
Excited to share our new paper! We developed a method to visualize proteasomal degradation at the single–molecule level in live cells, enabling us to dissect distinct modes of substrate engagement, probe co-factor dependence, and study proteasome–ribosome collisions.
www.biorxiv.org/content/10.6...
20.01.2026 12:27 —
👍 48
🔁 15
💬 1
📌 1
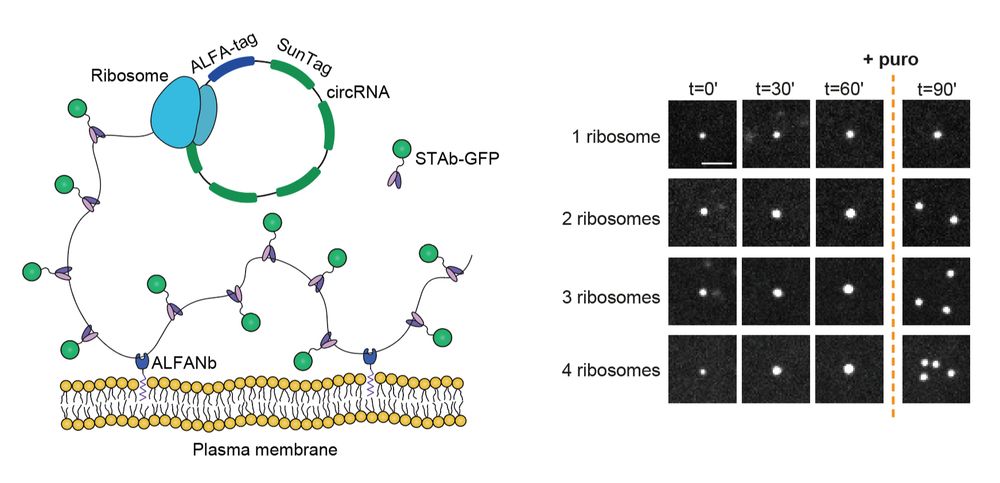
Our paper is out! We delevoped a method to follow individual translating ribosomes for hours in living cells, and discovered that ribosomes are great friends and help each other in problematic situations:
www.cell.com/cell/fulltex...
03.02.2025 08:31 —
👍 43
🔁 11
💬 0
📌 0
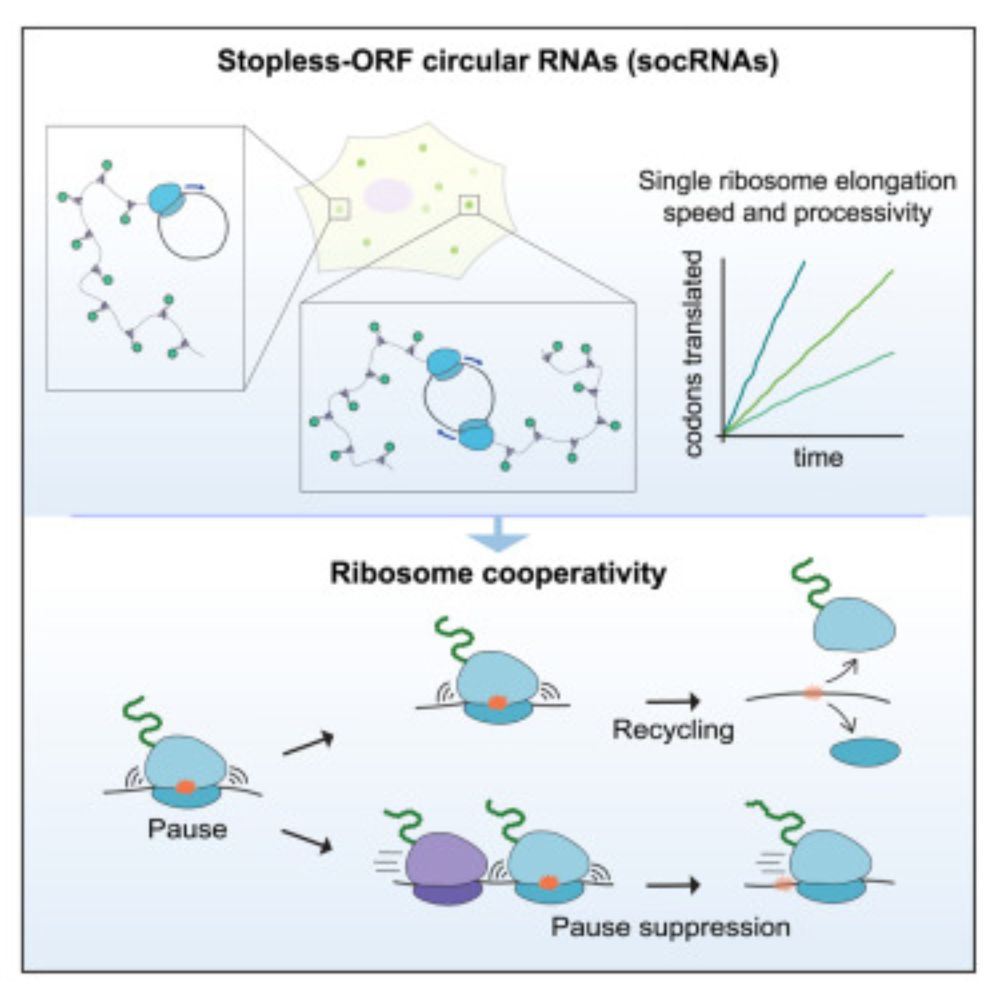
Long-term imaging of individual ribosomes reveals ribosome cooperativity in mRNA translation
Ribosomes cooperate through transient collisions to ensure efficient translation.
Our paper on Stopless-ORF Circular RNAs (socRNAs) is now out in Cell. By high-res tracking and comparing translation by either single or multiple ribosomes, we find that ribosomes cooperate to overcome pausing to ensure fast and efficient translation
www.cell.com/cell/fulltex...
03.02.2025 07:38 —
👍 114
🔁 47
💬 4
📌 6
Very happy to share our preprint on visualizing the life cycle of Influenza viruses using single-molecule imaging! 🥳 We developed two techniques to visualize infections of unmodified influenza viruses in live cells from endosomal release to budding of new viruses. For more details&videos see below ⬇️
21.01.2025 10:16 —
👍 65
🔁 14
💬 2
📌 1



