0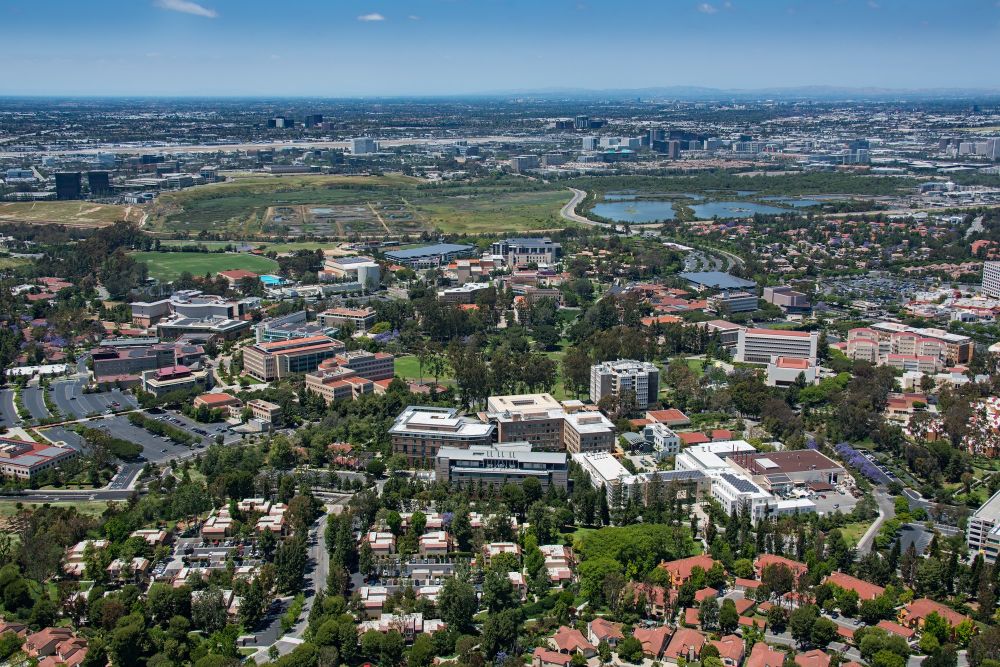
The Xue lab at UC Irvine is looking for a staff scientist to support our work investigating how microbes interact and evolve in the gut microbiome! Open to a wide range of previous experience levels, see ad for more.
recruit.ap.uci.edu/JPF09601

[0/8] Stoked to share our work with @arjunraj.bsky.social on tissue organization in the gastruloid. We use lineage tracing and spatial transcriptomics to show that diversity among stem cell clones promotes, rather than hinders, gastruloid development: www.biorxiv.org/content/10.1...
15.07.2025 15:44 — 👍 26 🔁 9 💬 1 📌 3
Very excited to share our new work on gastruloids by the incredible Cat Triandafillou! We mapped gene expression across 26 individual gastruloids at single-cell resolution and discovered some pretty amazing patterns about how these "mini-embryos" organize themselves.
www.biorxiv.org/content/10.1...
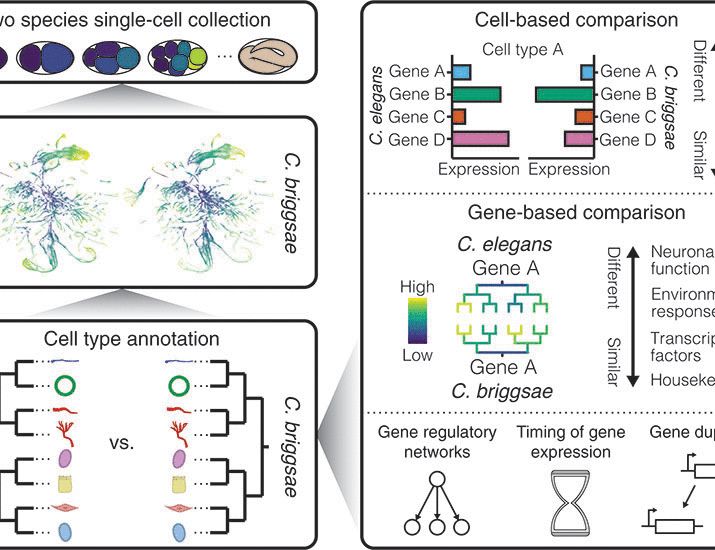
Happy to announce our paper comparing embryonic gene expression between C. elegans and C. briggsae, work led by Christopher Large with Rupa Khanal and in collaboration with Junhyong Kim and Bob Waterston. www.science.org/doi/10.1126/...
20.06.2025 20:25 — 👍 81 🔁 32 💬 5 📌 5Team Avenir forever!
09.07.2025 17:15 — 👍 1 🔁 0 💬 0 📌 0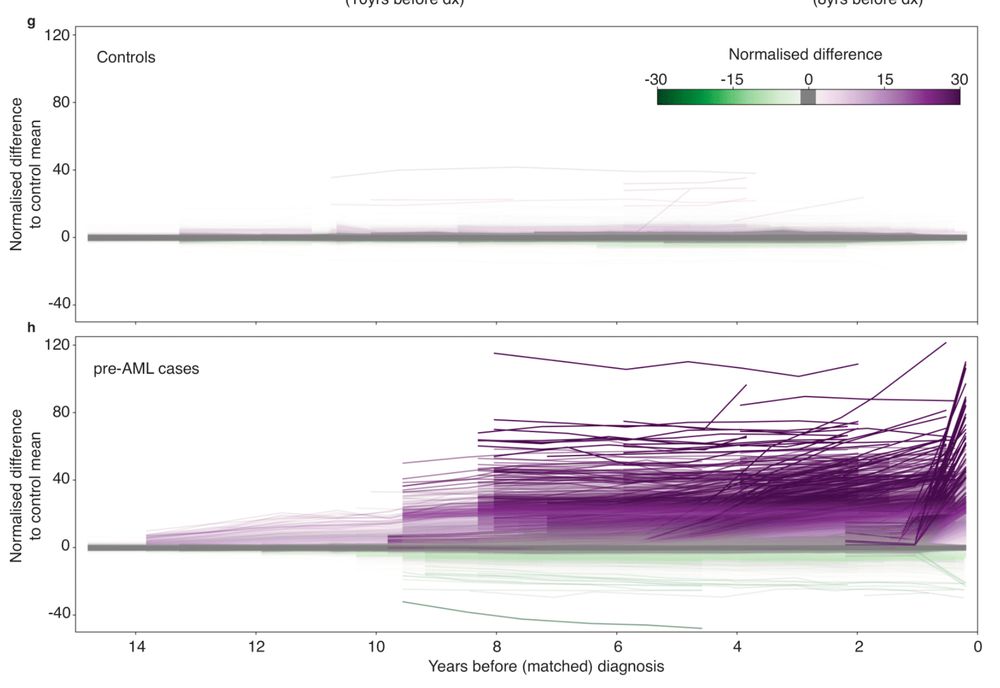
Delighted to share our latest on longitudinal methylation dynamics preceding cancer. Epigenetic signs of AML appear in blood DECADES before Dx.
👉 Early cancer detection
👉 Methylation drivers
👉 Epimutation rates
👉 CpG lineage tracing
www.biorxiv.org/content/10.1...
@jamie-blundell.bsky.social
09.07.2025 15:58 — 👍 2 🔁 0 💬 0 📌 0