Junior research scientist in modelling for genomic prediction using complex data
CR26-GA-1 - You will join the Genetics, Physiology and Livestock Systems (GenPhyse, about 150 permanent staff) joint research unit, where researchers aim to contribute to the agroecological transition of livestock systems through better understanding of livestock biology, genetic bases of traits, and selection schemes to achieve resilient populations. The unit brings together skills in biology, physiology, genomics, genetics, statistics, and bioinformatics. Within the unit, you will join the Chamade team (Characterization and management of genetic diversity) of the Diversity and Selection group comprising methodologists in quantitative genetics (genomic prediction, selection and evolution) and population genomics, as well as statisticians. Within your research team, you will be in charge of developing a new research programme in applied statistics to integrate high-throughput, heterogenous, and multiscale data into genetic and genomic evaluation methods. You will conduct your research to improve genomic prediction models by integrating new information relative to genome function (e.g. functional annotation), molecular phenotypes (e.g. transcriptomics, methylation), and high-throughput or intermediate phenotypes (e.g. longitudinal data, high throughput sensors).To integrate different types of data into current genomic prediction models, you will draw on a variety of statistical modelling, for example modelling SNP effects according to their functional annotation category, or including random effects capturing the inter-individual covariance for intermediate phenotypes. The modelling could draw, for example, on hierarchical models, meta-analysis methods, mediation analysis or machine learning, possibly simulation-based. In addition, as new high-throughput data may not necessarily be available on the same individuals as traditional data, their integration will require the implementation of suitable statistical techniques. The predictive performance of the developed models will be evaluated using numerical simulations, for instance based on real breeding programmes. The models will also be tested on real data from experimental and commercial programmes of livestock species. These data will be available through existing projects and partnerships within the unit and the division to initiate your research project. Computational efficiency must be taken into consideration in your developments to ultimately ensure their practical use in genetic and genomic evaluation. To develop your research, you will benefit from the proximity of experts in statistics, computer science, quantitative, molecular and population genetics within GenPhySE. In accordance with INRAE's policy for open science, in addition to scientific publications, you will promote your work by distributing free software implementing the new methods developed to ensure their wide dissemination to the international community.
A new permanent position is opened in our group at the Animal Genetics Division of @inrae-france.bsky.social to work on the development of new prediction models for genomic selection in livestock.
Job description is here : urls.fr/FZNWJQ
Don't hesitate to contact me for more information !
28.01.2026 17:25 —
👍 3
🔁 7
💬 0
📌 0
Pleasure giving a recap of a wonderful PopGroup conference, and the opportunity to talk about my research.
🔥
03.02.2026 11:49 —
👍 21
🔁 11
💬 1
📌 0
Junior research scientist in modelling for genomic prediction using complex data
CR26-GA-1 - You will join the Genetics, Physiology and Livestock Systems (GenPhyse, about 150 permanent staff) joint research unit, where researchers aim to contribute to the agroecological transition of livestock systems through better understanding of livestock biology, genetic bases of traits, and selection schemes to achieve resilient populations. The unit brings together skills in biology, physiology, genomics, genetics, statistics, and bioinformatics. Within the unit, you will join the Chamade team (Characterization and management of genetic diversity) of the Diversity and Selection group comprising methodologists in quantitative genetics (genomic prediction, selection and evolution) and population genomics, as well as statisticians. Within your research team, you will be in charge of developing a new research programme in applied statistics to integrate high-throughput, heterogenous, and multiscale data into genetic and genomic evaluation methods. You will conduct your research to improve genomic prediction models by integrating new information relative to genome function (e.g. functional annotation), molecular phenotypes (e.g. transcriptomics, methylation), and high-throughput or intermediate phenotypes (e.g. longitudinal data, high throughput sensors).To integrate different types of data into current genomic prediction models, you will draw on a variety of statistical modelling, for example modelling SNP effects according to their functional annotation category, or including random effects capturing the inter-individual covariance for intermediate phenotypes. The modelling could draw, for example, on hierarchical models, meta-analysis methods, mediation analysis or machine learning, possibly simulation-based. In addition, as new high-throughput data may not necessarily be available on the same individuals as traditional data, their integration will require the implementation of suitable statistical techniques. The predictive performance of the developed models will be evaluated using numerical simulations, for instance based on real breeding programmes. The models will also be tested on real data from experimental and commercial programmes of livestock species. These data will be available through existing projects and partnerships within the unit and the division to initiate your research project. Computational efficiency must be taken into consideration in your developments to ultimately ensure their practical use in genetic and genomic evaluation. To develop your research, you will benefit from the proximity of experts in statistics, computer science, quantitative, molecular and population genetics within GenPhySE. In accordance with INRAE's policy for open science, in addition to scientific publications, you will promote your work by distributing free software implementing the new methods developed to ensure their wide dissemination to the international community.
A new permanent position is opened in our group at the Animal Genetics Division of @inrae-france.bsky.social to work on the development of new prediction models for genomic selection in livestock.
Job description is here : urls.fr/FZNWJQ
Don't hesitate to contact me for more information !
28.01.2026 17:25 —
👍 3
🔁 7
💬 0
📌 0
Portail Emploi CNRS - Offre d'emploi - Ingénieur d'études en bioinformatique (H/F)
Un poste d’IE en bioinformatique est actuellement ouvert au CEFE à Montpellier pour travailler dans le cadre de l'ERC RegEvol.
Détails ici:
emploi.cnrs.fr/Offres/CDD/U...
Début 01/01/2026 pour 18 mois potentiellement renouvelable.
candidature jusqu'au 29/11
N’hésitez pas à diffuser largement
18.11.2025 15:09 —
👍 3
🔁 14
💬 0
📌 0

Cross-species cloning in ants 🐜
These two males belong to different species—but share the same mother. How? Why?
To celebrate the print release of our last paper in this week’s @nature.com (issue 8084), here’s a thread summarizing the results. Why? Let’s dive in🧵👇 www.nature.com/articles/s41...
14.10.2025 12:00 —
👍 30
🔁 19
💬 1
📌 0
Population Genetics group 59
Exciting news!
The next #PopGroup meeting will take place in Lille 🍟, France, 7–9 January 2026 – just 1 hour by train from London, Brussels, and Paris.
This year, PopGroup will also host ALPHY, the annual meeting of Evolutionary Genomics.
More info: populationgeneticsgroup.org.uk
See you there !
29.09.2025 08:52 —
👍 46
🔁 53
💬 1
📌 2
“We are not working on building an AI agent. We are working to protect Signal from the invasion of AI agents that threaten privacy and that are being implemented in irresponsible ways”
16.09.2025 06:58 —
👍 563
🔁 166
💬 2
📌 1
Redirecting
1/
Out now in Cell: Our new study uncovers the ancient origins of a genetic mutation that protects against HIV — and rewrites the story of its surprising high frequency in Europe.
Link: doi.org/10.1016/j.ce...
Let’s dig in (popular science first, jump to 16 for technical details)
06.05.2025 13:51 —
👍 9
🔁 4
💬 1
📌 2

Hybridization and introgression are major evolutionary processes. Since the 1940s, the prevailing view has been that they shape plants far more than animals. In our new study (www.science.org/doi/10.1126/...
), we find the opposite: animals exchange genes more, and for longer, than plants
12.09.2025 07:54 —
👍 204
🔁 121
💬 3
📌 3
Talk:Linkage disequilibrium - Wikipedia
Just want to share this wild revision history of the Wikipedia page on Linkage Disequilibrium where Felsenstein himself debated people on the correct use of notations & definitions.
After 100 years of #popgen people does not seem to be totally agree on what LD is?
en.wikipedia.org/wiki/Talk:Li...
22.07.2025 14:44 —
👍 16
🔁 9
💬 0
📌 0
Integration of pedigrees and methods for demographic inference in livestock population genomics
Demographic inference is a central tool for reconstructing the genetic history of a population. Recently, several new approaches have brought this tool into a new era, that of the exhaustive use of whole-genome sequence. These approaches can be divided into two categories: demographic inference based on ancestral recombination graphs (ARGs1) and deep learning based inference2,3. However, without any other source of information, it is difficult to assess the actual degree of accuracy of these methods. In the case of livestock species, we usually have access to an additional information, rather unique and valuable, the pedigree over several generations. This set of relationships makes it possible to compare inferred demographic histories with the exact, albeit incomplete, history of the population.In this thesis project, we aim to use the pedigree information on the one hand, to assess the results obtained with the new approaches of demographic inference, and, on the other hand, to refine methods for ARG estimation. Therefore, we propose a research program divided into three main tasks: (i) to appropriate new approaches (ARG, deep learning) in demographic inference on goat and sheep datasets, (ii) to compare these approaches together, in particular to evaluate their inference of the effective size of a population in the light of genealogical information, and (iii) to integrate this information into the inference of ARGs.Kelleher, J. et al. Inferring whole-genome histories in large population datasets. Nature Genetics 51, 1330–1338 (2019).Schrider, D. R. & Kern, A. D. Supervised Machine Learning for Population Genetics: A New Paradigm. Trends in Genetics 34, 301–312 (2018).Korfmann, K., Gaggiotti, O. E. & Fumagalli, M. Deep learning in population genetics. Genome Biology and Evolution 15, evad008 (2023).You will be welcomed in the CHAMADE team (“CHAracterization and MAnagement of Diversity”) of the GenPhySE research unit (https://genphyse.inrae.fr/), located in the Occitanie-Toulouse research centre (31320, Castanet-Tolosan). The CHAMADE team is part of the “Diversity and Selection” scientific division of the research unit. The team is interested in methodological issues in the field of population genomics, genetic evaluation of livestock species and quantitative and evolutionary genetics. On deep learning approaches for demographic inference, collaboration is also planned with the BioInfo team from the Laboratoire Interdisciplinaire des Sciences du Numérique (LISN, Paris-Saclay University).
I'd like to advertise a PhD Position opened in our INRAE lab in Toulouse with Pierre Faux and myself to work on evaluating methods to infer demography of livestock populations 🐐 🐏 ...
More details here : jobs.inrae.fr/en/ot-25908 and even more after contacting us :) Applications are open !
18.06.2025 14:56 —
👍 1
🔁 1
💬 0
📌 0
In 2010 (SMBE Lyon) I gave a talk about the impact of GC-biased gene conversion (gBGC) on functional sequence evolution. I argued that, because gBGC promotes G and C alleles irrespective of their fitness effect, it should generate some genetic load. 1/5
11.01.2025 14:35 —
👍 31
🔁 21
💬 2
📌 1

a cartoon says hey everybody an old man 's talking while bart simpson looks on
ALT: a cartoon says hey everybody an old man 's talking while bart simpson looks on
Bluetorial-A dream and a bit of a nightmare
Serving as Editor-in-Chief at Science was fascinating. I greatly enjoyed working with talented and committed editorial, news, graphics, and production staff. But the inside look into scientific publishing and AAAS was also deeply disillusioning.
21.01.2025 17:56 —
👍 479
🔁 192
💬 17
📌 77
Partitioning the phenotypic and genetic variances of reaction norms
Excited to share the news that our article on the partition of variance of reaction norm with @lmchev.bsky.social has been recommended by @peercommunityin.bsky.social Evolutionary Biology: 👇🧵
ecoevorxiv.org/repository/v...
evolbiol.peercommunityin.org/articles/rec...
#phenplas #quantgen
21.01.2025 07:57 —
👍 30
🔁 13
💬 2
📌 2

Genome Editing and Eugenics
The one hundred and third Take:
Thoughtful piece by Greg Gibson pushing strongly back on the "Heritable polygenic editing" article
genomestake.substack.com/p/genome-edi...
16.01.2025 22:49 —
👍 51
🔁 36
💬 4
📌 4
Yup
13.12.2024 15:52 —
👍 2483
🔁 495
💬 23
📌 30

We MUST keep vaccines.
have you written your Senator to let them know?
14.12.2024 20:03 —
👍 37037
🔁 5994
💬 358
📌 176
Today I.J. Good is getting spicy about hypothesis testing.
All quotes are from "Good Thinking", store.doverpublications.com/products/978....
12.12.2024 15:34 —
👍 1
🔁 1
💬 1
📌 0
“Null hypotheses are usually known in advance to be false, and the point of significance tests is usually to find out wether they are nevertheless approximately true.”
12.12.2024 15:34 —
👍 1
🔁 1
💬 1
📌 0
Ce n'est pas parce qu'on manifeste contre un président accusé de haute trahison qu'on ne peut pas le faire avec le sens de l'humour : au cours de la semaine passée, plusieurs drapeaux de manifestants ont attiré l’attention pour leur côté décalé. Traditionnellement, ces drapeaux étaient là pour
09.12.2024 17:58 —
👍 633
🔁 347
💬 19
📌 60

I am looking for a postdoc! Pls share
#postdoc #genomics #insects #inversions
genomicsofoutbreaks.com/?page_id=141
10.12.2024 12:14 —
👍 19
🔁 29
💬 1
📌 1
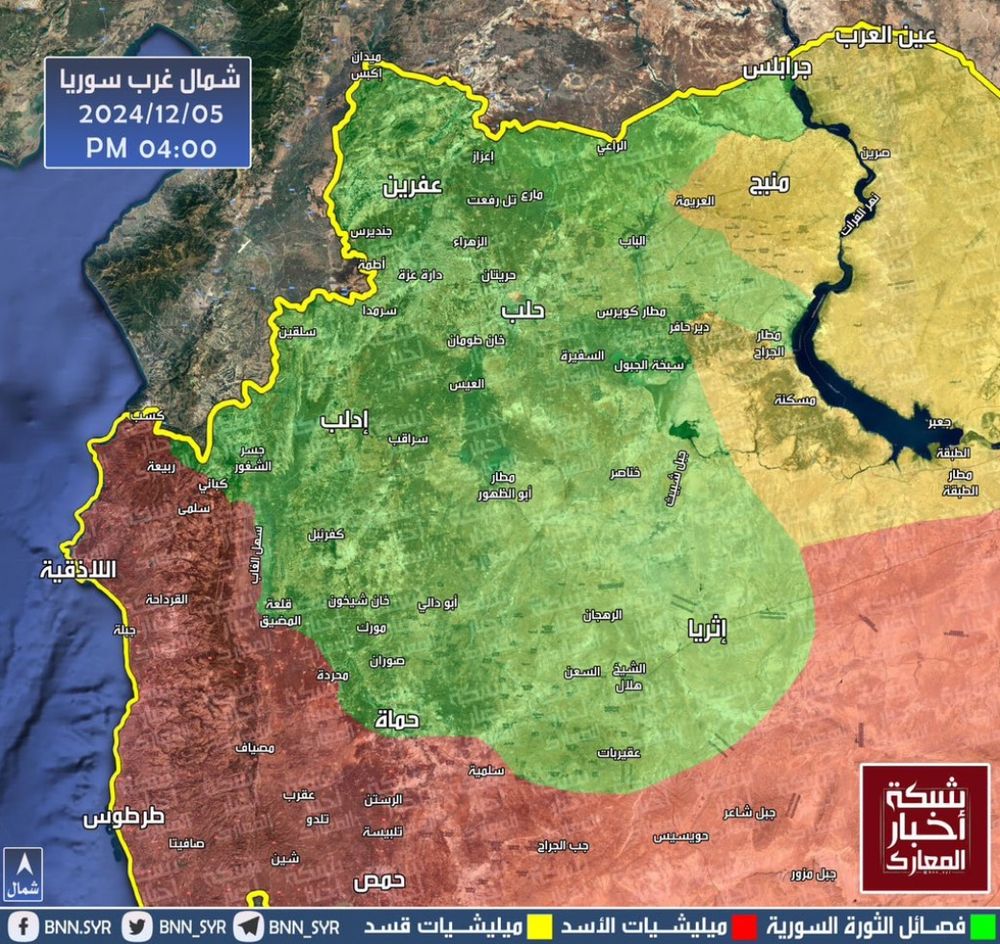
Having worked on #Syria full-time since the crisis began nearly 14yrs ago, there really is no understating how remarkable the losses imposed on #Assad's regime have been over the past week.
A large reason for this lies with #HTS — a 🧵:
05.12.2024 16:02 —
👍 2519
🔁 813
💬 77
📌 216

🧬🔍There are 50 petabases of freely-available DNA sequencing data. We introducing Logan Search which allows you to search for any DNA sequence in minutes, bringing Earth’s largest genomic resource to your fingertips.
🏔️ logan-search.org 🏔️
#Genomics #Bioinformatics #OpenScience
11.11.2024 19:29 —
👍 108
🔁 56
💬 2
📌 4
GitHub - MichelNivard/awesome-complex-trait-genetics: A list of awesome tools for complex trait genetics.
A list of awesome tools for complex trait genetics. - MichelNivard/awesome-complex-trait-genetics
🚨 This will become a curated list of awesome tools for complex trait genetics, **add yours**! it may become a review in which case those who contribute are invited as co-authors.
28.11.2024 09:21 —
👍 80
🔁 43
💬 7
📌 4















