
Our great collaboration with the lab of Wei Yang and Kaoru Sugasawa is now online in Nature. We’re glad we could contribute some cell biology to complement this beautiful structural and biochemical work.
www.nature.com/articles/s41...
@jervdberg.bsky.social
post-doc with Alexander van Oudenaarden @hubrecht | PhD with René Medema @NKI | Canman lab @ Columbia NYC | single cell sequencing | scEdU-seq| DNA replication | CRISPR | Genomic instability & Cell Fate | he/him

Our great collaboration with the lab of Wei Yang and Kaoru Sugasawa is now online in Nature. We’re glad we could contribute some cell biology to complement this beautiful structural and biochemical work.
www.nature.com/articles/s41...

Excited to share our new Nature study! 🧬
We (sidrituruci.bsky.social et al) discovered that CFAP20 helps clear stalled RNAPII, preventing harmful clashes with DNA replication machinery. This protects cells from R-loops and genome instability.
Full paper: www.nature.com/articles/s41...

🧬 Preprint alert!
"Distinct ATRX functions cooperate with 9-1-1 and CST complexes to safeguard replication and telomere integrity."
Read the manuscript here 👉 www.biorxiv.org/content/10.1...

New method paper:
Bennie Lemmens et al @scilifelab.se describe 3D-SPARK, a super-resolution microscopy method that maps changes in DNA replication nanostructures in response to genetic perturbations or drugs, linking spatial cell biology with DNA synthesis dynamics
www.embopress.org/doi/full/10....

Are you a postdoc looking for a new challenge and do you match the profile? Apply to join us! Especially, if you have experience with addressing the mechanisms underlying telomere maintenance, DNA replication or DNA repair. www.werkenbijavl.nl/vacatures/po...
06.10.2025 13:25 — 👍 7 🔁 6 💬 0 📌 1
Histone acetylation - not always about transcriptional activation: H4K16ac safeguards the genome replication program by repressing premature replication of heterochromatin. Well done Marta Milan! tinyurl.com/yuy9m8t3
30.09.2025 12:14 — 👍 31 🔁 13 💬 2 📌 0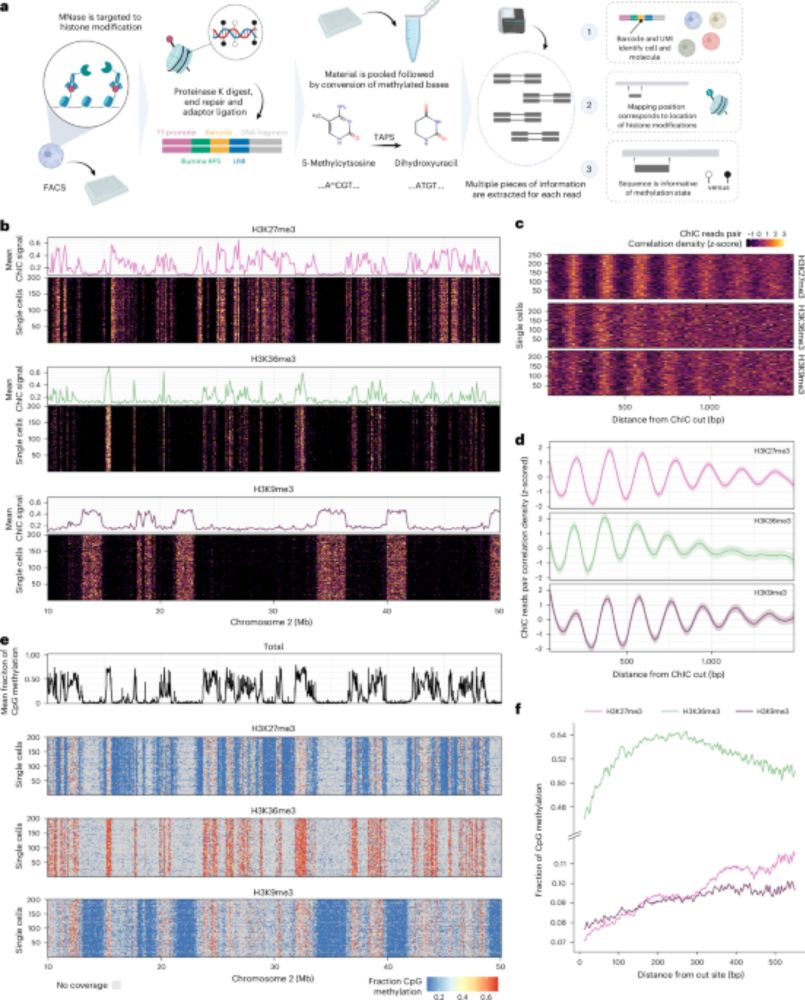
scEpi2-seq: a method for simultaneous single-cell profiling of DNA methylation and histone modifications.
@jervdberg.bsky.social @hubrechtinstitute.bsky.social
www.nature.com/articles/s41...




🧬 Hubrecht researchers have developed a method that can detect DNA methylation and histone modifications simultaneously in single cells. They can now for the first time study how their interaction influences gene activity. https://www.nature.com/articles/s41592-025-02847-4
@jervdberg.bsky.social
big shoutout to Christoph Geisenberger, Alexander van Oudenaarden and all co-authors!
link to paper 👉 www.nature.com/articles/s41...
New method 🚀: scEpi2-seq lets us read DNA methylation+histone marks in the same single cell. Found that methylation “memory” after replication depends on chromatin+histones, and TF promoters repressed by H3K27me3 carry distinct methylation. So stoked to share this as first+co-corresponding author 🎉
25.09.2025 09:02 — 👍 18 🔁 5 💬 2 📌 0The latest work from ours and @vram142.bsky.social lab is out! True teamwork to visualize nascent chromatin with strand resolution, using a fully reconstituted system. Very proud of superstar-PhD student Bruna, and @palindromephd.bsky.social. Learning so much from Vijay’s amazing technologies! RT
21.09.2025 07:12 — 👍 36 🔁 12 💬 2 📌 0Teamwork at its best!! Thank you Vijay, for sharing your technologies and computational talent, for the fun discussions and work together, at random hours of the day or night 😃
21.09.2025 07:25 — 👍 8 🔁 2 💬 0 📌 0such a tour de force! great stuff, drop eveything and first read this, you wont be disappointed! congrats to all authors @vram142.bsky.social @fmattiroli.bsky.social
22.09.2025 09:05 — 👍 8 🔁 0 💬 1 📌 0
Some (+)ve news to lighten another heavy weekend: our latest preprint (c/o Mattiroli + Ramani labs) is up!
www.biorxiv.org/content/10.1...
A tour-de-force by 1st authors Bruna Eckhardt & @palindromephd.bsky.social, focusing on chromatin replication. RTs welcome; tweetorial in 3,2...(1/n)

Image of Niels Mejlgaar (CEO of DNRF), Anja Groth (EpiC director) & Jesper Fisker (CEO of DCS).
We have lift off!! 🚀 Yesterday was the official opening of EpiC! The DNRF Center for Epigenetic Cell Memory at the Danish Cancer Institute!
Our EpiC team with director Anja Groth @groth-anja.bsky.social, were joined by Jesper Fisker, CEO @cancer.dk & Niels Mejlgaar, CEO @dg.dk to celebrate 🎉

Image of Toronto
JOB ALERT 🚨 We are hiring TWO principal investigators in cell, molecular, systems, or chemical biology in Toronto, Canada at @sinaihealth.bsky.social. We provide a generous startup, fully funded salary and academic appointment at U of Toronto.
www.nature.com/naturecareer...
Please repost!

Aspiring PhD students, this is your chance to join us at the NKI to work on genomic instability and telomeres!: www.werkenbijavl.nl/vacatures/ph...
15.08.2025 09:42 — 👍 3 🔁 6 💬 0 📌 0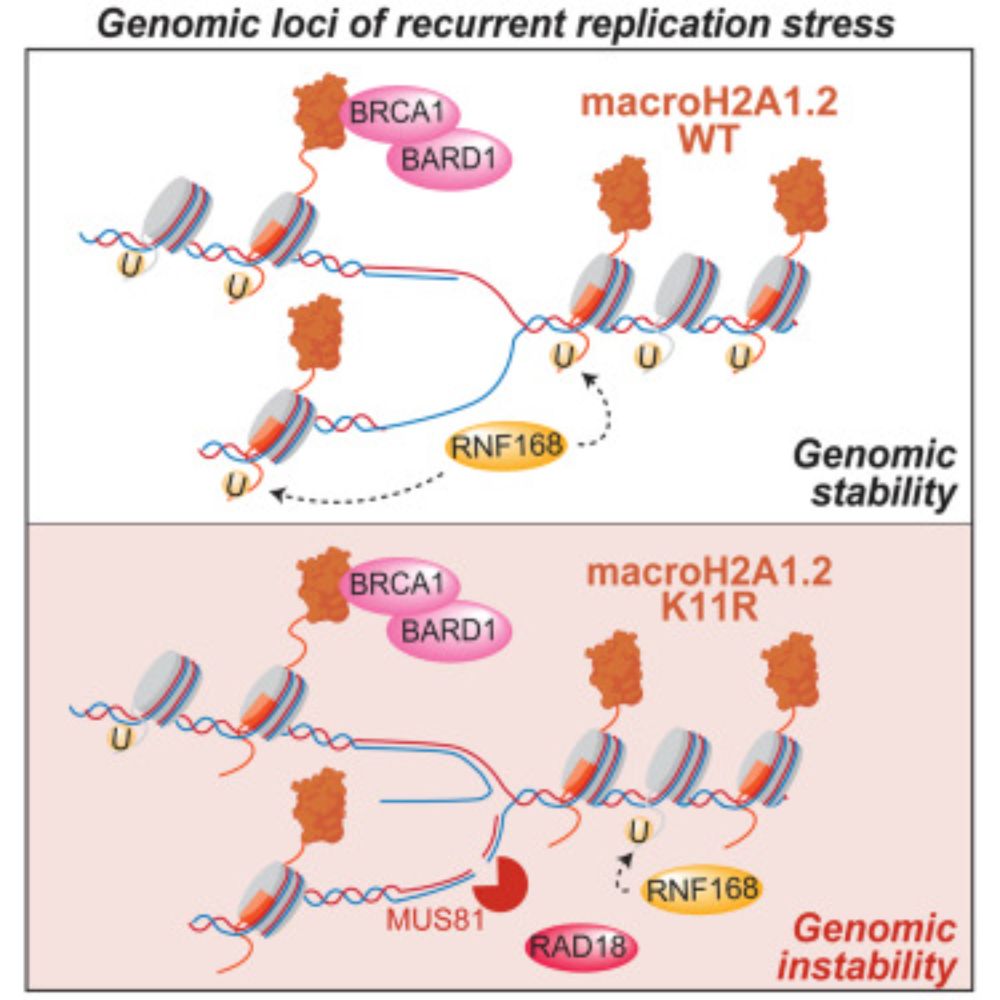
I am very excited to share the newest paper of the lab, a huge amount of work led by our talented PhD student Maxime Galloy, in close collaboration with Andréanne Blondeau, our dedicated research assistant for 10 years! 🙌
www.cell.com/molecular-ce...
I am thrilled to share with you my first co-corresponding author Nature paper with Jos Jonkers. This project was led by the extraordinary @sarahmoser.bsky.social . Congratulations, and thank you to all the authors, @nkinl.bsky.social , and @oncodeinstitute.bsky.social for their support.
13.08.2025 15:57 — 👍 30 🔁 5 💬 4 📌 0Congrats on this great paper!
13.08.2025 17:10 — 👍 2 🔁 0 💬 0 📌 0
Thrilled to share our latest work, just published in @nature.com ⬇
www.nature.com/articles/s41...
We discovered that PARP inhibitors 💊 trigger histone eviction from the chromatin and this creates a hidden vulnerability in PARPi resistant tumors.
🧵 (1/8)
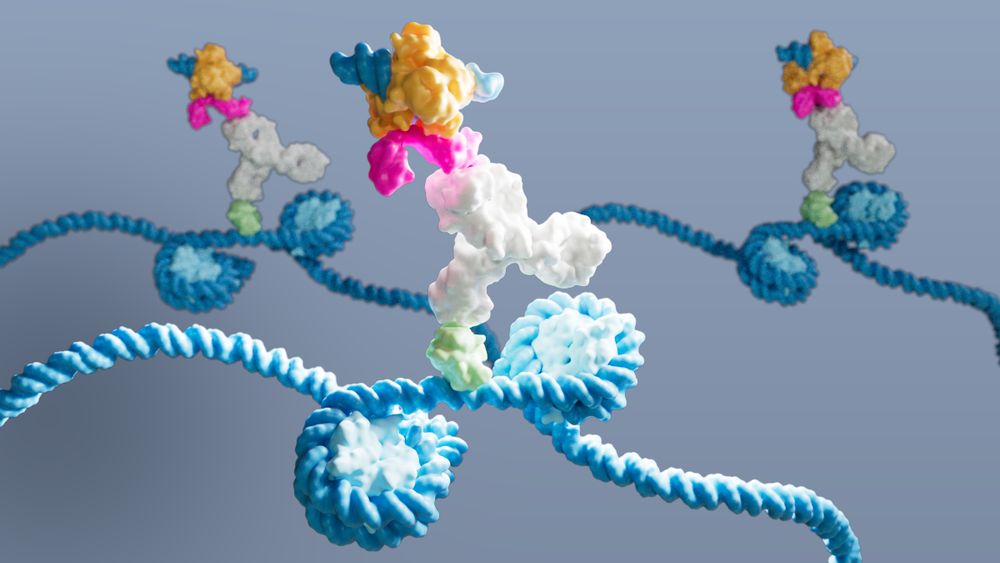
Image credit: @gloglita.bsky.social @lifescienceeditors.bsky.social captured DynaTag in action: a pA-Tn5 probe (multicoloured) binds an antibody (white), which binds p53 DNA-binding domain (green) on DNA (blue) within 2 nucleosomes
🧪Move over CUT&Tag, there’s a new #TranscriptionFactor mapping method in town.
Our newly developed DynaTag is faster, cleaner, more sensitive than #ChIPseq, #CUT&RUN and #CUT&Tag.
🔗 Our @natcomms.nature.com paper: www.nature.com/articles/s41...
🧵Let’s break down what makes DynaTag so powerful (1/7)

Less than two weeks left to apply!
Postdoc positions in my lab to study
1. initiation of DNA replication.
2. chromatin replication/epigenetic inheritance.
Great for biochemists, biophysicists & structural biologists.
Deadline 3 August 2025.
crick.wd3.myworkdayjobs.com/External/job...

Very proud of this international collaborative team effort published in EJHG defining ERCC1-hepatorenal syndrome as a severe multisystem DNA repair disorder with high mortality and pediatric liver cancer risk. Thanks to all families and collaborators.
www.nature.com/articles/s41...
Check out this new review (with animations, natch) of mechanisms of licensing origins of DNA replication - a wonderful (and continuing!) collaboration with Bruce Stillman @cshlnews.bsky.social and John Diffley @crick.ac.uk! www.nature.com/articles/s41...
02.07.2025 17:01 — 👍 81 🔁 44 💬 6 📌 2
Cytarabine has been the mainstay for AML treatment for over 50 years. This chemotherapy can lead to problems with movement and balance. Here we explain this neurotoxic side-effect. www.nature.com/articles/s41... Work led by Jia-Cheng Liu, Donpeng Wang and Elsa Callen and terrific collaborators!
25.06.2025 15:59 — 👍 24 🔁 10 💬 0 📌 0
We’re #hiring, please share! A fully funded #postdoc position is available in my lab to work on DNA damage response mechanisms.
Please get in touch with your CV if you are interested; closing date: 27th June 2025.
More details here: candidate.hr-manager.net/ApplicationI...
Big shoutout to this amazing work from our colleagues in the Knipscheer lab. Congratulations on discovering and dissecting yet another new molecular mechanism that keeps our genome safe! 👏🍾
13.06.2025 09:42 — 👍 10 🔁 2 💬 0 📌 0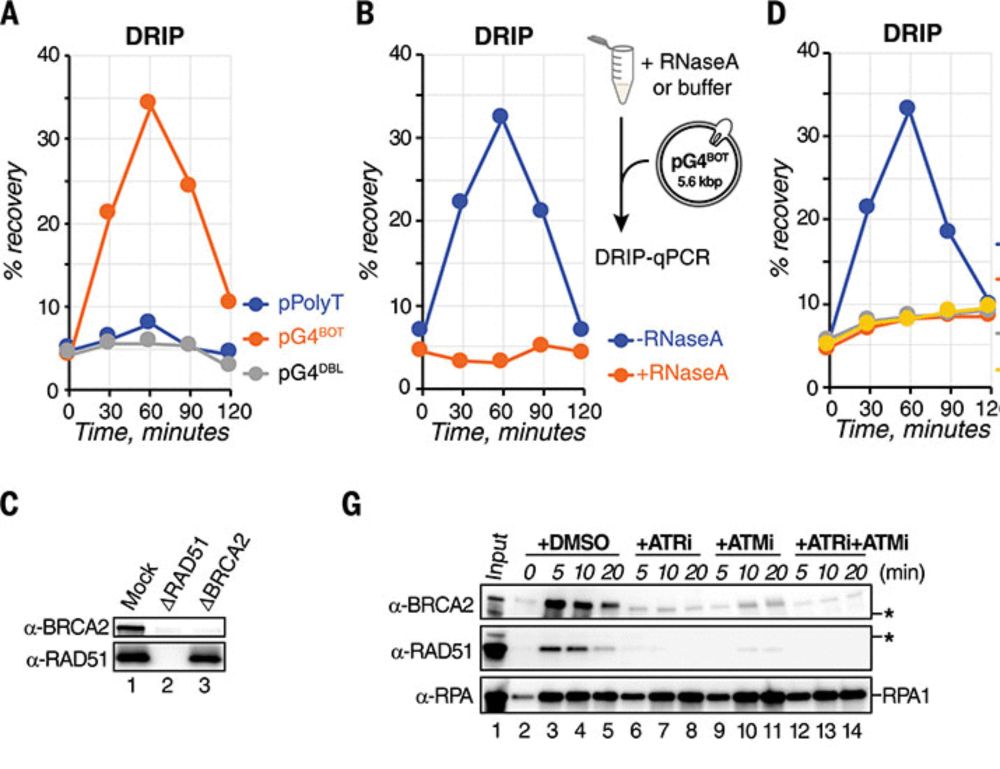
Excited our paper is out! G-quadruplex (G4) unwinding via an intricate G-loop assembly and disassembly mechanism maintains genome integrity. Pioneered by Koichi Sato with great collaborators Jing Lyu and @simonelsasser.bsky.social Check it out: www.science.org/doi/10.1126/science.adr0493 🧵(1/4)
13.06.2025 09:33 — 👍 77 🔁 22 💬 6 📌 3We found G4s are recognized as DNA lesions leading to homology-directed invasion of RNA opposite the G4 to form a G-loop structure. This G-loop initiates G4 unwinding by DHX36 and FANCJ. Subsequent nucleolytic incision followed by renewal of the RNA-containing strand drive G-loop disassembly (2/4)
13.06.2025 09:33 — 👍 1 🔁 1 💬 1 📌 0