
NR5A2 overexpression improves SCNT efficiency and enhances ZGA by promoting H3K27ac deposition. Great work!
journals.plos.org/plosbiology/...
@wkobayashi.bsky.social
PI at the University of Dundee / Wellcome CDA holder / Pre-implantation development / Genomics / TFs / Chromatin / Biochemistry / Cryo-EM Postdoc: MPIB, IMBA Ph.D: Waseda Univ. Google Scholar: https://scholar.google.com/citations?hl=ja&user=tw4M2dgAAAAJ

NR5A2 overexpression improves SCNT efficiency and enhances ZGA by promoting H3K27ac deposition. Great work!
journals.plos.org/plosbiology/...
Happy New Year! Check out 👀our new protocol paper on the analysis of transcription factor-nucleosome interactions! Hope you find it useful 🔬🧬Was a pleasure to write up with you @wkobayashi.bsky.social and Kikuë Tachibana!
31.12.2025 09:25 — 👍 8 🔁 5 💬 0 📌 0
Happy to share a protocol for analysis of TF-nucleosome interactions and cryo-EM structure determination in STAR Protocols. I had a great opportunity to describe this protocol with @aliciakmichael.bsky.social. Thank you very much, all!
www.sciencedirect.com/science/arti...

Excited to announce that our peer-reviewed manuscript is online in the Development journal @dev-journal.bsky.social. The manuscript was much improved through the peer-review process, and I am grateful to the reviewers for their valuable feedback.
journals.biologists.com/dev/article/...
Our first cell-free DNA paper is now published! Thank you to all co-authors and collaborators for their contributions.
13.11.2025 06:01 — 👍 3 🔁 1 💬 1 📌 0
I'm looking forward to presenting at the Third Scottish Cryo-EM Symposium tomorrow. A nice opportunity to meet cryo-EM folks in Scotland.
01.09.2025 14:03 — 👍 1 🔁 0 💬 0 📌 0It was extremely challenging to identify transcription factor binding profiles using mouse pre-implantation embryos due to the limited availability of material. I was fortunate to establish an optimised CUT&Tag protocol for this purpose in 2020.
30.05.2025 14:01 — 👍 0 🔁 0 💬 0 📌 0
This method chapter describes the CUT&Tag approach for profiling transcription factors during murine ZGA. I hope it will be helpful for others studying TF binding profiles in similar contexts.
link.springer.com/protocol/10....
Thank you very much!
24.04.2025 10:43 — 👍 1 🔁 0 💬 0 📌 0Thank you, Johanna! See you at the conference!
22.04.2025 17:52 — 👍 0 🔁 0 💬 0 📌 0
Combining expertise with low-input genomics, biochemistry, and cryo-EM analysis, my group cross the scale to understand this fundamental question.
Dundee is a surprisingly sunny spot in Scotland, and I’m looking forward to enjoying life in the UK, surrounded by its beautiful nature!
I’m pleased to announce that I have now officially started my research group at the University of Dundee, UK. My laboratory focus on how transcription factors orchestrate epigenetic reprogramming in mammalian embryos.
22.04.2025 15:09 — 👍 30 🔁 4 💬 3 📌 0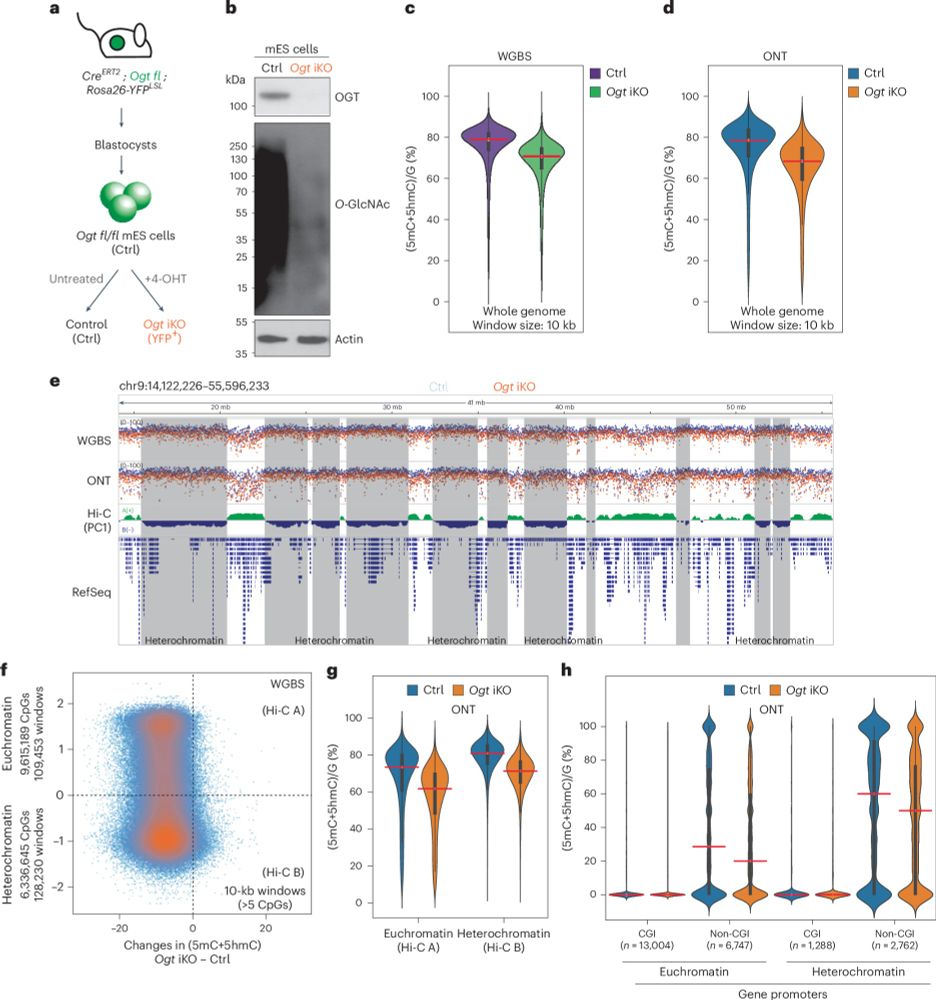
Our study with Anjana Rao's lab was just published at NSMB 🥳 Shows how OGT prevents TET proteins from activating retrotransposons, particularly those in heterochromatin. Paper has 6-base and ONT sequencing data from OGT iKO mESCs, and also interesting OGT inhibitor data 👀.
rdcu.be/efuho
Our second cell-free DNA manuscript is here! In collaboration with Dr. Takada’s group, we explore how abemaciclib treatment response in metastatic breast cancer can be monitored via circulating chromatin fragments: www.researchsquare.com/article/rs-5...
24.03.2025 04:34 — 👍 3 🔁 2 💬 0 📌 0Thank you very much. I moved to UK yesterday. It’s gonna be slow starting.
20.02.2025 12:51 — 👍 0 🔁 0 💬 0 📌 0Thank you, Ana!
20.02.2025 09:41 — 👍 0 🔁 0 💬 0 📌 0Feed-forward loops by NR5A2 ensure robust gene activation during pre-implantation development https://www.biorxiv.org/content/10.1101/2025.02.14.638292v1
20.02.2025 00:30 — 👍 0 🔁 1 💬 0 📌 0
Finally, this work is a great collaboration within the Tachibana lab!
🎉 Huge thanks to:
Chad for all bioinformatic analysis. Eda (a talented student!) helped CUT&Tag and biochemistry. Adarsh helped testing KD. (11/n)
This work represents technical culmination in my postdoc, combining embryology, genomics, biochemistry, and cryo-EM. Huge thanks to Kikuë and the fantastic core facilities at @MPI_Biochem ! (10/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0Taken togther, NR5A2 regulates the expression of KLF5 and GATA6, which in turn function as NR5A2 co-regulators to ensures robust gene activation through feed-forward loops. (9/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0Biochemical data further revealed that NR5A2 co-binds to nucleosomes with KLF5 and GATA6 in vitro, providing strong support that these pioneer factors simultaneously engage with chromatin to drive transcriptional activation in vivo. (8/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0
Surprisingly, Xist was down-regulated upon Nr5a2 KD. We found that NR5A2 regulates Gata1 and Gata6 expression and further binds Xist enhancers with GATA6, indicating that NR5A2 directly or indirectly regulates X-chromosome inactivation. Please also see www.nature.com/articles/s41... (7/n)
20.02.2025 07:43 — 👍 1 🔁 0 💬 1 📌 0Perturbation of both Nr5a2 and Klf5 caused severe developmental defects compared to that of Nr5a2 alone, highlighting synergistic roles of NR5A2 and KLF5. Mechanistically, NR5A2 promotes chromatin accessibility and facilitates H3K27ac deposition in cooperation with KLF5. (6/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0Thus, NR5A2 acts upstream of these factors. Interestingly, these three TFs co-occupancy correlated with the gain of H3K27ac levels at the morula stage. We then addressed how NR5A2 functions together with KLF5 or GATA6. (5/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0We determined NR5A2 chromatin binding profiles from 2-cell to morula stage and identified KLF and GATA families as potential NR5A2 co-regulators. Klf5 and Gata6 were highly expressed in later developmental stages with their expression partially regulated by NR5A2. (4/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0We previously found that NR5A2 targets cell-type specific (putative) enhancers between 2-cell embryos (totipotency) and ESCs (pluripotency). We hypothesized that NR5A2 may cooperate with other TFs to control distinct transcriptional networks during totipotency-to-pluripotency transition. (3/n)
20.02.2025 07:43 — 👍 0 🔁 0 💬 1 📌 0
NR5A2 is a pioneer transcription factor essential for pre-implantation development in mice. We recently reported that NR5A2 facilitates gene expression by unwrapping nucleosomal DNA through minor groove anchor competition. (2/n)
www.nature.com/articles/s41...
🚀Excited to share preprint as my last work in the Tachibana lab! Our study shows how the pioneer factor NR5A2 ensures robust gene activation through feed-forward regulatory loops with lineage-determining factors. A quick summary follows (1/n)
www.biorxiv.org/content/10.1...