A huge thank you goes to all co-authors for advancing the project alongside me! Vittorio Rainaldi, Nitin Bohra, Katsutaka Suzuki, Titouan DANET, Huda Kasim, Ari Satanowski, Hai He, Elena Rossini, Seung Hwan Lee, Beau Dronsella, Shanshan Luo, @nicoc-micsynmet.bsky.social
21.01.2026 19:52 —
👍 0
🔁 0
💬 0
📌 0
Realizing the MEVIS workflow as central piece of my PhD work took a village - I am grateful to Tobias Erb for giving me the freedom to pursue this vision, and to the Bosch Research Foundation for financing the work.
21.01.2026 19:52 —
👍 0
🔁 0
💬 1
📌 0
⭐ Why does it matter: MEVIS severely shortened the engineering timeline for achieving methylsuccinate assimilation - this should be transferable across modules ! Ultimately, shortening the DBTL cycles of synthetic biology and biotechnology is fundamental to building a circular bioeconomy.
21.01.2026 19:52 —
👍 0
🔁 0
💬 1
📌 0
📈 Seeing how ALE and MEVIS appeared orthogonal in tuning either the host metabolism or the synthetic module, we wondered whether both strategies might function synergistically. Indeed, combining both improved strain growth more than 200 % compared to either strategy alone !
21.01.2026 19:52 —
👍 0
🔁 0
💬 1
📌 0
While constitutive pathway enzyme expression made ALE essential to obtain growth with methylsuccinate, MEVIS enabled comparable growth without evolution, and within just weeks !
21.01.2026 19:52 —
👍 2
🔁 0
💬 1
📌 0
🏃♀️ To work efficiently, we combined machine learning and automation in the MEVIS workflow to directly optimize enzyme abundances and substrate concentrations in vivo!
21.01.2026 19:52 —
👍 1
🔁 0
💬 1
📌 0

✅ Our solution: We got curious whether informed tuning of a synthetic metabolic module for methylsuccinate assimilation, which is central to synthetic CO2 fixation cycles, (module-centric engineering) could bypass time-intensive ALE.
21.01.2026 19:52 —
👍 0
🔁 0
💬 1
📌 0
What happened repeatedly in evolutions towards C1-trophy of E. coli was a tuning of the host’s endogenous metabolic network allowing to accommodate the synthetic pathway better (host-centric tuning). But is this truly essential to achieve the desired growth?
21.01.2026 19:52 —
👍 0
🔁 0
💬 1
📌 0
❌ The problem: Metabolic engineering heavily relies on adaptive laboratory evolution to functionalize synthetic metabolism, for example for more efficient C1 assimilation, in vivo. However, desired phenotypes often emerge only after years - thus, metabolic engineering turnover times aren’t great.
21.01.2026 19:52 —
👍 0
🔁 0
💬 1
📌 0
Amazing work led by Vittorio Rainaldi from the lab of @nicoc-micsynmet.bsky.social on realizing the majority of the rMASP cycle for #synthetic #co2 #fixation in E. coli!
#Ccr, the key carboxylase of the CETCH, THETA and HOPAC cycles finally works #invivo !
25.11.2025 18:44 —
👍 1
🔁 0
💬 0
📌 0
Congrats !
24.10.2025 21:39 —
👍 0
🔁 0
💬 0
📌 0
Redirecting
🔥🔥🔥 Friday Evening, Paper Out, Feierabend 🔥🔥🔥
When evolution and enzyme engineering team up, they can convince microbes to assimilate sustainable carbon substrates preparing them for a life in a circular bioeconomy.
doi.org/10.1016/j.ym...
24.10.2025 17:00 —
👍 7
🔁 6
💬 3
📌 0
Good luck and congratulations Gerrich, I am sure you will do amazing!
09.10.2025 10:30 —
👍 2
🔁 0
💬 1
📌 0
Please repost!
Postdoc in adaptive laboratory evolution and C1 synthetic metabolism!
Location: DTU Biosustain
🦠🧬🔬
Starting in 01/2026!
Fell free to reach out if you have questions!
efzu.fa.em2.oraclecloud.com/hcmUI/Candid...
12.09.2025 11:43 —
👍 8
🔁 16
💬 1
📌 0
4) Many thanks to @beaubd.bsky.social and Tobias Erb for all the interesting discussions. In addition, want to thank Hai He, María Suárez Diez, and Elad Noor for constructive feedback on this work!
12.07.2025 16:44 —
👍 1
🔁 0
💬 0
📌 0
3) My hope is that this is a starting point to compile selection strains into a readily accessible collection. Let me know if you’d be interested !
12.07.2025 16:00 —
👍 1
🔁 0
💬 1
📌 0
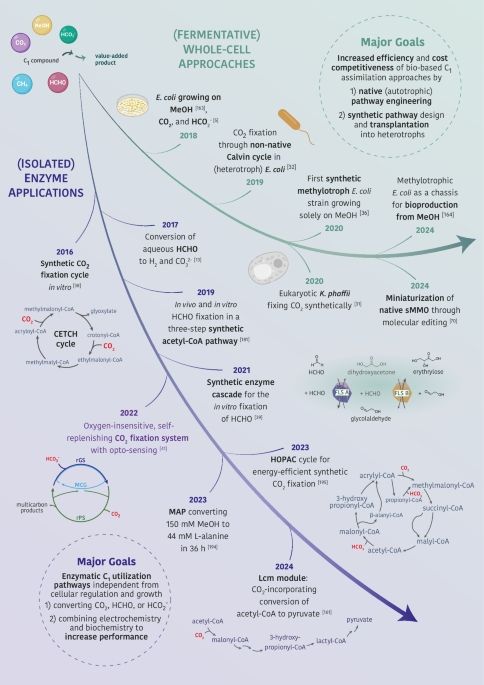
1) We dive into how strain growth parameters and pathway attributes are correlated, update some of the most central #schemes to our work (which I first got to work with in the Bar-Even lab!) and reflect on future developments in the field.
12.07.2025 15:58 —
👍 0
🔁 0
💬 1
📌 0

Escherichia coli selection strains for growth-coupled metabolic engineering
Synthetic metabolism has the potential to transform carbon capture, bioremediation, or bioproduction strategies. To transfer metabolic designs from an…
#Growth-coupled #selection just got an update ! In the last years, E. coli selection strains for #central , #amino #acid and #energy metabolism intermediates were created by the community. So, we compiled them and revisit the core concepts of growth coupling.
www.sciencedirect.com/science/arti...
12.07.2025 15:57 —
👍 10
🔁 5
💬 1
📌 0
#ME16
This is quite subjective , but imo
@helenasm.bsky.social just gave the most impressive talk of the conference so far and she is "only" in the third year of her PhD.
If you are interested in synthetic pathway engineering or C1 fixation you should check out her research.
16.06.2025 13:45 —
👍 5
🔁 2
💬 1
📌 0
Thank you so much Gerrich ! I am so honoured to show my work to such a supportive and interested audience and genuinely enjoyed discussing with you afterwards. Looking forward to see you shape tools for difficult-to-access hosts !
17.06.2025 10:51 —
👍 1
🔁 0
💬 1
📌 0
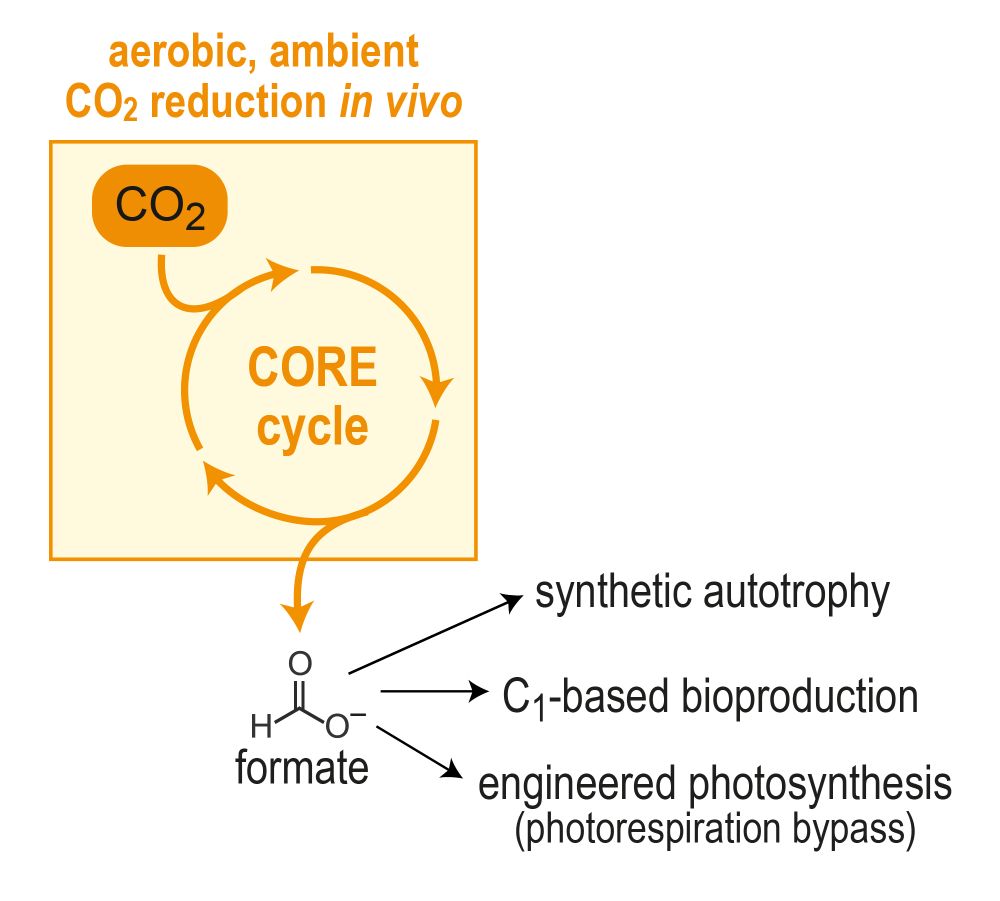
Excited to share a main project from my PhD, out now in
@naturecomms.bsky.social! 📝
We've designed and brought to life the “CORE cycle” – a new-to-nature pathway that provides a novel route for biological CO2 capture 🦠🌱
nature.com/articles/s41...
Take a look! Thread below... 🧵
04.04.2025 10:03 —
👍 39
🔁 17
💬 4
📌 4
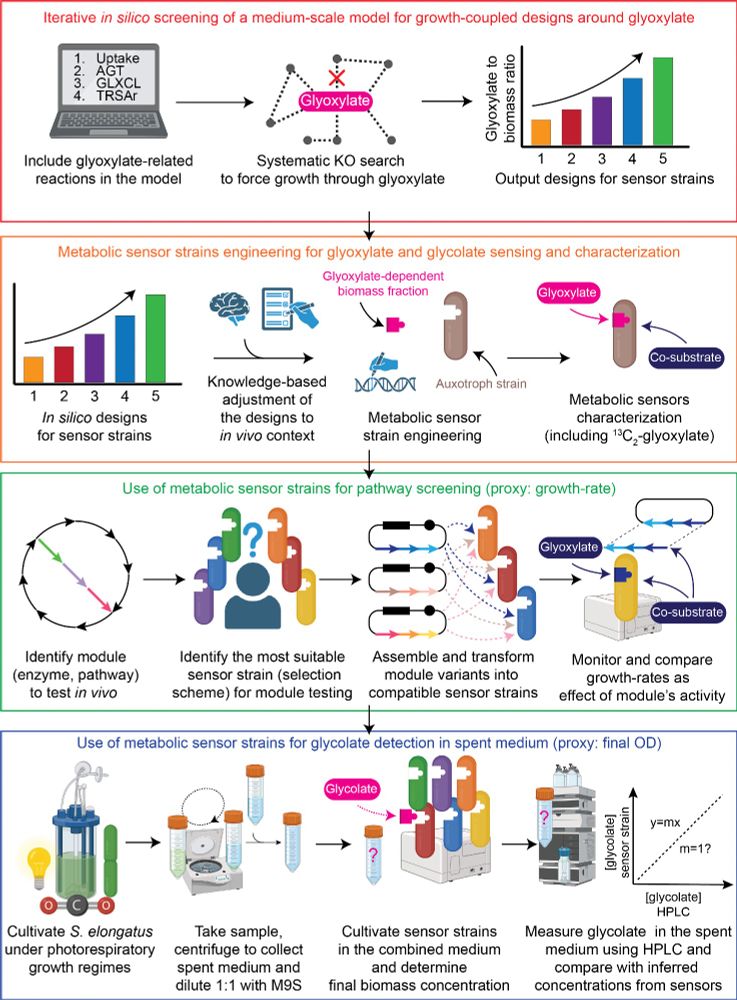
We just #published an end-to-end work for the computer-aided design💻, construction🛠️, validation✅, characterization🔬 & application💪 of #auxotroph metabolic sensors, the work-horses of in vivo growth-coupled selection screening systems-using glycolate as test case. 🧵👇1/
www.nature.com/articles/s41...
04.03.2025 20:38 —
👍 12
🔁 9
💬 1
📌 2
9) It was a true pleasure to work with Enrico Orsi, Hai He, Elad Noor, Ari Satanowski, Pablo Nikel, Tobias Erb and all co-authors involved. The origins of the strains go way back to the Bar-Even lab and Charlie Cotton creating the first strains - I am more than grateful to see it out !
04.03.2025 18:09 —
👍 2
🔁 0
💬 0
📌 0
8) The sensor strains are supposed to become a #community resource – so please feel free to reach out to us if you could apply them in your projects! This work would not have been possible without Enrico Orsi doing a tremendous job at leading it.
04.03.2025 18:08 —
👍 3
🔁 0
💬 1
📌 0
7) Of course, you can also use them to detect glyoxylate or glycolate in #environmental applications. For example, we could quantify how much glycolate was produced by #cyanobacteria during #photorespiratory growth based on the OD our sensors reached.
04.03.2025 18:06 —
👍 1
🔁 0
💬 1
📌 0
6) What can you use those sensors for ? Amongst other things, to select for synthetic metabolic modules. As an example in the context of #C1assimilation, we engineered a module of the HOPAC cycle and show that we can determine the best #expression strength for the catalysts.
04.03.2025 18:06 —
👍 1
🔁 0
💬 1
📌 0
5) Next, we characterised the strains by verifying expected metabolic fluxes by isotope tracing and determining their selective demands with #glyoxylate and #glycolate.
04.03.2025 18:06 —
👍 1
🔁 0
💬 1
📌 0