
Detecting foldback artifacts in long-reads - BMC Genomics
Long-read sequencing data is useful for detecting large and complex structural variations; however, technical artifacts can lead to false structural variant calls. In our analyses, we became aware of ...
Our paper on foldback artifacts in long-read sequencing is now published in BMC Genomics!
We introduce Breakinator to flag foldback and chimeric artifacts across library types, sequencers, and chemistries.
Paper: link.springer.com/article/10.1...
With Matthew Meyerson and @lh3lh3.bsky.social
24.02.2026 16:02 —
👍 17
🔁 6
💬 1
📌 0

Nallo: a Nextflow pipeline for comprehensive human long-read genome analysis
AbstractMotivation. Long-read sequencing (LRS) is increasingly used for human medical research and clinical diagnostics, due to its capacity to generate co
Say hello to Nallo - our Nextflow pipeline for long-read WGS analysis!👋 It handles both ONT and PacBio data and we’re using this for rare disease and population projects in Sweden. A big team effort by Felix Lenner, Anders Jemt et al.🧬💻 academic.oup.com/bioinformati...
20.02.2026 09:04 —
👍 11
🔁 4
💬 0
📌 0
Just in time for Karneval, we present coelsch ! 🎭🥳
A platform-agnostic framework for single-cell recombination analysis. We study crossover variation in mutants & natural lines, identifying the largest natural inversion in A. thaliana to date!
📄 doi.org/10.64898/202...
🐱 github.com/schneeberger...
05.02.2026 12:21 —
👍 7
🔁 5
💬 1
📌 1
That looks great! To avoid errors due to starting with spotty gene annotations (gene loss vs. not annotated), I would suggest re-annotating every genome used with TOGA2 from the Hiller lab:
github.com/hillerlab/TO...
Many annotations out there are not optimal and TOGA2 has great power and accuracy.
30.01.2026 14:58 —
👍 4
🔁 1
💬 0
📌 0

Vergrößerte Aufnahme der fünf neu entdeckten Krebstiere eingefärbt in Rot, Orange, Gelb, Türkis und Blau vor schwarzem Hintergrund.

Vergrößerte Aufnahme eines der neu entdeckten, blass-weißen Krebstiere, einmal von oben, einmal von der Seite vor schwarzem Hintergrund.
📣 #Forschungsnews: Feine Unterschiede – 5 neue Krebsarten in der Nordsee entdeckt 🌊🦐
In der Deutschen Bucht, einem der weltweit bestuntersuchten Meeresgebiete, hat ein Senckenberg-Forschungsteam 5 neue Arten winziger Krebstiere nachgewiesen. Unbekannte Vielfalt vor der Haustür!
👉 https://sgn.one/1df
28.01.2026 12:01 —
👍 17
🔁 7
💬 0
📌 0
Excited to share our final accepted version of the CiFi method out today: www.nature.com/articles/s41...
08.12.2025 17:55 —
👍 51
🔁 16
💬 1
📌 1

Day 2: bringing the consortium researchers & experts together is on the theme of Functional & comparative genomics
Keynote #StefanMundlos: Functional Genomics of Bat Limb Development
With Genome Chalk team talks #LucasCanesin, #YuryMalovichko, #AlejandroGonzales, #BernhardBein & #MicheleAlbertini
02.12.2025 11:14 —
👍 1
🔁 1
💬 0
📌 0
⚡ We've released a new version of rdeval! v0.0.8 includes full support for @pacbio.bsky.social CiFi read preprocessing, visualization report improvements, and upgrades to performance and parallelization!
🔗 Release notes and source code: github.com/vgl-hub/rdev...
25.11.2025 14:02 —
👍 4
🔁 1
💬 0
📌 0

Happy to present TOGA2, developed by Yury Malovichko @ymalovichko.bsky.social, the faster, memory-efficient & more accurate TOGA1 successor (github.com/hillerlab/TO...). And annotations, orthologs & gene loss/dup data generated with 4 references for 883 placental mammal and with 5 refs for 676 ...
23.11.2025 21:49 —
👍 28
🔁 15
💬 1
📌 0
📢 New #preprint alert 📢
https://www.biorxiv.org/content/10.1101/2025.11.18.689136v1
Several lineages of #birds and #mammals appear to have independently gained or lost the ability to enter #torpor. 🧵1/4
19.11.2025 15:04 —
👍 0
🔁 1
💬 1
📌 0

The poster session was really great and lots of fun @eseb2025.bsky.social! Many interesting discussions and ideas! Thanks to everyone who stopped by at my poster! Looking forward to more interesting talks and posters in the following days!
19.08.2025 20:32 —
👍 8
🔁 1
💬 1
📌 0
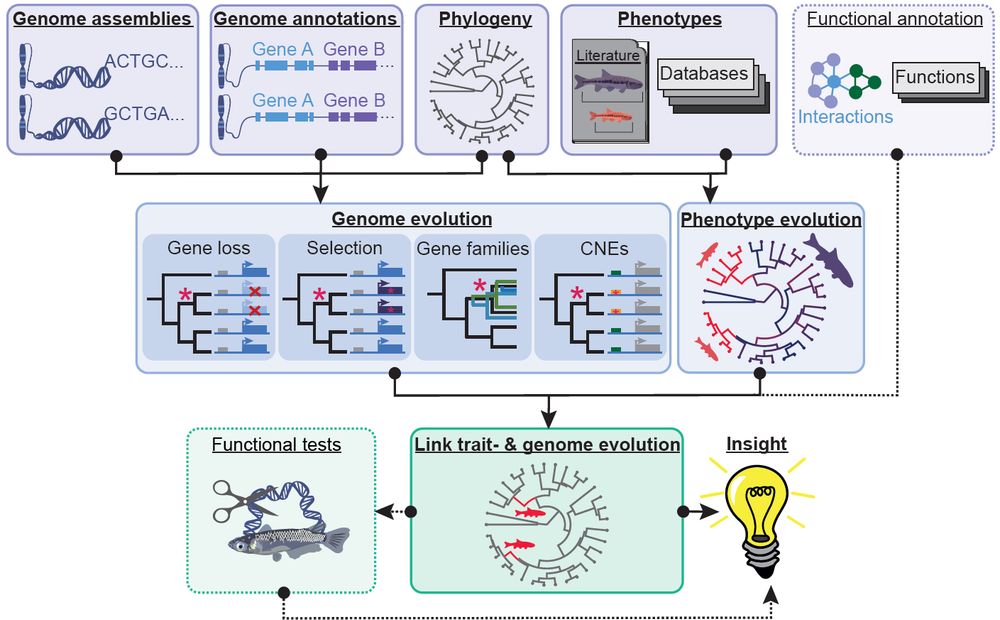
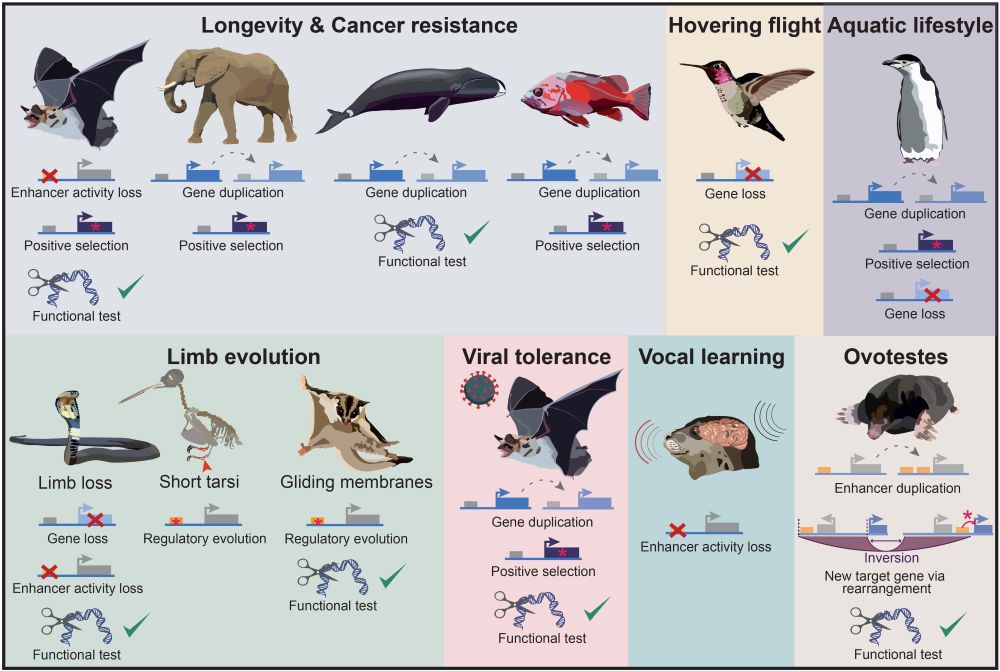
You link #phenotype 🦜 to #genotype 🧬 with #comparative #genomics 💻?
This #review is for you 📜: authors.elsevier.com/sd/article/S...
We review new #methods, remaining #challenges and #future directions and highlight recent key studies.
Thanks @hillermich.bsky.social!
Please share! 🙂
24.07.2025 08:00 —
👍 42
🔁 22
💬 1
📌 1
new from the lab -- we investigate 100% ethanol, DMSO salt solution (DESS), EDTA, RNAlater, and AllProtect as possible approaches for short term room temperature preservation of mosquito HMW DNA, nuclei for Hi-C, and RNA towards high quality reference genome creation. the key findings/tips are ....
07.07.2025 07:20 —
👍 24
🔁 13
💬 3
📌 0

Four new freshwater crab species of the genus Megapleonum Huang, Shih & Ahyong, 2018 (Crustacea, Decapoda, Potamidae) from Guangdong, China
Four new species of the poorly known genus Megapleonum Huang, Shih & Ahyong, 2018, are described from Guangdong Province, China: Megapleonum falx sp. nov. from Huizhou City, M. yangdongense sp. no...
6 years ago, a dream came true when Chao took me along on a field trip to look for freshwater crabs in Guangdong. Now, the new species we found is formally described (𝘔𝘦𝘨𝘢𝘱𝘭𝘦𝘰𝘯𝘶𝘮 𝘺𝘢𝘯𝘨𝘥𝘰𝘯𝘨𝘦𝘯𝘴𝘦 in the article).
Congrats!!
zookeys.pensoft.net/articles.php...
#FreshwaterCrab #Biodiversity #Potamidae
06.07.2025 20:18 —
👍 6
🔁 2
💬 0
📌 0
Exciting news! Good to see that findings from our study on ethanol-preserved specimens are now applied to formalin-fixed cancer samples. Looking forward to more of these interdisciplinary applications regarding amplification-aided sequencing (i. e. scWGA).
#PCRAmplification #LongReadSequencing
20.05.2025 14:37 —
👍 1
🔁 0
💬 0
📌 0

#ERGAReads | Thinking of sequencing a genome in your lab? 🧬 This review offers practical guidance on how to start a #genome project for non-model species - covering DNA extraction, sequencing tech, costs, and more!
🔗 rdcu.be/eiHRG
@sgn.one @leibnizlib.bsky.social #biodiversity #genomics
23.04.2025 08:11 —
👍 17
🔁 10
💬 0
📌 0

Still no reading material for the Easter break🌞🌻🐰? Then I have a suggestion: "Establishing genome sequencing and assembly for non-model and emerging model organisms: a brief guide" (lnkd.in/dsRmSBDE). It was a pleasure to work on this new publication with TilmanSchell & @lpodsiadlowski.bsky.social🙏!
17.04.2025 10:39 —
👍 6
🔁 2
💬 0
📌 0

Evaluating the Benefits and Limits of Multiple Displacement Amplification With Whole‐Genome Oxford Nanopore Sequencing
Multiple displacement amplification (MDA) outperforms conventional PCR in long fragment and whole-genome amplification, making it attractive to couple MDA with long-read sequencing of samples with li...
Research about ultra-low input DNA seq and MDA I recently found:
1. MDA on pg input DNA for ONT rapid seq of pathogens + Tool to detect concatemers
doi.org/10.1111/1755...
2. MDA for metagenomics of low biomass samples
doi.org/10.1093/isme...
Exciting Stuff, small inputs + ampli. going strong!
21.03.2025 14:26 —
👍 1
🔁 0
💬 0
📌 0

Ein Kraken der Gattung Muusoctopus schwimmt am Meeresgrund

Ein Blauer Drache (Glaucus atlanticus), eine leuchtend blaue Meeresnacktschnecke, schwebt in dunklem Wasser. Seine fingerartigen Fortsätze erstrecken sich symmetrisch vom Körper. Darunter ist eine blau-weiße, kreisförmige Kolonie einer Staatsqualle zu sehen.

Eine Hawaiische Schwarzfuß-Napfschnecke (Cellana exarata) guckt mit ausgestreckten Fühlern aus ihrer Muschelschale heraus, die auf sandigem Boden nahe einer dunklen Felswand liegt.
🐙 Jetzt abstimmen: Molluske des Jahres 2025! 🐌
5 beeindruckende Weichtiere stehen im Finale. Dem Gewinner-Weichtier winkt die Entschlüsselung seines kompletten Genoms! 🧬🏆
Erfahrt mehr über die Kandidaten und stimmt bis zum 31.3. für euren Favoriten ab 👉 moty.senckenberg.science
03.03.2025 14:11 —
👍 68
🔁 18
💬 4
📌 4


























