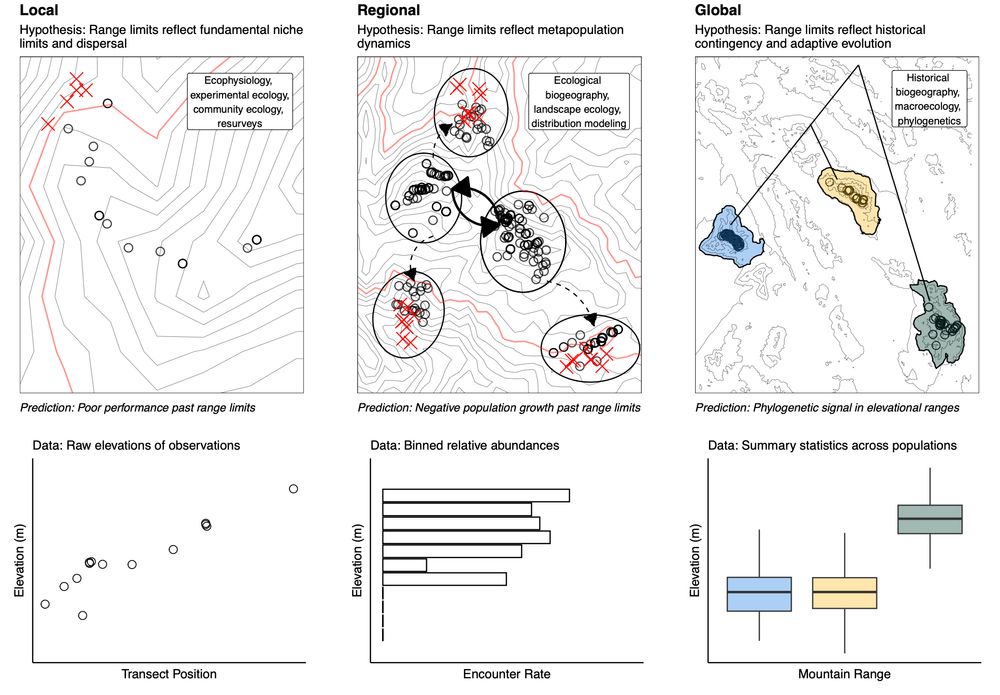
maybe my favorite paper I've written, I have a synthesis out today early access in @asn-amnat.bsky.social today that attempts to answer a simple but slippery question: what is an elevational range? doi.org/10.1086/737130
11.06.2025 21:44 —
👍 118
🔁 43
💬 6
📌 2
Thanks, Kara! The feeling is mutual! ☺️
17.07.2025 22:09 —
👍 0
🔁 0
💬 0
📌 0

Incredibly excited to announce that I will be starting as an assistant professor and curator of vertebrate collections at Northern Arizona University! I feel very fortunate to be joining such a wonderful department and also to live in a southwestern mountain town like Flagstaff — a dream come true!
16.07.2025 16:26 —
👍 43
🔁 2
💬 6
📌 1
the ad is up! applications due 7/15: evol.mcmaster.ca/brian/evoldi...
07.07.2025 14:06 —
👍 16
🔁 19
💬 1
📌 0


This work couldn’t have been done thanks to collaborations with Oscar Flores and others at UNAM and Joe Townsend at IUP. It’s been a wonderful privilege to work with collaborators across Mexico and Central America to shed some light on the incredible biodiversity there!
06.05.2025 15:58 —
👍 2
🔁 0
💬 0
📌 0
PNAS
Proceedings of the National Academy of Sciences (PNAS), a peer reviewed journal of the National Academy of Sciences (NAS) - an authoritative source of high-impact, original research that broadly spans...
Thrilled to see the last of my dissertation work out in @pnas.org ! We addressed some long-standing questions in leopard frogs across Mexico and Central America, discovering three undescribed species and taxonomic overinflation in 10 others. 🐸
www.pnas.org/doi/full/10....
06.05.2025 15:48 —
👍 28
🔁 9
💬 2
📌 0

Right, we messed up our own email address in yesterday's post. Let's try again: do you love evolutionary biology and art? Then our new logo contest may be for you!
06.09.2024 15:51 —
👍 3
🔁 8
💬 0
📌 0


Bears Ears always delivers, this time with a greater short-horned lizard! I’ve never found one this colorful before 😍
20.08.2024 13:44 —
👍 17
🔁 1
💬 0
📌 0

Please share: @UBCScience is searching for 15 TT faculty in diverse areas of expertise science.ubc.ca/about/careers. Included are 2 positions in Quantitative and Environmental Science as part of a hiring cluster for black faculty; you will join 6 current UBC faculty science.ubc.ca/blackscholars.
27.10.2023 20:00 —
👍 103
🔁 145
💬 2
📌 7
Thanks for the shoutout!! So glad you enjoyed the talk ☺️
27.10.2023 20:31 —
👍 1
🔁 0
💬 0
📌 0
ALGATR! I saw this presented at Evolution in Albuquerque and got very very excited
27.10.2023 15:57 —
👍 6
🔁 2
💬 3
📌 0

A figure with four examples of algatr’s added functionality: easily interpretable output tables with useful summary statistics for each method, kriged ancestry coefficients from TESS are now provided as rasters, variable loadings can act as legends for GDM maps, and the option for easy variance partitioning in RDA.
7/ Lastly, because @anushabishop.bsky.social and I love data visualization so much (and think a good figure is key to any paper!), we’ve added in functionality to make lots of fun tables and figures with algatr which we hope users will enjoy!
27.10.2023 20:24 —
👍 0
🔁 0
💬 0
📌 0

A screenshot showing several parts of the MMRR vignette, including lines to run the method as well as some useful data visualizations.
6/ algatr has detailed vignettes that will walk you through each of these methods and algatr’s new functionality:
27.10.2023 20:24 —
👍 0
🔁 0
💬 0
📌 0

A figure showing that algatr can calculate geographic distances and extract environmental variables at sampling coordinates, detect collinearity and perform a raster PCA on environmental layers, and perform LD-pruning, impute missing values, and calculate genetic distances from genetic data.
5/ algatr includes everything you need for processing coordinates, environmental layers, or genetic data for any of its methods 🧹:
27.10.2023 20:23 —
👍 0
🔁 0
💬 0
📌 0

A figure with three benefits of individual-based sampling: the first is lower impact on natural populations, showing all frogs collected at a pond (population-based sampling) vs. one frog collected per locality (individual-based sampling); the second is greater geographic and environmental coverage, showing that with individual-based sampling you can capture more heterogeneity across the environment because you can scatter your samples out more; and the third is better spatial resolution, showing example structure plots demonstrating how individual-based sampling can capture where a landscape barrier occurs because of increased coverage of sampling across the landscape.
4/ We also discuss the benefits of ind-based sampling for these methods – it’s great, it works, and is likely the future of LG studies! Below are some of the main benefits of this type of sampling 🗺️
27.10.2023 20:23 —
👍 0
🔁 0
💬 0
📌 0

A figure showing four key conservation questions alongside each of algatr’s methods. The questions are “How is genetic variation distributed?” answered using wingen (Bishop et al. 2023), “How do we delineate population units for management?” answered using TESS (Caye et al. 2016), “What are the drivers of population connectivity?” answered using multiple matrix regression with randomization (MMRR; Wang 2013) and generalized dissimilarity modeling (GDM; Fitzpatrick & Keller 2015), and “How can we describe and protect adaptive genetic variation?” answered using redundancy analysis (RDA; Capblancq & Forester 2021) and latent factor mixed modeling (LFMM; Caye et al. 2019).
3/ We really also wanted to prioritize methods that can inform conservation decisions so we framed the package in such a way as to answer key questions in conservation biology 🧬. algatr makes use of a lot of existing packages, so please be sure to cite them if you do use algatr!
27.10.2023 20:23 —
👍 2
🔁 1
💬 0
📌 0

An illustration of an alligator in the water; the back of the alligator has an abstract DNA double helix and the ripples of the water are landscape contour lines.
2/ In it, we introduce the algatr 🐊 package, openly available on GitHub. algatr builds on six landscape genomics methods that work for individual-based sampling: github.com/TheWangLab/a...
27.10.2023 20:21 —
👍 2
🔁 0
💬 0
📌 0
Individual‐based landscape genomics for conservation: An analysis pipeline
Molecular Ecology Resources is a broad journal publishing computer programs, statistical & molecular advances & more for studies in evolution, ecology & conservation.
Want to use landscape genomics for conservation? Curious about how to apply these methods to individual-based sampling? Not to worry– use algatr! Our new paper w/ co-first author @anushabishop.bsky.social and coauthor Ian Wang is out in MER! More below ⬇️ #OA
onlinelibrary.wiley.com/doi/10.1111/...
27.10.2023 20:09 —
👍 27
🔁 15
💬 6
📌 0













