Not sure - but keen to see an answer to this 👍
07.01.2026 11:53 — 👍 2 🔁 0 💬 0 📌 0Not sure - but keen to see an answer to this 👍
07.01.2026 11:53 — 👍 2 🔁 0 💬 0 📌 0
This paper by Celentano et al. on scalable simulation under the birth death is one of my favourites for the year! It introduces some elegant thinning that has been sorely needed. Fantastic to pave the way forward for future simulation-heavy inference, including neural Bayes!
doi.org/10.1073/pnas...
Looks interesting!
04.12.2025 08:08 — 👍 1 🔁 0 💬 0 📌 0
Our study on SARS-CoV-2 dynamics and long COVID in Nepal has made it to preprint 🎉 Huge thanks everyone from BNMT, Center for Molecular Dynamics Nepal (CMDN), and the Centre for Pathogen Genomics at The Peter Doherty Institute for Infection and Immunity 👏👏👏
www.medrxiv.org/content/10.1...

The BEAST X logo - an octopus wrapping its noodley appendages round the letter X.
BEAST X v10.5.0 finally released – you can't just do beta releases forever. github.com/beast-dev/be...
Details and instructions for installing on the BEAST website: beast.community

New paper from our lab in collaboration with Blaskovich lab led by former Phd student Anggia Prasetyoputri using in vitro selection to identify drug resistance conferring murations in S. aureus academic.oup.com/jacamr/artic...
26.06.2025 04:23 — 👍 1 🔁 1 💬 0 📌 0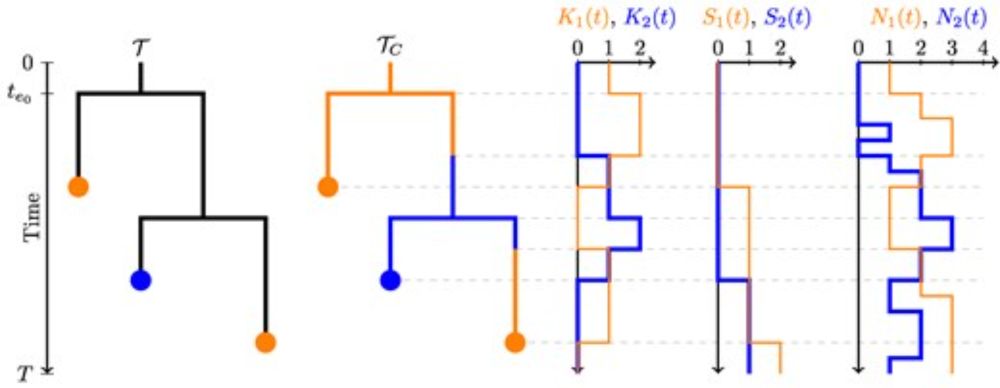
Vaughan & @tanjastadler.bsky.social develop a method to infer multitype population trajectories and apply it to MERS-CoV, revealing transmission patterns between camels and humans.
🔗 doi.org/10.1093/molbev/msaf130
#evobio #molbio #virus
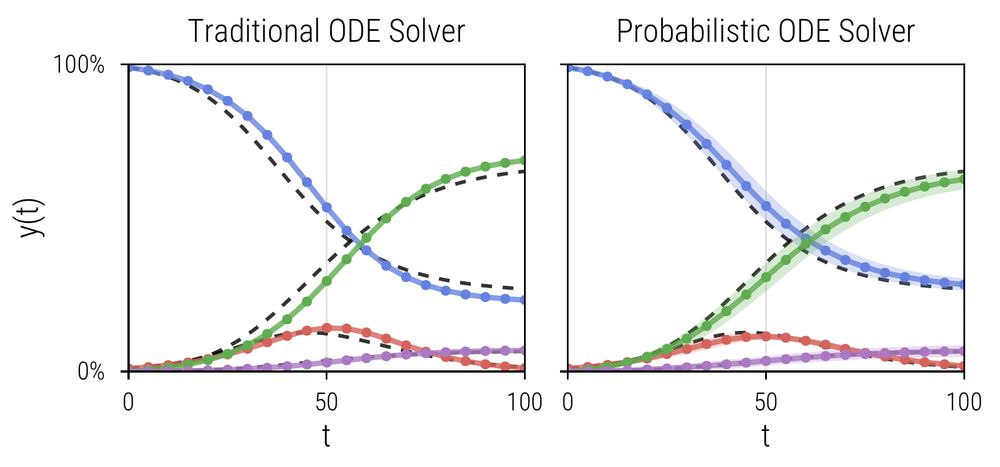
🎉 My PhD dissertation is now online! Traditional ODE solvers compute a single solution estimate - Probabilistic solvers also tell you how reliable they are! In my PhD, I established them as a Flexible and Efficient Framework for Probabilistic Simulation and Inference.
📄 tinyurl.com/mt3sffb
🧵 1/6
Please move genomes to an open pathogen platform like pathoplexus.org!
29.05.2025 05:08 — 👍 4 🔁 1 💬 0 📌 0Very exciting!
23.05.2025 23:36 — 👍 1 🔁 0 💬 0 📌 0
How can one efficiently simulate phylodynamics for populations with billions of individuals, as is typical in many applications, e.g., viral evolution and cancer genomics? In this work with M. Celentano, @wsdewitt.github.io , & S. Prillo, we provide a solution. doi.org/10.1073/pnas...
1/n

Ancestral sequence reconstruction often assumes unrealistic homogeneous substitution models; however @rmuniztrejo.bsky.social et al. find that reconstruction accuracy is determined by phylogenetic signal, not model choice.
🔗 doi.org/10.1093/molb...
#evobio #molbio

New preprint from the lab, describing an approach to enrichment of transcript isoforms for nanopore cDNA sequencing, using CRISPR-Cas9 www.biorxiv.org/content/10.1...
15.04.2025 04:41 — 👍 9 🔁 5 💬 1 📌 0
@aezarebski.bsky.social will give 2 talks in 🇨🇭:
🔹 Bern – 31/3 15:15
🔹 Basel – 16/4, 11:15
💡 Talk: Narrow AI for Phylodynamics
This visit is supported by SIB’s Speaker Roadshow Initiative, which coordinates multiple talks at partner institutions and promotes sustainable travel.