Author Correction: Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes - Nature
Nature - Author Correction: Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes
Today we published a Correction on our 2023 @nature.com paper reporting the heimdallarchaeial ancestry of eukaryotes: www.nature.com/articles/s41...
Corrected paper: www.nature.com/articles/s41...
Importantly, the re-analyses of the corrected dataset are consistent with the original findings.
11.02.2026 21:10 —
👍 52
🔁 15
💬 1
📌 0


Comparative Genomics of Unicellular Eukaryotes (San Feliu, Spain):
Abstract submission is open, with a short deadline! comparativegenomics2026.com
Join us as we explore the most diverse, surprising, and still largely uncharted branches of the eukaryotic tree.
02.02.2026 18:49 —
👍 20
🔁 12
💬 0
📌 1

Microbial eukaryotes keep challenging assumptions about eukaryotic cell and genome biology.
Our upcoming workshop explores the frontier of protist genomics and how it shapes cell biology, ecology and evolution.
Please save the date! Talks, posters and ECR events
Websites and details coming soon
13.12.2025 10:19 —
👍 94
🔁 38
💬 1
📌 0
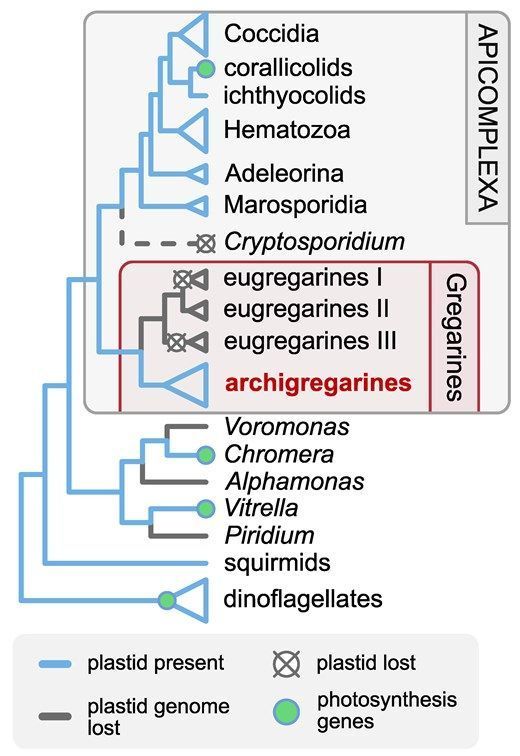
Divergent Plastid Genomes in the Deepest-Branching Apicomplexan Parasites #protists #protistsonsky academic.oup.com/gbe/article/...
11.07.2025 10:01 —
👍 23
🔁 9
💬 0
📌 1
Beautiful work!
26.06.2025 06:06 —
👍 7
🔁 0
💬 1
📌 0
Happy to see our work published and glad to have contributed together with @maxraas.bsky.social ! Looking forward to all the projects that will come out of this work!
23.06.2025 11:18 —
👍 19
🔁 6
💬 0
📌 0
We haven’t. But we have follow up projects that dig more specifically in the archaeal side of things!
23.06.2025 01:01 —
👍 1
🔁 0
💬 1
📌 0
A huge shoutout to @jvhooff.bsky.social who poured her soul into this paper, and our co-authors @eelcotromer.bsky.social and Max Raas!
22.06.2025 00:15 —
👍 5
🔁 0
💬 0
📌 0
Redirecting
If you’re into chromosome biology, genome organization, or evolutionary cell biology, give it a read and share your thoughts!
🔗 doi.org/10.1016/j.celrep.2025.115855
#EvoCellBio #Genomics #SMC #LECA #Chromatin
22.06.2025 00:15 —
👍 7
🔁 0
💬 1
📌 0
Going deeper, duplications of SMC genes happened before eukaryotes: they trace back to the TACK + Asgard archaeal ancestor. This hints at sophisticated chromosome management in the archaeal lineage that gave rise to eukaryotes. #eukaryogenesis #archaea
22.06.2025 00:15 —
👍 3
🔁 0
💬 1
📌 0
Surprise: condensin II was lost independently >30 times across eukaryote evolution—making it one of the most frequently discarded cellular machineries we know. This highlights major shifts in genome organization throughout eukaryotic history.
22.06.2025 00:15 —
👍 8
🔁 0
💬 1
📌 0
SMC complexes (condensin I/II, cohesin, SMC5/6) are the molecular machines that fold chromosomes. By scanning >1 000 genomes we show LECA already housed all four—pointing to a remarkably sophisticated ancestral eukaryote.
22.06.2025 00:15 —
👍 8
🔁 1
💬 1
📌 0

Excited to share our new paper in @cellreports.bsky.social that reshapes our understanding of chromosome organization's deep evolutionary roots! Our work dives into the origins of the machinery that structures our very genomes.
🔗: doi.org/10.1016/j.ce...
#Genomics #Evolution #CellBiology #LECA
22.06.2025 00:15 —
👍 181
🔁 74
💬 4
📌 2
Thank you, Jacob!
19.06.2025 23:00 —
👍 1
🔁 0
💬 0
📌 0
Because it’s done on a computer, some think it doesn’t require expertise and that anyone can do it through button pushing. Just because there are arguments doesn’t mean all arguments have equal value. And in fact, over time, many things in phylogenetics have settled even when they started hot.
19.06.2025 19:45 —
👍 3
🔁 0
💬 1
📌 0
Meaning what? 🫣
19.06.2025 18:07 —
👍 0
🔁 0
💬 1
📌 0
With @deemteam.bsky.social @andrewjroger.bsky.social and more!!
17.06.2025 14:25 —
👍 2
🔁 1
💬 1
📌 0
Why it matters
🎯 Helps to place DPANN firmly in the archaeal tree, tidying up a long-standing evolutionary puzzle
💡 Highlights cross-domain gene transfer as a key driver in archaeal innovation
🌱 Opens doors to explore how free‑living ancestors gave rise to symbiotic lifestyles
17.06.2025 14:25 —
👍 0
🔁 1
💬 1
📌 0
4/ We uncovered ancient horizontal gene transfer events from symbiotic bacteria—namely Patescibacteria and Omnitrophota—that likely empowered DPANN’s shift to their unique episymbiotic lifestyles
17.06.2025 14:25 —
👍 0
🔁 1
💬 1
📌 0
3/ Altiarchaeota emerge as the earliest‑diverging branch in DPANN, suggesting an origin from free‑living ancestors
17.06.2025 14:25 —
👍 0
🔁 1
💬 1
📌 0
2/ Key finding: DPANN forms a monophyletic clade within Euryarchaeota, resolving a long-standing debate on their evolutionary placement
17.06.2025 14:25 —
👍 0
🔁 1
💬 2
📌 0
1/ We used 126 conserved proteins across 11 known DPANN phyla, with careful taxon sampling and sophisticated phylogenomic analysis
17.06.2025 14:25 —
👍 1
🔁 1
💬 1
📌 0
Every night should be duck night Chez Gladines ! 😋
25.05.2025 20:44 —
👍 1
🔁 0
💬 0
📌 0

New vacancy in my team!
PhD student position on microbial genome evolution, focusing on the evolutionary principles underlying bacterial genome architecture.
Please repost and share with talented MSc students in #evobio, bioinformatics or related :)
www.uu.nl/en/organisat...
#MEvoSky #MicroSky
14.05.2025 14:07 —
👍 84
🔁 105
💬 3
📌 3
Fair enough, my initial post was hastily worded.
And yes, when at least one reviewer defeats the point of peer-review by saying teh authors shouldn’t do more work and the debate should happen in public, that’s not helping anyone.
12.05.2025 18:17 —
👍 2
🔁 0
💬 0
📌 0

The marine ciliate Mesodinium rubrum (bottom right) is famed for its ability to steal chloroplasts and nuclei from red cryptophyte algae in the Teleaulax/Gemingera clade. But even these prey specialists can make "mistakes," ingesting blue-green Hemiselmis pacifica (upper left) and transiently retaining their plastids. Here, a M. rubrum cell points its feeding tentacles (labeled with an anti-centrin antibody, yellow) at an H. pacifica cell. The cryptophyte's flagella and ciliate's cilia (labeled with an anti-tubulin antibody, purple) and nuclei (labeled with DAPI, blue) are visible in this expansion micrograph taken with a Nikon spinning disk confocal microscope. Numerous stolen prey nuclei are visible inside of the M. rubrum cell. See Moeller et al. https://doi.org/10.1111/jeu.13066
Check out exciting protist research in JEM! Holly Moeller et al. explore the feeding habits of Mesodinium rubrum and provide a tool for studying its cell biology and photophysiology in their open access paper in our March/April issue!
doi.org/10.1111/jeu....
#protistsonsky
08.05.2025 12:00 —
👍 17
🔁 5
💬 0
📌 2
(Unlike you, the authors here do imply that the phylogenetic conclusions were driven by this contamination)
09.05.2025 17:56 —
👍 0
🔁 0
💬 0
📌 0
To be clear, I’m not “defending” these MAGs, and take your point regarding Fig 2c.d I don’t think figure 2a b is proving anything though (see above post). Curious to hear your take on 2a and b if any?
09.05.2025 17:55 —
👍 1
🔁 0
💬 2
📌 0








