
I made a map of 3.4 million Bluesky users - see if you can find yourself!
bluesky-map.theo.io
I've seen some similar projects, but IMO this seems to better capture some of the fine-grained detail

I made a map of 3.4 million Bluesky users - see if you can find yourself!
bluesky-map.theo.io
I've seen some similar projects, but IMO this seems to better capture some of the fine-grained detail
Our study of the evolution of the ParB NTP binding domain across the Tree of Life is now published! An awesome collaboration with @tunglejic.bsky.social led by @jovanakaljevic.bsky.social at the John Ines Center. Thanks to editors and reviewers. www.pnas.org/doi/10.1073/...
05.12.2025 09:05 — 👍 18 🔁 4 💬 0 📌 1
Paper alert!
We have created a bacterium that eats plastic! We named it PETBuster! Great work by PhD student Dekel Freund @dekel-freund.bsky.social.
www.biorxiv.org/content/10.1...
Early Microbial Evolution
"The origin of life on Earth remains one of the greatest and most pervasive mysteries in science. We know the story in broad strokes: Around 4 billion years ago, simple chemical compounds gave rise to living cells, which later formed..."
🦠
asm.org/articles/202...



Alternate between predicting structure with AF3 and designing sequence with ProteinMPNN to generate proteins, including protein, small molecule and nucleic acid binders
@yehlincho.bsky.social @sokrypton.org
www.biorxiv.org/content/10.1...
Thrilled to see this work in its preprint form! A great collaboration with Jovana and Tung to explore the evolution and properties of the surprisingly versatile nucleotide binding domain of ParB.
11.10.2025 13:02 — 👍 19 🔁 2 💬 1 📌 0
New preprint!
Ever wondered why only a fraction of genomes encode CRISPR immunity? 🧬 🦠
Turns out CRISPR is rarely beneficial against virulent phages, being most beneficial against those for which resistance mutations are rare!
An epic effort by Rosanna Wright
www.biorxiv.org/content/10.1...

#NewResearch
Structure of a functional archaellum in bacteria of the Chloroflexota phylum
#MicroSky
@archaellum.bsky.social @sshamphavi.bsky.social @loumollat.bsky.social @mariejoest.bsky.social @sgribaldo.bsky.social
www.nature.com/articles/s41...
hey bluesky 👋 visa hurdles mean I’m looking for opportunities outside the US. I’m a computational biologist (bacterial + phage genomics, postdoc in Koonin’s group @ NIH). I am interested in teaming up on funding apps. reach out if this resonates!
15.09.2025 17:26 — 👍 69 🔁 89 💬 1 📌 3
Sometimes you meet absolutely incredible bioinfo-magicians.
It was a huge privilege when @shenwei356.bsky.social
joined our group for a year on an @embl.org sabbatical.
While here, he developed a new way of aligning to
millions of bacteria, called LexicMap 1/n
www.nature.com/articles/s41...

Please re-post:
Interested in chromatin and its evolution? Good news! There's still time to join us in beautiful Catalonia (9-12 Dec) to discuss eukaryotic, bacterial, archaeal, and viral chromatin and how it all hangs together meetings.embo.org/event/24-evo...
Abstract deadline: 30 September

Preprint: De-novo design of proteins that inhibit bacterial defenses
Our approach allows silencing defense systems of choice. We show how this approach enables programming of “untransformable” bacteria, and how it can enhance phage therapy applications
Congrats Jeremy Garb!
tinyurl.com/Syttt
🧵

Please re-post: If you know (or are!) somebody who might fancy doing a PhD (Oct 2026 start) in my group @oxfordbiochemistry.bsky.social, working on chromatin evolution in prokaryotes (or other things we're interested in), please have a look at www.bioch.ox.ac.uk/supervisors-...
01.09.2025 09:27 — 👍 42 🔁 50 💬 0 📌 1
Do plasmids evolve faster 🐇, slower 🐢, or just like chromosomes 🧬?
In our new paper, we tackled this question using theory, simulations, bioinformatics, and experiments!
👇 Check out all the details in Paula’s thread!
Hint: 🐇 (most of the time)
I was #TWiMAdjacent about the much missed Elio Schaechter. I tried to discuss an article later, but I was stressed for time. The first part about Elio is so worth your time, as is the paper I discussed, about how archaea make enzymes that break down the bacterial cell wall!
asm.org/podcasts/twi...

Awesome to see the transparent, systematic, and evidence-based approach behind the WHO Bacterial Priority Pathogens List 2024 out in the Lancet Infectious Diseases www.thelancet.com/journals/lan...
27.08.2025 16:13 — 👍 45 🔁 16 💬 1 📌 1
I cannot fully put into words what publishing this Review has meant to me, so I leave you with how we closed the paper.
"The humble bacterium is still a relevant tool for the study of the underlying mechanisms that are conserved throughout life."
🧪🧫🧬📚
doi.org/10.1093/gene...
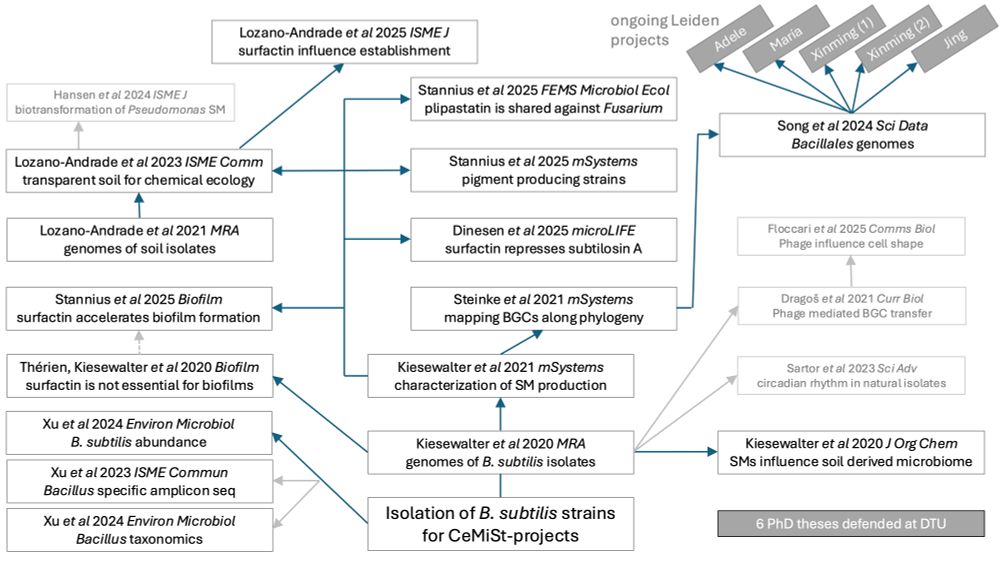

When we started sampling soil to isolate Bacilli and other bacterial species within @cemist.bsky.social, I did not expect that so many direct and indirect publications and numerous chapters in 6 PhD theses in my group will originate from those efforts
🧵 [1/n]

"We show that unrelated proteins have a universal tendency towards convergent evolution of secondary and tertiary motifs, causing an excess of high-scoring FP alignment... previous methods routinely overestimate significance by up to six orders of magnitude."
www.biorxiv.org/content/10.1...

🚨New paper 🚨
Can protein language models help us fight viral outbreaks? Not yet. Here’s why 🧵👇
1/12
📣 ABSTRACT DEADLINE EXTENDED UNTIL 18TH AUGUST!
Register here: register.oxfordabstracts.com/event/75604?...
Abstract submission: app.oxfordabstracts.com/stages/79029...
Not something we tested explicitly but the catalytic domains involved have not been shown to cleave pseudomurein. Also, very few pseudomurein producers encode PGHs, ruling out auto-lysis as the main function.
15.08.2025 12:37 — 👍 2 🔁 0 💬 1 📌 0
Left: Results of spotting various supernatants onto lawns of Halalkalibacterium halodurans, Phycicoccus endophyticus, and Virgibacillus salexigens. a,b: top fraction (>3 kDa) of supernatant from Haloferax volcanii expressing an intact (a) or catalytic mutant (b) version of Woldo; c,d: bottom fraction (<3 kDa) of supernatant from H. volcanii expressing an intact (c) or catalytic mutant (d) version of Woldo; e: unfiltered supernatant from Halogranum salarium B-1; f,g: top fraction (>3 kDa) of supernatant from H. volcanii expressing (f) Danwoldo or (g) and empty plasmid control; h,i: bottom fraction (<3 kDa) of supernatant from H. volcanii expressing (h) Danwoldo or (i) and empty plasmid control. Top right: Structural overlay of zoocin A from Streptococcus zooepidemicus (UniProt ID: O54308) and Woldo, a candidate PGH from Halogranum salarium B-1 (UniProt ID: J2ZZK6), highlighting homologous M23 domains but divergent cell wall binding domains. TRD: target recognition domain. PG: peptidoglycan. Bottom right: Images of bacterial cells following exposure to H. volcanii supernatants expressing Woldo, Danwoldo or the control (as above). Bacteria are stained using LIVE/DEAD BacLight bacterial viability staining kit with ingress of red dye into bacterial cells indicative of cells with terminally compromised cell wall integrity.
Archaea-on-bacteria action! @romainstrock.bsky.social @tobiaswarnecke.bsky.social &co show that many #archaea encode #peptidoglycan hydrolases, which specifically target #bacterial cell walls, experimentally confirming the killing capacity of 2 of these enzymes @plosbiology.org 🧪 plos.io/4lrJBBa
15.08.2025 08:13 — 👍 16 🔁 8 💬 1 📌 1PGHs are unlikely to be the sole kind of weapons deployed by archaea to kill bacteria. We close by discussing a road map to uncover more of the archaeal arsenal deployed against bacteria. Have a read for more details! 6/6
15.08.2025 11:19 — 👍 0 🔁 0 💬 0 📌 0
Plate with bacterium Halalkalibacterium halodurans growing on top. Four inhibition zones are visible, caused by peptidoglycan hydrolase proteins secreted by archaeon Halogranum salarium B-1.
We experimentally show that two PGHs encoded by salt-loving archaeon Halogranum salarium B-1 do kill one of the predicted targets, salt-tolerant bacterium Halalkalibacterium halodurans. 5/6
15.08.2025 11:19 — 👍 0 🔁 0 💬 1 📌 0
Pipeline to predict the putative bacterial targets of archaeal epptidoglycan hydrolases.
We developed a structural homology-based pipeline to infer the putative bacterial targets of these proteins and found that most seem to target monoderm bacteria living in the same environment as the producer. 4/6
15.08.2025 11:19 — 👍 1 🔁 0 💬 1 📌 0
Heatmap of the presence of peptidoglycan hydrolases in archaea.
Here, we show that about 5% of archaea encode peptidoglycan hydrolases (PGHs) in their genomes, though most archaea do not use peptidoglycan. They are enriched on plasmids and display a high level of structural conservation with their bacterial counterpart, hinting at their use as weapons. 3/6
15.08.2025 11:19 — 👍 0 🔁 0 💬 1 📌 0Archaea and bacteria live in the same environments across the biosphere, yet only a handful of studies describe antagonistic interactions between the two domains of life (Atanasova et al., 2013 is particularly relevant). 2/6
15.08.2025 11:19 — 👍 1 🔁 0 💬 1 📌 0
This first-author publication is my… first! Archaea kill bacteria by targeting their Achilles’ heel: peptidoglycan. Big shoutout to @ahocher.bsky.social, @valeriesoo.bsky.social, Pauline Misson, @tobiaswarnecke.bsky.social and MRC LMS Proteomics. A thread 🔽
journals.plos.org/plosbiology/...
@romainstrock.bsky.social work is out!
Why should you care? Because it helps shift our view of archaea — from rare extremophiles to not-so-passive bystanders. Some archaea can also fend off bacteria.
Glad to be part of this cool project from @tobiaswarnecke.bsky.social. Opens lots of Questions.