
Wanna compare dynamics across neural data, RNNs, or dynamical systems? We got a fast and furious method🏎️
The 1st preprint of my PhD 🥳 fast dynamical similarity analysis (fastDSA):
📜: arxiv.org/abs/2511.22828
💻: github.com/CMC-lab/fast...
I’ll be @cosynemeeting.bsky.social - happy to chat 😉
08.01.2026 16:07 —
👍 114
🔁 35
💬 1
📌 4

netneurotools: a trainee-oriented approach to network neuroscience | doi.org/10.1101/2025...
Our lab’s internal toolkit for accomplishing everyday tasks in brain imaging ⤵️
18.12.2025 14:58 —
👍 74
🔁 36
💬 1
📌 3
Thanks for reading! Do reach out to us if you plan to use cuBNM in your study, we are happy to help 🙂
And many thanks to @sofievalk.bsky.social, @sbe.bsky.social, @borismontreal.bsky.social, @wanb.bsky.social and all the collaborators on this project 🙏🏻
21/21
14.11.2025 17:29 —
👍 2
🔁 0
💬 0
📌 0
This started as a small weekend curiosity and turned into a very fun project.
I learned a lot along the way, and I hope cuBNM helps others with their modeling studies.
19/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

A few technical details: The frontend is written in Python (for ease of use), but to optimize efficiency, the simulation core is written in CUDA and C++. And not just simulations, but also calculation of FC, FC dynamics and goodness of fit measures is done on GPUs 🚀
17/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

As examples of the utility of these individualized models, we investigated test-retest reliability and heritability of simulated (and empirical) measures in the HCP dataset, and found simulated features to be fairly reliable and heritable.
15/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
This took a total of 1,426 hours on A100 GPUs (still considerable!) but multi-core supercomputer CPU nodes would have required 11x more time, 31x more cost and 14x more energy to do the same!
14/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
A key advantage of cuBNM’s GPU implementation is to make it possible to fit large numbers of individualized BNMs, in batches!
To demonstrate this, we built individualized models for 430 HCP participants, running >28 million simulations.
13/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

cuBNM also supports regional heterogeneity of parameters, using map-based and node-based approaches.
To demonstrate these approaches in the preprint, we applied them to fit group-level models of the HCP data, and as expected, found them to outperform the homogenous model.
12/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0


Two optimization approaches are included in cuBNM:
1️⃣ grid search
2️⃣ evolutionary optimizers (via pymoo, with CMA-ES as the featured method)
We demonstrate both in the preprint by fitting the rWW model to the group-averaged HCP data.
11/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

cuBNM currently includes five commonly used models (with rWW model featured in tutorials and the preprint).
New models can be added via YAML model specification files. And a guide for contributing new models is included in the documentation.
10/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
Tutorials - cubnm documentation
Check out our comprehensive tutorials cubnm.readthedocs.io/en/stable/tu... and read the preprint to learn how to 😊
And feel free to get in touch if you needed help.
9/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

Using cuBNM, if you have access to a few GPUs*, you can quickly and easily create individualized BNMs for your own dataset. All you need is: resting-state BOLD & structural connectivity (from each subject’s DWI, or a template SC).
* CPUs are also supported, but are not our focus.
8/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

Why are GPUs so much faster?
GPUs are designed for parallel processing, so computations across both *simulations* and *nodes* can be massively parallelized 🧠⚙️
7/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

GPU implementation also makes it feasible to simulate denser networks with thousands of nodes: Compute time with number of nodes increases near-quadratically on CPUs, but near-linearly on GPUs.
6/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0

On GPU, compared to single-core CPU, simulations can run up to 1000+ times faster (typically: 400-800x faster).
For example, running 32,768 simulations would take 3.8 *days* on a single-core CPU, but only 5.6 *minutes* on a data-center A100 GPU ⚡
5/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
So we thought: why not use GPUs to parallelize simulation calculations and make them faster? We tried it, and … it paid off really well 🥳Then we turned it into the cuBNM toolbox!
4/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
We experienced this directly during my PhD work on adolescent maturation of E-I ratio. We needed to fit individualized models for several hundred subjects, using more complex models, and quickly ran into the limits of CPU-based implementations.
3/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
Brain network models (BNMs) are great tools for investigating latent neural properties at micro-/mesoscale (such as excitation-inhibition ratio) using simulations. Despite their usefulness, the computational cost of simulation has limited their application at larger scales.
2/n
14.11.2025 17:29 —
👍 0
🔁 0
💬 1
📌 0
✨ New preprint ✨
Introducing cuBNM, a toolbox we developed to run brain simulations *much* more efficiently on GPUs 🎉🚀
Preprint: www.biorxiv.org/content/10.1...
Documentation: cubnm.readthedocs.io/en/stable/
GitHub: github.com/amnsbr/cubnm
1/n 🧵
14.11.2025 17:29 —
👍 9
🔁 3
💬 1
📌 0
Excited to share that our work introducing the Reproducible Brain Charts (RBC) data resource is now published in Neuron!! 🎉
📚 Read the paper: authors.elsevier.com/c/1lpaF3BtfH...
🧠 Explore the RBC dataset: reprobrainchart.github.io
22.09.2025 21:51 —
👍 71
🔁 39
💬 3
📌 2
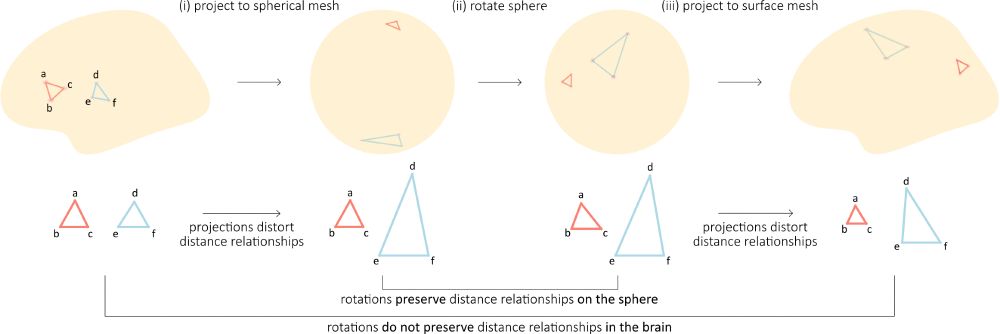
The effect of spherical projections on spin tests for brain maps | doi.org/10.1162/IMAG...
Spin tests are the de facto null model for map-to-map comparisons in brain imaging. Why don't they perfectly control false positives? @vincebaz.bsky.social explores ⤵️
10.09.2025 13:06 —
👍 25
🔁 14
💬 1
📌 1

🚨 Excited to share the latest preprint form the lab ➡️https://tinyurl.com/32d3be9f
Here we tackle a long-standing chicken-or-egg 🐣🥚question in #autism and developmental neuroscience
➡️ Is excitation–inhibition (E:I) imbalance a "cause" or a "consequence" of #autism?
Check out what we found!
🧵1/n
10.09.2025 15:08 —
👍 64
🔁 34
💬 1
📌 5
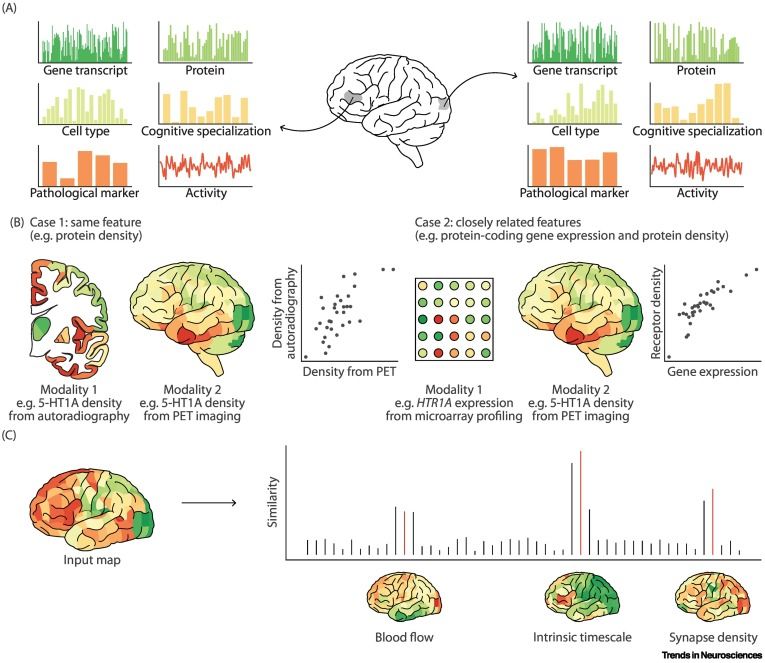
Integrating and interpreting brain maps | doi.org/10.1016/j.ti...
Imaging and recording technologies make it possible to map multiple biological features of the brain. How can these features be conceptually integrated into a coherent understanding of brain structure and function? ⤵️
04.08.2025 14:26 —
👍 74
🔁 42
💬 1
📌 0
My lab is at @ohbmofficial.bsky.social in Brisbane! Find us at the @gradientsworkshop.bsky.social - great talks by @wanb.bsky.social Katerina Manoli (and others)
Then we are at the educational sessions where @claraweber.bsky.social will present @amnsbr.bsky.social work on E/I balance.
21.06.2025 11:47 —
👍 16
🔁 4
💬 1
📌 0
... and functional changes e.g., faster and stronger IPSCs, and an increase in parvalbumin expression which (at least in one study) was not mirrored by an increase in PV cell counts. I can recommend Caballero 2014 Brain Struct Func, Caballero 2016 Neuro Biobeh Rev, and Hoftman 2017 Biol Psychiatry.
06.06.2025 18:22 —
👍 1
🔁 0
💬 1
📌 0

















