
Wow. Catalonia has more active clinical trials than the entirety of the UK or Germany. That's wild. From the latest @biocat.cat annual report.
21.02.2026 06:56 — 👍 9 🔁 3 💬 0 📌 0@benlehner.bsky.social
Solve Biology Head of Generative Genomics, Wellcome Sanger Institute, Cambridge, UK Systems + Synthetic Biology, CRG, Barcelona http://barcelonacollaboratorium.com http://allox.bio https://www.sanger.ac.uk/programme/generative-and-synthetic-genomics/

Wow. Catalonia has more active clinical trials than the entirety of the UK or Germany. That's wild. From the latest @biocat.cat annual report.
21.02.2026 06:56 — 👍 9 🔁 3 💬 0 📌 0
The allosteric landscape of the Src kinase
by Toni Beltran, @ajfaure.bsky.social , @crg.eu @sangerinstitute.bsky.social @alloxbio.bsky.social
www.science.org/doi/10.1126/...
Looking to start your lab in generative biology / AI?
Come join us at the @sangerinstitute.bsky.social
Sanger is core-funded so you can generate data at scale to train the next generation of models and understanding. Design/Engineering/Chemistry/Proteins/Pathways!
pls RT
tinyurl.com/GenGenFaculty

TF-MAPS: fast high-resolution functional and allosteric mapping of DNA-binding proteins by @XianghuaLi2
Are Transcription Factors really 'undruggable'?
www.biorxiv.org/content/10.1...
Latest pre-print from our lab is out. Great work by @alexbendel.bsky.social and team.
Full deep mutational scan of an entire human protein interaction domain family
2M quantitative measurement of protein-protein interaction -> deep learning model to predict PPI from sequence
More details 👇

Under which circumstances do genomic neural networks learn motifs and their interactions?https://www.biorxiv.org/content/10.1101/2025.07.25.666754v1
01.08.2025 06:18 — 👍 28 🔁 6 💬 0 📌 0
New preprint from the allostery team
www.biorxiv.org/content/10.1...
🎲 Our paper on the genetics, energetics, and allostery in proteins with randomized cores and surfaces is out today @science.org!
🧬 By charting a protein’s sequence universe, we could rationalize which versions were kept through evolution – and why many stable ones were not.

New preprint: Allostery is a widespread cause of loss-of-function variant pathogenicity by the great
@xt117.bsky.social
biorxiv.org/content/10.1...

Only a few days left to apply to become a group leader in our Generative Biology programme at @sangerinstitute.bsky.social
www.sanger.ac.uk

Arriving at the airport to fly to an AI conference and this happens #AIxBio25
23.06.2025 16:58 — 👍 5 🔁 0 💬 0 📌 0
The evolution of allostery in a protein family.
Aina's new preprint containing seven complete comparative allosteric maps biorxiv.org/content/10.1...
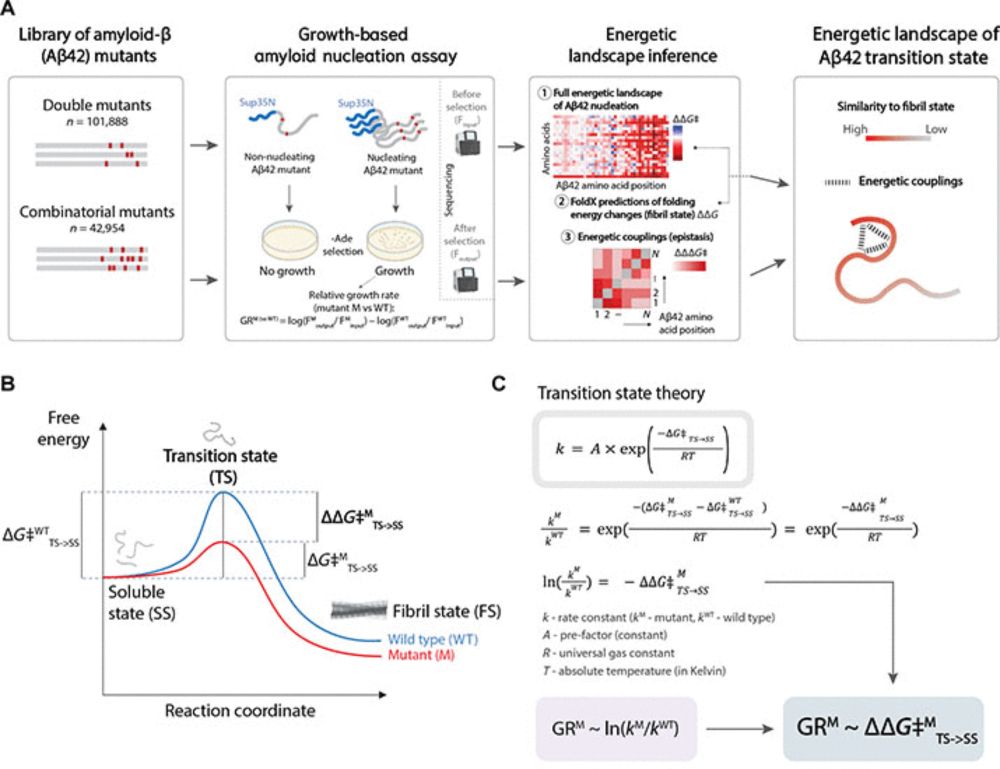
Alzheimer’s disease 🧠starts with a molecular domino effect - but what triggers the first piece to fall?
In our new study we cracked open the black box of early protein aggregation, and the findings could reshape how we fight neurodegeneration. 🧵👇
www.science.org/doi/10.1126/...

1 plant hormone receptor ☘️
3,500 mutants, to single-site saturation 🧬
>45,000 binding and abundance measurements 📶
Very happy to present our latest work – where deep mutational scanning meets the world of small molecules.
www.biorxiv.org/content/10.1...
With @benlehner.bsky.social
[1/7]

Shoutout to the Maxi @maxstammnitz.bsky.social for pioneering dose-response methods and check out his pre-print too: www.biorxiv.org/content/10.1...
01.06.2025 07:48 — 👍 7 🔁 2 💬 0 📌 0
GPCR-MAPS is now live! www.biorxiv.org/content/10.1...
A new platform for high resolution functional mapping of GPCRs with massive mutagenesis. We identify the core activation network, residues involved in biased signaling, and generate >7,000 full dose response curves. With @benlehner.bsky.social
Want to improve your protein or genomic language model’s performance at zero-shot variant effect prediction? We propose a simple adjustment to likelihood-based predicton
26.05.2025 17:33 — 👍 24 🔁 6 💬 0 📌 0
Our "Atlas of Variant Effects 2030 Roadmap" is live: zenodo.org/records/1542...
1/n
Countdown officially started! #VariantEffect25 @varianteffect.bsky.social
09.05.2025 14:37 — 👍 12 🔁 4 💬 0 📌 0

Come and join us! We’re hiring a new Group Leader in Generative Biology at the @sangerinstitute.bsky.social
Building AI models or the data to train them?
Core funding of >$130M a year for a faculty of ~30.
www.nature.com/naturecareer...
acrobat.adobe.com/id/urn:aaid:...
pls RT!
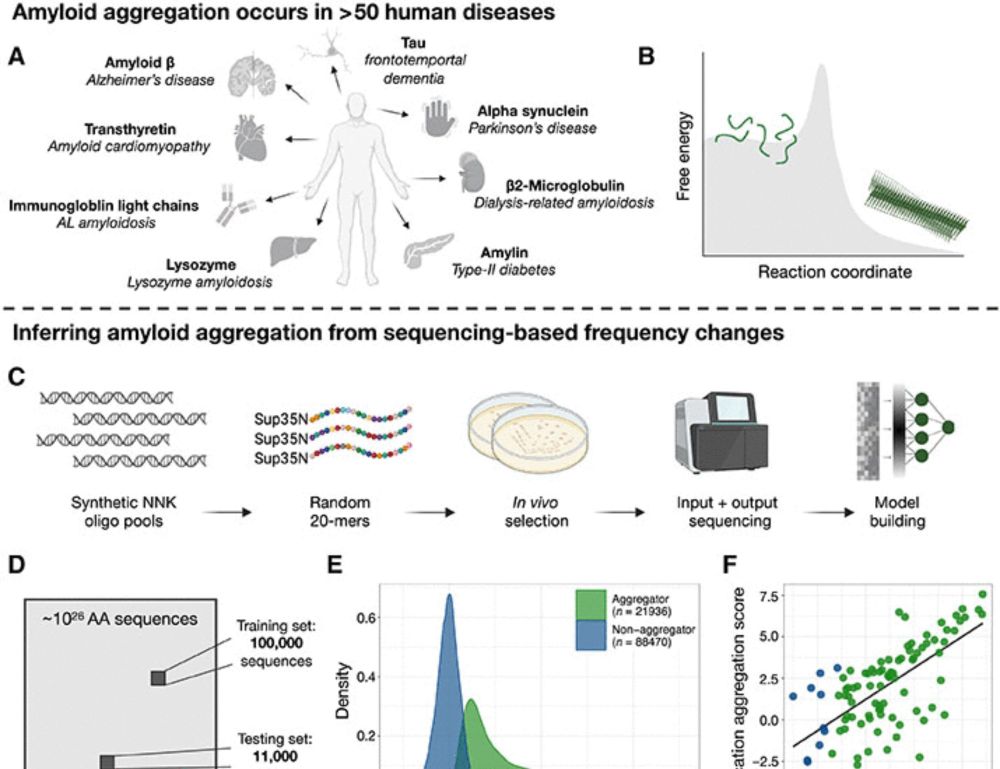
We quantify the aggregation of >100,000 random protein sequences to train CANYA, a convolution-attention hybrid neural network to predict aggregation from sequence. With @bennibolo.bsky.social www.science.org/doi/10.1126/...
04.05.2025 10:40 — 👍 77 🔁 29 💬 1 📌 2
Deadline for abstract submissions and early bird reg for #VariantEffects25 is 1 March 2025.
Organised by @bennibolo.bsky.social @jonnyfrazer.bsky.social @benlehner.bsky.social @muffley.bsky.social @muffley.bsky.social and Mafalda Dias.
More info: events.ibecbarcelona.eu/mutational-s...
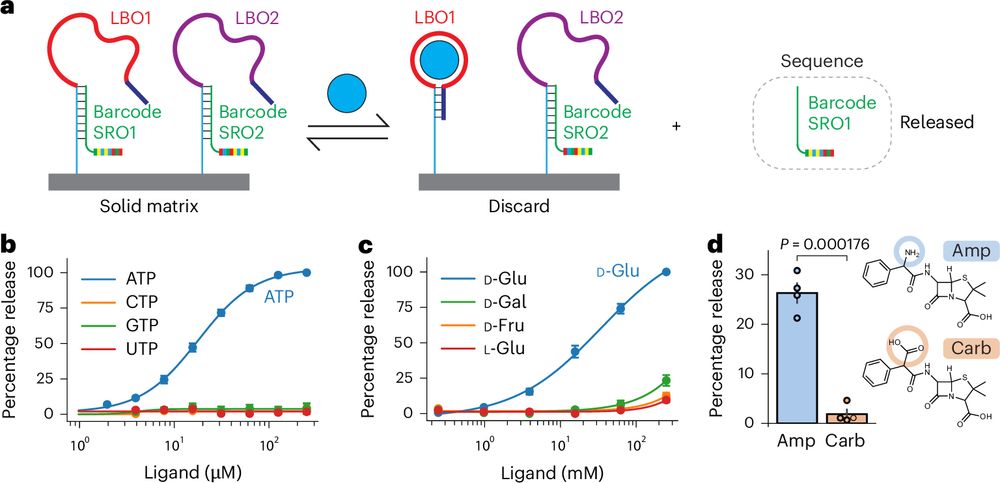
Delighted to share our latest paper describing a method to read the levels of hundreds of metabolites or drugs in parallel using DNA sequencing. This method, which we call ‘smol-seq’ (Small MOLecule sequencing), harnesses the power of DNA sequencing for metabolite detection:
rdcu.be/d8xLv (1/6)
3 great computational positions available at @alloxbio.bsky.social to work with the great @ajfaure.bsky.social @juliadiumenge.bsky.social and team
03.02.2025 12:01 — 👍 1 🔁 3 💬 0 📌 0
At ALLOX @alloxbio.bsky.social in Barcelona we now have 3 computational positions available!
📌Structural Bioinformatics Scientist 👉 tinyurl.com/3rs46j4t
📌Platform Bioinformatician 👉 tinyurl.com/nyvazbke
📌Computational Chemist 👉 tinyurl.com/5n6vpem3
Apply or repost to help us spread the word 🙏
I am incredibly excited to announce that our project with @benlehner.bsky.social, @thomaswilhelm-blue.bsky.social, and the @crg.eu #TBDO has been awarded an ERC Proof of Concept Grant from @ercresearch.bsky.social! Huge thanks to everyone involved for their ongoing support – let's boost the science!
23.01.2025 14:07 — 👍 11 🔁 2 💬 2 📌 0If you’d like to work on such projects, our team is hiring postdocs! We are part of the new Generative and Synthetic Genomics programme with @benlehner.bsky.social and Jussi Taipale, combining data generation at scale with AI to solve biology.Check out sanger.wd103.myworkdayjobs.com/en-US/Wellco...
31.01.2025 13:48 — 👍 8 🔁 5 💬 0 📌 0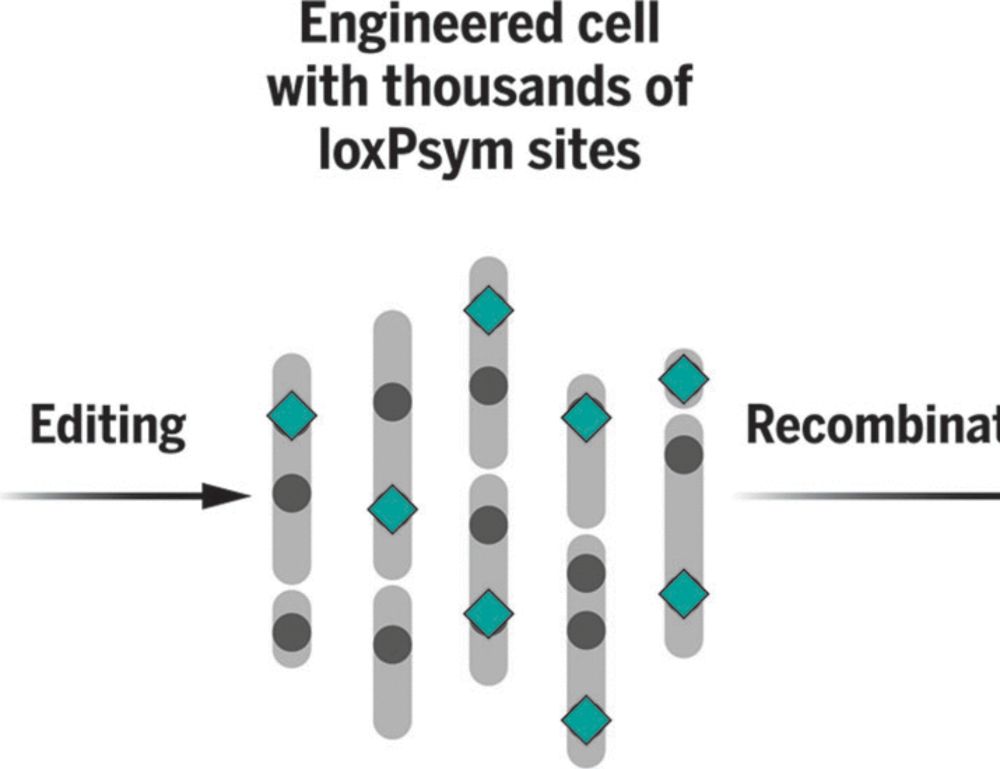
We're delighted to share our work on scrambling the human genome using prime editing, repetitive elements, and recombinases in @science.org , led by @jonaskoeppel.bsky.social , @f-raphael.bsky.social , with @proftomellis.bsky.social and George Church.
www.science.org/doi/10.1126/...
Nice highlighting of @taylor-mighell.bsky.social's GPCR pharmacochaperone preprint by @dereklowe.bsky.social @science.org www.biorxiv.org/content/10.1...
19.01.2025 14:05 — 👍 11 🔁 4 💬 0 📌 0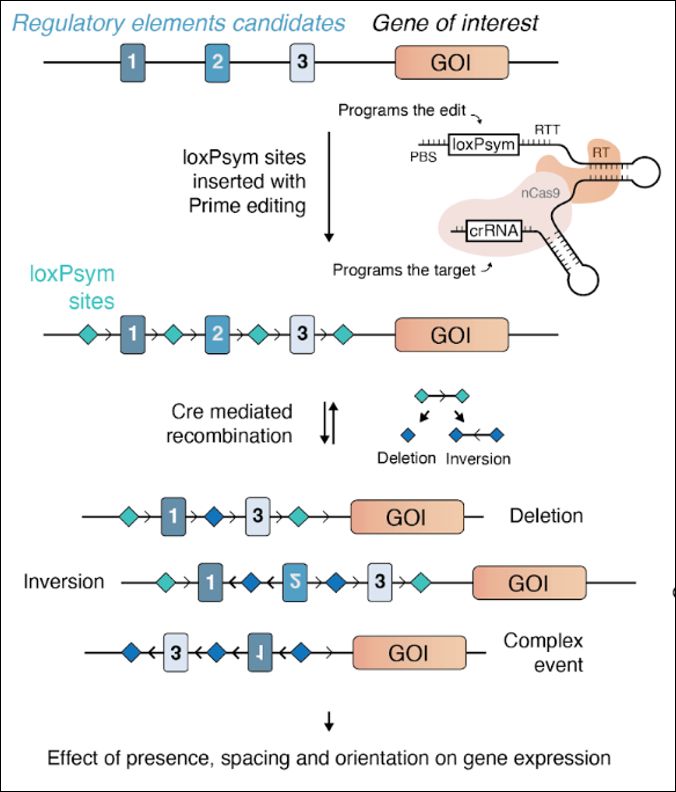
Enhancer scrambling strategy
We are happy to share our enhancer scramble story, a strategy to create hundreds of stochastic deletions, inversions, and duplications within mammalian gene regulatory regions and associate these new architectures with gene expression levels 🧵
www.biorxiv.org/content/10.1...