
Announcing the joint @idpseminars.bsky.social / BPS IDP Subgroup Trainee Symposium Speakers! Join us on Mar 27, 12–2pm CST for 6 exciting talks focused on Intrinsically Disordered Proteins as part of #BophysicsWeek2025 with the @biophysicalsoc.bsky.social Sign up now using the QR code below!
17.03.2025 15:37 —
👍 8
🔁 3
💬 0
📌 1

Announcing the joint @idpseminars.bsky.social / BPS IDP Subgroup Trainee Symposium Speakers!
Join us on Mar 27, 12–2pm CST for 6 exciting talks focused on Intrinsically Disordered Proteins as part of #BophysicsWeek2025 with the @biophysicalsoc.bsky.social Sign up now using the QR code below!
17.03.2025 15:38 —
👍 16
🔁 8
💬 0
📌 1

IDPSeminars/BPS IDP Subgroup symposium. March 27th 2025 at noon central time. Speakers are Ella Mozier, Sara Bologna, Borna Novak, Hamidreza Ghafouri, Ankith Sharma and Ananya Chakravarti!
Announcing the joint IDPSeminars / BPS IDP Subgroup Trainee Symposium Speakers!
Join us on Mar 27, 12–2pm CST for 6 exciting talks focused on Intrinsically Disordered Proteins as part of #BophysicsWeek2025 with the @biophysicalsoc.bsky.social
Sign up now at docs.google.com/forms/d/1L_K...
17.03.2025 16:14 —
👍 20
🔁 10
💬 0
📌 4
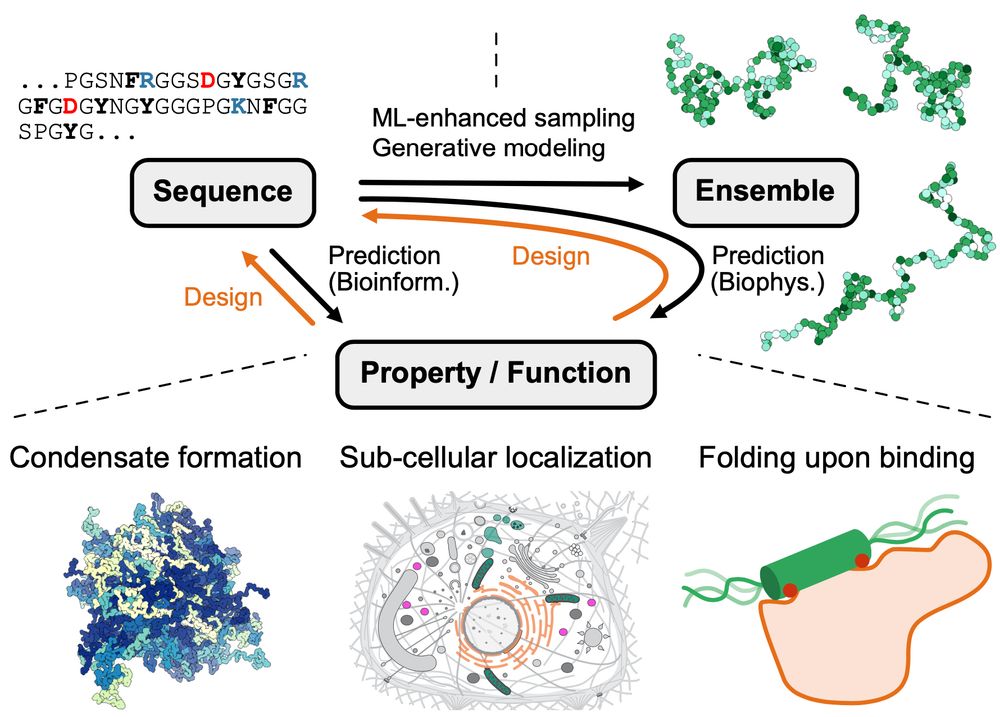
Figure from the paper illustrating sequence–ensemble–function relationships for disordered proteins. ML prediction (black) and design (orange) approaches are highlighted on the connecting arrows. Prediction of properties/functions from sequence (or vice versa, design) can include biophysics approaches via structural ensembles, or bioinformatics approaches via other hetero- geneous sources. The lower panels show examples of properties and functions of IDRs for predictions or design targets. ML, machine learning; IDRs, intrinsically disordered proteins and regions.
Our review on machine learning methods to study sequence–ensemble–function relationships in disordered proteins is now out in COSB
authors.elsevier.com/sd/article/S...
Led by @sobuelow.bsky.social and Giulio Tesei
12.03.2025 21:37 —
👍 91
🔁 27
💬 0
📌 1
Now published in JCIM! Thanks to reviewers and our editor for pushing us to (hopefully) clarify a few things and ensure it's clear what PENGUIN is trying to do (and what it's NOT trying to do).
Also, in case you were unsure, graphic design is our passion.
pubs.acs.org/doi/full/10....
05.03.2025 14:50 —
👍 35
🔁 9
💬 1
📌 1

HIF-1α ensemble dimensions change less under osmotic stress than with specific mutations. In contrast, CITED2 shows significant ensemble changes under stress. This suggests cellular environments might naturally regulate AD function through changes in ensemble dimensions.
26.02.2025 07:03 —
👍 0
🔁 0
💬 1
📌 0

Mutations expanding the HIF-1α ensemble correlated with higher activity, while compaction correlated with reduced activity. For CITED2, ensemble size didn’t impact activity. Our findings suggest that ensemble dimensions may influence transcriptional activity in some ADs.
26.02.2025 07:03 —
👍 0
🔁 0
💬 1
📌 0

We studied two activation domains from HIF-1α and CITED2, using live-cell FRET microscopy to measure changes in their ensemble dimensions.
26.02.2025 07:03 —
👍 0
🔁 0
💬 1
📌 0
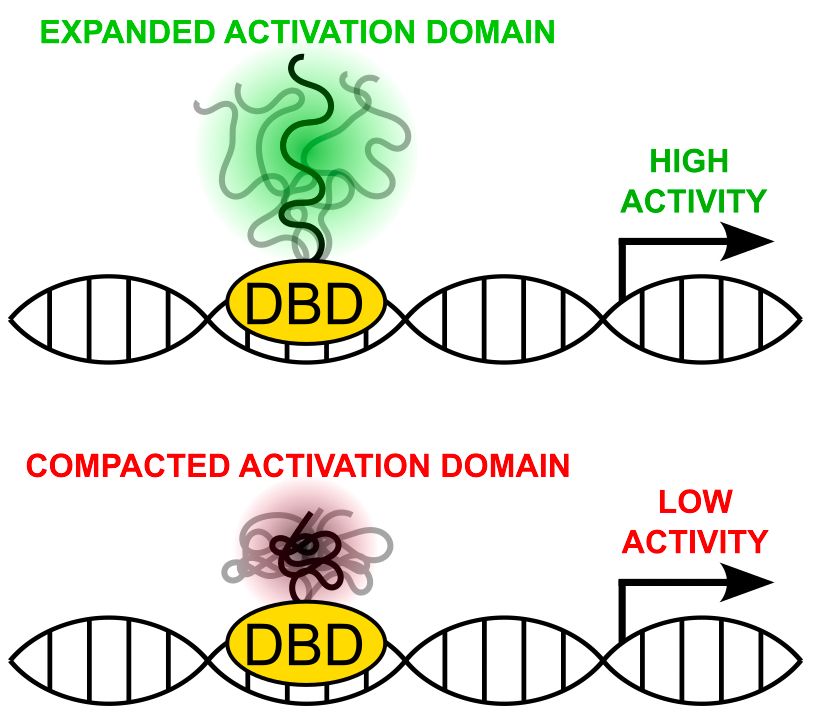
📢 Excited to share our research with @shaharsu.bsky.social & @mvstaller.bsky.social on how activation domain (AD) structure affects gene expression! 🧬 TLDR: HIF-1α & CITED2 ADs suggest ensemble dimensions may regulate transcription in unique ways.
www.cell.com/biophysrepor...
🧵 Highlights:
26.02.2025 07:03 —
👍 10
🔁 3
💬 1
📌 0
mvstaller.bsky.social
I’m grateful to Max Staller (@mvstaller.bsky.social) for his insights, discussions, and for providing the activity data for our correlations. Deep thanks to (blueskyless?) Aleah Camacho, Estefania Cuevas-Zepeda, Mary McCoy, and Feng Yu for their contributions throughout the project!
26.02.2025 06:54 —
👍 0
🔁 0
💬 0
📌 0

HIF-1α ensemble dimensions change less under osmotic stress than with specific mutations. In contrast, CITED2 shows significant ensemble changes under stress. This suggests cellular environments might naturally regulate AD function through changes in ensemble dimensions.
26.02.2025 06:54 —
👍 0
🔁 0
💬 1
📌 0

Mutations expanding the HIF-1α ensemble correlated with higher activity, while compaction correlated with reduced activity. For CITED2, ensemble size didn’t impact activity. Our findings suggest that ensemble dimensions may influence transcriptional activity in some ADs.
26.02.2025 06:54 —
👍 0
🔁 0
💬 1
📌 0

We studied two activation domains from HIF-1α and CITED2, using live-cell FRET microscopy to measure changes in their ensemble dimensions.
26.02.2025 06:54 —
👍 0
🔁 0
💬 1
📌 0












