Robin Herbert
PhD Candidate, UC Berkeley
ORCHID ID: 0000-0002-9174-1562
Robin Herbert
PhD Candidate, UC Berkeley
ORCHID ID: 0000-0002-9174-1562
If you're looking for someone to use large-scale genetic data to understand how the ecology of the microbiome impacts it's function I would love to talk!
26.11.2025 19:40 — 👍 1 🔁 2 💬 1 📌 0Publications include work in Microbial Cell Factories, Environmental Toxicology and Chemistry, Frontiers in Microbiology, Nature Reviews Microbiology, etc.
26.11.2025 19:40 — 👍 0 🔁 0 💬 1 📌 0My long term goals are to use functional genomics, barcoding, and microbial ecology to learn how microorganisms cooperate, compete, and colonize their hosts!
26.11.2025 19:40 — 👍 0 🔁 0 💬 1 📌 0
In doing so I've developed skills in:
--High-throughput sequencing and analysis
--Strain engineering (RB-TnSeq, Cas9, VcDART)
--Plant-microbe / community interaction experiments
--Analysis of large datasets in R and Python
I've also genetically modified microorganisms for a variety of purposes, from the production of commodity chemicals to measuring the genes involved in colonization of the Sorghum rhizosphere using RB-TnSeq
26.11.2025 19:40 — 👍 0 🔁 0 💬 1 📌 0Specifically, I developed BarTn7, a transposon-based DNA barcoding system which enables sub-species level tracking in model and non-model bacteria, which I used to draw some novel conclusions about Switchgrass root colonization (see thread below)
26.11.2025 19:40 — 👍 0 🔁 0 💬 1 📌 0My work generally involves synthetic biology, microbial ecology, and plant-microbe interactions. I've worked with genetic and computational tools to track bacterial lineages and understand how bacteria adapt and interact during #microbiome assembly
26.11.2025 19:40 — 👍 0 🔁 0 💬 1 📌 0More exciting news! I'm finishing my PhD in plant biology from @ucberkeleyofficial.bsky.social this December and am looking for #postdoc positions starting in early 2026!
26.11.2025 19:40 — 👍 1 🔁 0 💬 1 📌 1And thank you to everyone in the Deutschbauer lab and Dr.s @tlowepower.bsky.social and Johan Leveau for the mentorship and advice! This was a great project to work on and I can’t wait to apply it to fun host-microbe interactions!
25.11.2025 22:38 — 👍 1 🔁 0 💬 0 📌 0If you are interested in using BarTn7 for your own ecological or evolutionary work please feel free to reach out to amdeutschbauer@lbl.gov. We would love to collaborate!
25.11.2025 22:38 — 👍 2 🔁 0 💬 1 📌 0
Finally, we made “magic pools” of these BarTn7 vectors, which use a parts-based strategy to screen promoters and antibiotic markers in new species. These magic pools identified vector designs which enabled BarTn7 in bacteria which were recalcitrant to the original plasmids!
25.11.2025 22:38 — 👍 1 🔁 0 💬 1 📌 0
We then took steps to optimize the system by reversing cargo orientation to prevent host toxicity and combining transposon and transposase into one vector, improving conjugation efficiency by ~10X
25.11.2025 22:38 — 👍 0 🔁 0 💬 1 📌 0
BarTn7 was also more accurate than 16S sequencing at measuring community composition when different barcoded species were combined and outgrown in different media, and comparable to Illumina shotgun whole-genome sequencing (WGS) at a fraction of the cost
25.11.2025 22:38 — 👍 0 🔁 0 💬 1 📌 0
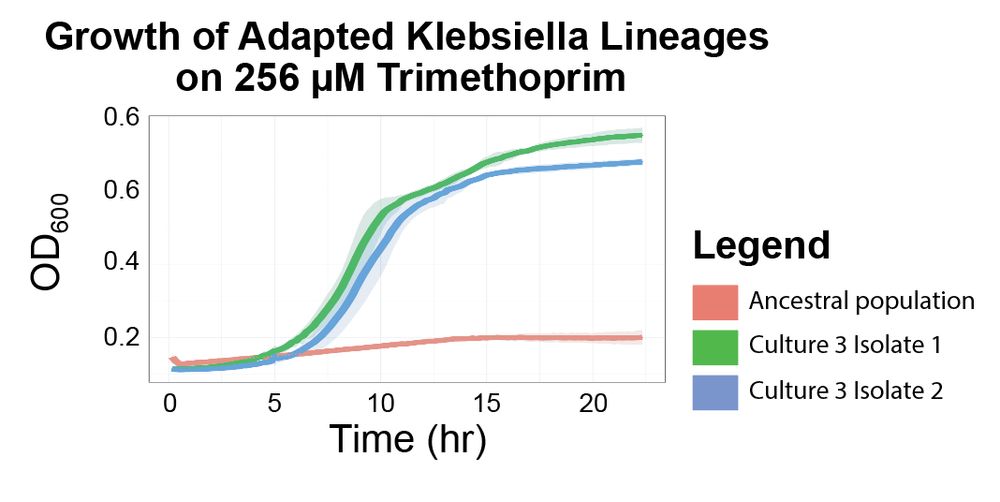
Next, we show that BarTn7 enables evolution experiments by identifying adaptive lineages (and the rise and fall of rare lineages). We detected selective sweeps linked to causal folA mutations in E. coli and Klebsiella treated with trimethoprim
25.11.2025 22:38 — 👍 0 🔁 0 💬 1 📌 0
Then we use BarTn7 to measure the colonization of Switchgrass roots by three plant-associated bacteria and find that lineage recruitment was species-specific: Pseudomonas (WCS417) often outcompeted lineages of the same species, while Paraburkholderia (Bfirm) colonized more evenly
25.11.2025 22:38 — 👍 0 🔁 0 💬 1 📌 0
First, we confirm that each lineage grows neutrally, showing population-level changes are due to drift or ecological dynamics rather than barcode bias
25.11.2025 22:38 — 👍 1 🔁 0 💬 1 📌 0

By chromosomally integrating DNA barcodes using the Tn7 transposon we created near-isogenic populations tagged with unique barcodes to we can track lineage behavior at the sub-species level
25.11.2025 22:38 — 👍 0 🔁 0 💬 1 📌 0Populations of closely related bacteria interact in ways which change microbiome assembly, evolution, and host health/productivity, but can be hard to measure
25.11.2025 22:38 — 👍 1 🔁 0 💬 1 📌 0
Very excited to share a big part of my dissertation work with the Deutschbauer lab at LBNL and @ucberkeleyofficial.bsky.social! BarTn7: A method for bacterial lineage tracking at sub-species resolution in population, ecological, and evolutionary experiments.
www.biorxiv.org/content/10.1...