🧬 New paper out!
Thrilled to contribute to our study on CaV1.3 channel clusters in cardiac myocytes using DNA-PAINT (4 nm loc. precision!) revealing flexible (not rigid) cluster organization.
👏 Great work by Niko Schwenzer, @samratb.bsky.social & team!
👉 www.sciencedirect.com/science/arti...
10.11.2025 13:06 —
👍 2
🔁 1
💬 0
📌 0

It was fun to share my passion for science at the #EuropeanResearchersNight in Groningen!
✨ Curious visitors
✅matched 🧫 plates with microbes to where they were collected
✅inspected flowers, berries, and lichens under the microscope🔬
✅ created their own “bacterial selfies”📸
02.10.2025 16:22 —
👍 4
🔁 0
💬 1
📌 0

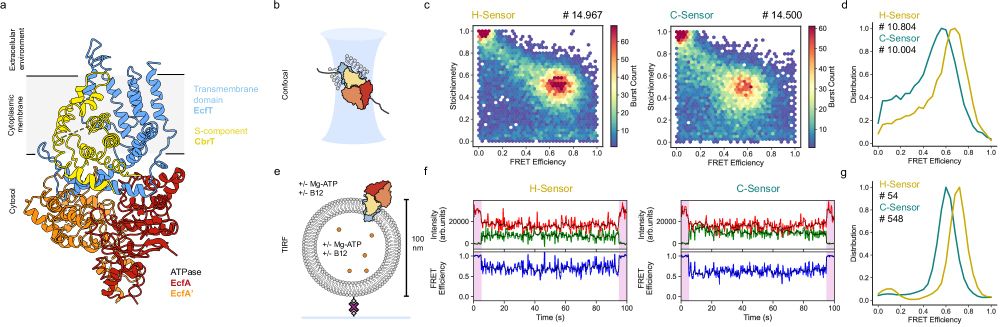
Finally, how to make sense of (1) sub-ms agonist-induced dynamics seen in confocal, and (2) long-lived states (>100ms) in ABEL trap. How do they not average out?
In our updated model, sub-ms dynamics occurs within long-lived states:
28.07.2025 13:27 —
👍 0
🔁 0
💬 1
📌 0

Maybe, these long-lived states can interconvert on much longer timescales - in fact, Wei et al. showed long-lived states with dwell times of seconds before with TIRF:
doi.org/10.1016/j.st...
28.07.2025 13:27 —
👍 0
🔁 0
💬 1
📌 0
We conclude that A2A receptors must have long-lived states with different FRET efficiencies with dwell times of at least hundreds of ms. This way, we extend our previous estimation, based on confocal burst data, ~100x fold!
28.07.2025 13:27 —
👍 0
🔁 0
💬 1
📌 0
it seems very much like "static heterogeneity":
within a "Gaussian" or FRET values, receptors on the left shoulder of the distribution stay there for 100s of milliseconds, as if they had "memory"
only metaphorically, of course ...
28.07.2025 13:27 —
👍 0
🔁 0
💬 1
📌 0
Instead of sharp PR peaks and step-like transitions, we see broad distributions and continuous FRET changes🤔 Is conformational space a spectrum?🙃
Our dynamics analysis (inspired by BVA, FRET-FCS, and RASP) showed that dynamics on 1-100 ms scale is significant but subtle, so...
28.07.2025 13:27 —
👍 0
🔁 0
💬 1
📌 0

With ABEL-trap, we can see individual receptors for up to 2 seconds instead of milliseconds! in solution and without tethers
Consistent with our previous data, agonist NECA increases FRET in double-labeled A2A
28.07.2025 13:27 —
👍 0
🔁 0
💬 1
📌 0

This is the final piece of my PhD studies on #smFRET with A2A adenosine receptors #GPCR!
In our previous study, we observed sub-ms agonist-induced conformational dynamics of the A2A, but longer transitions (if any) were invisible in burst-wise data
doi.org/10.1038/s420...
28.07.2025 13:27 —
👍 1
🔁 0
💬 1
📌 0
ABEL-FRET bridges the timescale gap in single-molecule measurements of the structural dynamics in the A2A adenosine receptor https://www.biorxiv.org/content/10.1101/2025.07.22.666123v1
26.07.2025 03:47 —
👍 3
🔁 1
💬 0
📌 0
It was a great Day 1 of the EMBL Symposium "New approaches and concepts in microbiology"! Stepping a little outside of my usual microscopy/biophysics bubble feels like the right kind of challenge - wish me more good ideas, inspiration, and coffee
#EESMicrobiology
24.06.2025 21:16 —
👍 6
🔁 0
💬 0
📌 0
⚛️Physics Olympiads hold a special place in my❤️
I cherish memories of my teachers and still enjoy the charm of the intellectual puzzles🥲
This year, I designed a few problems for the Dutch Physics Olympiads ⏳
Congratulations to the winners and good luck at #IPhO2025 🥳
11.06.2025 07:52 —
👍 1
🔁 0
💬 0
📌 0
beautiful pulsed labelings reveal where new wall form as bacteria grow
03.06.2025 04:33 —
👍 11
🔁 1
💬 0
📌 0
👏Loud applause to @linnikdmitrii.bsky.social, who came up with idea and combined so many methods to characterize the system from every possible angle! And thanks to everyone who contributed to the study! 🪅
@membraneenzymology.bsky.social 🔬🧫
@janstevens.bsky.social @cg-martini.bsky.social 🤖🧑💻
26.05.2025 11:39 —
👍 2
🔁 0
💬 0
📌 0
🚨 #LLPS: New trick for an old membrane protein!
Homo- and hetero- biomolecular condensates of a native membrane protein in E. coli tagged with PopTag 🔬🦠
LacY-PopTag is functional and localizes at cell poles or mobile foci in a curvature-dependent manner
🔗 doi.org/10.21203/rs.3.rs-6571918/v1
26.05.2025 11:25 —
👍 2
🔁 2
💬 1
📌 0
Giulia Viola, Torsten Wittmann @microtubule.bsky.social and colleagues @ctbatucsf.bsky.social use protein nanocages to benchmark fluorescent proteins in live cells.
Highlight: journals.biologists.com/jcs/article/...
Article: journals.biologists.com/jcs/article/...
#OpenAccess #ReadandPublish
23.05.2025 12:02 —
👍 13
🔁 4
💬 1
📌 0

Register: forms.gle/4hDdwKNhXMJj...
Happening TOMORROW!
Join us on April 25 at 7PM EAT for a powerful webinar with @atinygreencell.bsky.social on making synthetic biology accessible to all.
Hosted by @synbio4allafrica.bsky.social .
24.04.2025 07:45 —
👍 18
🔁 6
💬 0
📌 0

We are looking for a postdoc with microbiology background for #superresolution #singlemolecule #ERC tracking project. Apply by 27.4.2025. 🦠🔬🧫
Pls share!
www.aalto.fi/en/open-posi...
01.04.2025 07:30 —
👍 25
🔁 23
💬 1
📌 1

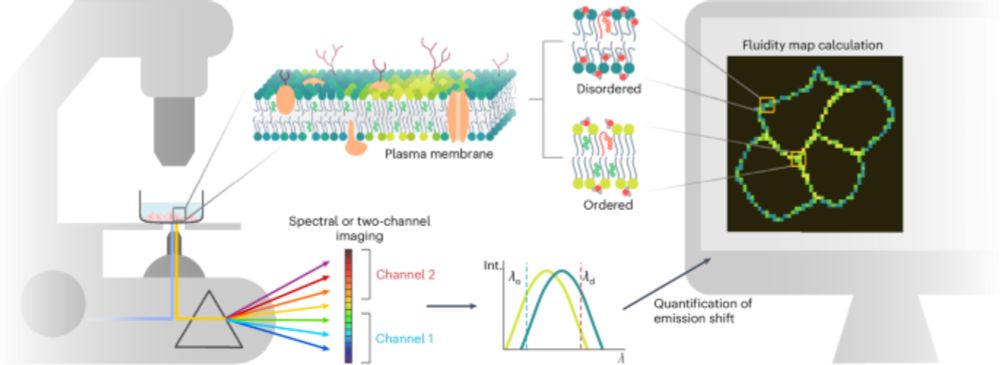
Lipid packing and cholesterol content regulate membrane wetting and remodeling by biomolecular condensates
Nature Communications - Membrane lipid packing, influenced by cholesterol, lipid chain length, and saturation, regulates the affinity of biomolecular condensates, enhancing their interactions with...
1/8 Exciting new work with Agu @mangiarotti.bsky.social @mpici.bsky.social ! 🚀 Our latest study explores how lipid packing and cholesterol regulate the interaction of biomolecular condensates with membranes. It turns out, it’s not just about membrane phase. Packing is the key! 🧵👇
rdcu.be/eelLq
21.03.2025 08:46 —
👍 40
🔁 11
💬 1
📌 0
New Night Science Podcast episode! David Baker was awarded the 2024 Nobel Prize in Chemistry. He explains how he socially engineers his lab’s "communal brain", where all individuals function like neurons, densely interconnected to maximize idea generation.
nightscience.buzzsprout.com/1744020/epis...
25.03.2025 07:30 —
👍 35
🔁 9
💬 3
📌 0

Protein function often depends on protein dynamics. To design proteins that function like natural ones, how do we predict their dynamics?
@hkws.bsky.social and I are thrilled to share the first big, experimental datasets on protein dynamics and our new model: Dyna-1!
🧵
20.03.2025 15:02 —
👍 103
🔁 38
💬 6
📌 5

A comprehensive review on covalent chemical strategies for protein self-labeling from @kjohnsson.bsky.social's lab: HaloTag, SNAP-tag et al. www.annualreviews.org/content/jour...
20.03.2025 20:56 —
👍 65
🔁 21
💬 1
📌 0

Single-molecule microscopy is full of surprises:
see how anisotropy can be used to follow the conformational dynamics! 🤯 #24-state-model
"Examination of conformational dynamics of AdiC transporter with fluorescence-polarization microscopy" @jgp.org 🔥🔬
rupress.org/jgp/article/...
26.02.2025 19:15 —
👍 3
🔁 0
💬 0
📌 0