The commit benefited tremendously by minimap2’s practical algorithmic solutions for chaining and rescue -- big credits to @lh3lh3.bsky.social and his minimap2. Also to my students Nicolas Buchin and Ivan Tolstoganov, as well as Marcel Martin 4/4
29.12.2025 05:49 —
👍 0
🔁 0
💬 1
📌 0
Strobealign can now map long reads. Still POC, i.e., PAF only -- no supplementary chains or piecewise extension (yet). 3/4
29.12.2025 05:49 —
👍 2
🔁 1
💬 1
📌 0
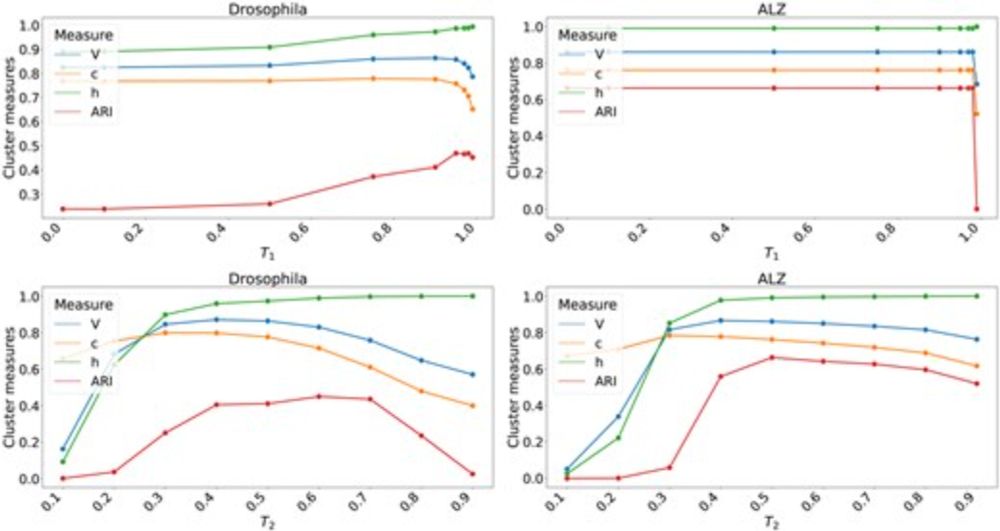
Accuracy upgrades: chaining instead of NAMs, smarter local rescue of repetitive hits (minimap2-style), and improved multi-context seeding 2/4
29.12.2025 05:49 —
👍 0
🔁 0
💬 1
📌 0
Thank you folks for your feedback on our survey about Hash functions in genomic sequence analysis. We've updated the paper and you can see the new version here: tinyurl.com/4kk9ccmt.
25.09.2025 13:21 —
👍 11
🔁 6
💬 0
📌 1
Preprint out for myloasm, our new nanopore / HiFi metagenome assembler!
Nanopore's getting accurate, but
1. Can this lead to better metagenome assemblies?
2. How, algorithmically, to leverage them?
with co-author Max Marin @mgmarin.bsky.social, supervised by Heng Li @lh3lh3.bsky.social
1 / N
07.09.2025 23:34 —
👍 114
🔁 80
💬 5
📌 5


Congratulations to Rayan Chiki, (Institut Pasteur) head of the “Sequence Bioinformatics” unit, for securing the ERC Proof of Concept 2025 for his project ENZYMINER! 👏
@rayan.chiki.bsky.social
#Bioinformatics
24.07.2025 15:10 —
👍 60
🔁 13
💬 4
📌 2
Incredible! 👏
25.07.2025 22:00 —
👍 1
🔁 0
💬 0
📌 0
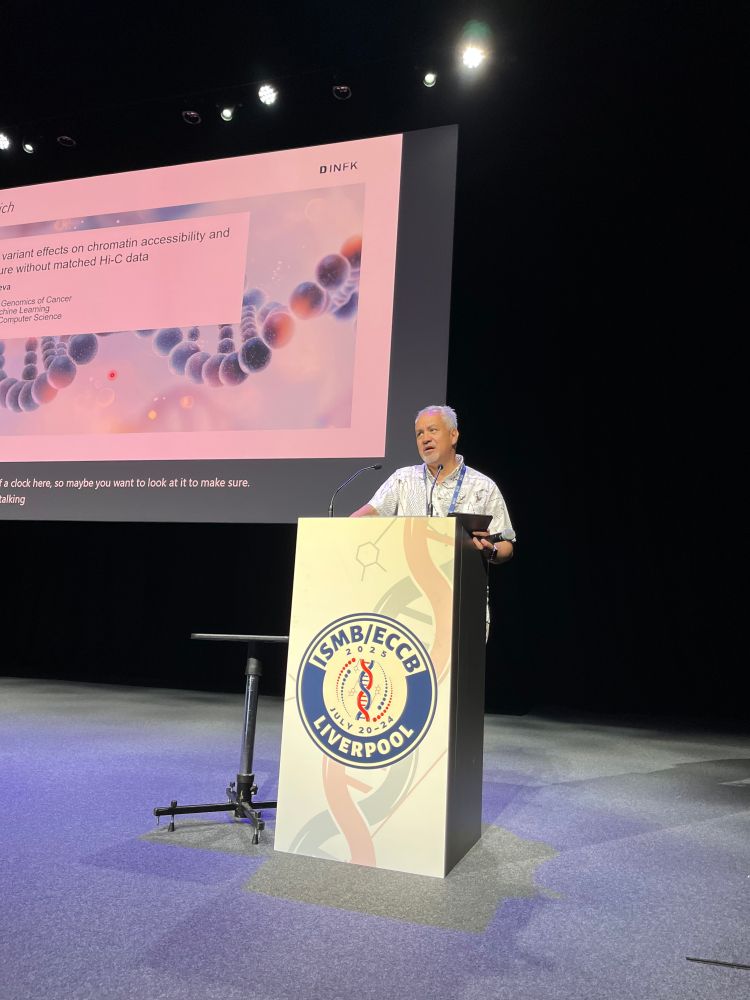
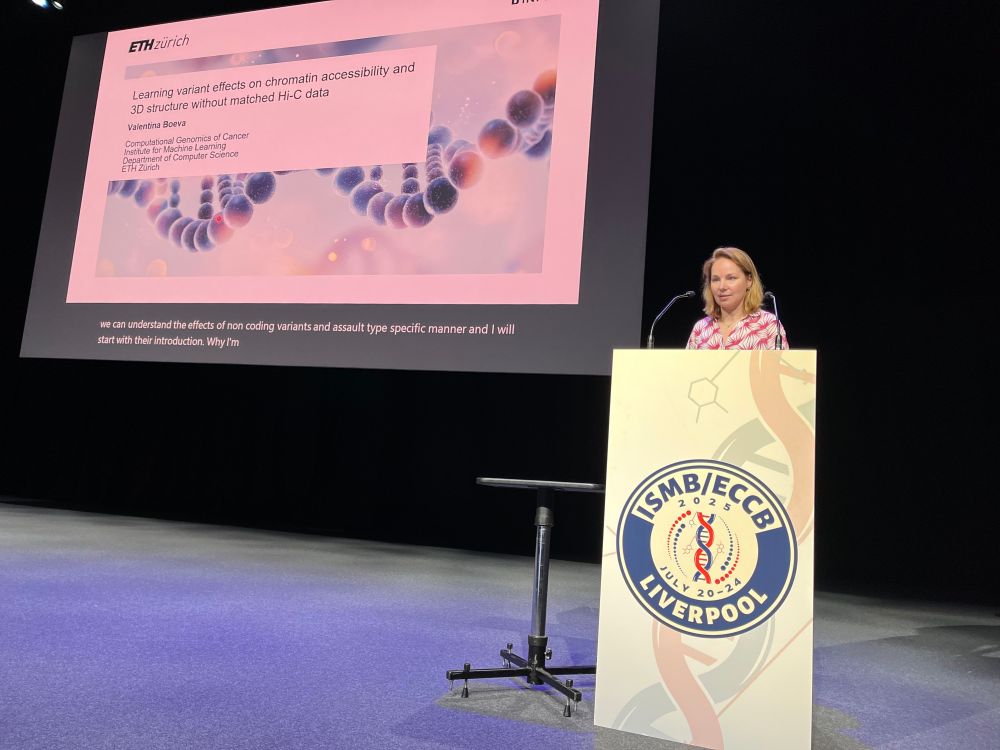


We have officially started #HitSeq track @hitseq.bsky.social at #ISMBECCB2025. Francisco de la Vega, introduces our first #keynote speaker Valentina Boeva @valboeva.bsky.social with her talk: "Learning variant effects on chromatin accessibility and 3D structure without matched Hi-C data"
23.07.2025 10:53 —
👍 5
🔁 2
💬 0
📌 0
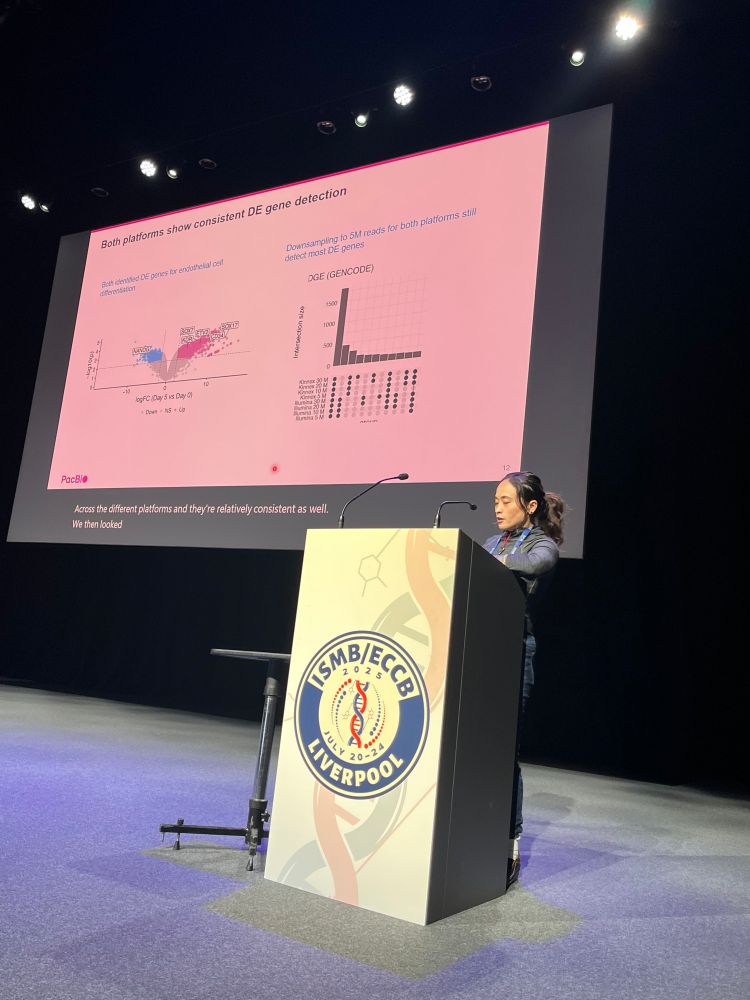
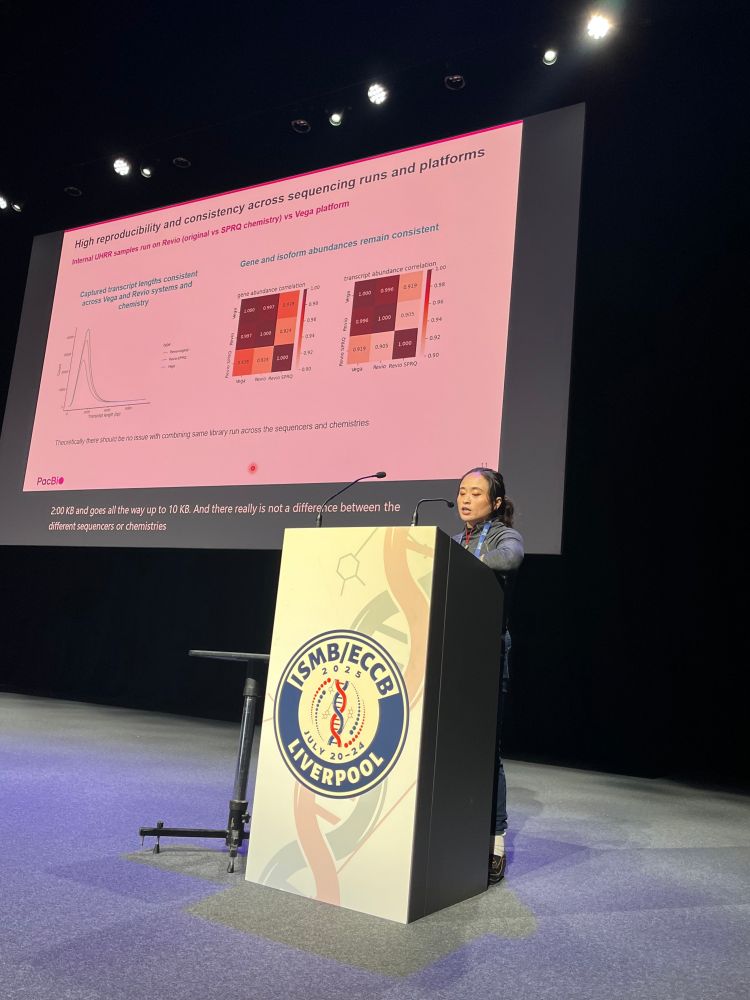
Meet our amazing sponsor PacBio @pacbio.bsky.social for @hitseq.bsky.social track at #ISMBECCB2025 represented by Elizabeth Tseng with her talk "Bioinformatics analysis for long-read RNA sequencing: challenges and promises" #hitseq #iscb #sequencing #application #iverpool #uk
23.07.2025 16:00 —
👍 3
🔁 1
💬 0
📌 0

Dont miss any of our #LongTREC communications at #ISMBECCB2025. Download this flyer to make catching all the latest & hottest long-read transcriptomics research simple.
@anaconesa.bsky.social
21.07.2025 12:30 —
👍 5
🔁 6
💬 0
📌 0


@hitseq.bsky.social is kicking off with our first keynote @valboeva.bsky.social talking about "Learning variant effects on chromatin accessibility and 3D structure without matched Hi-C data". #ISMBECCB2025
23.07.2025 10:38 —
👍 8
🔁 3
💬 0
📌 0
📽️ Next in the LongTREC Series: Mahmud Sami Aydin!
Sami is a Doctoral Candidate at @stockholm-uni.bsky.social , working under the supervision of @ksahlin.bsky.social .In this video, Sami shares his research and his role in the broader LongTREC collaboration across Europe.
#AlgorithmDevelopment
13.07.2025 08:28 —
👍 6
🔁 4
💬 0
📌 1
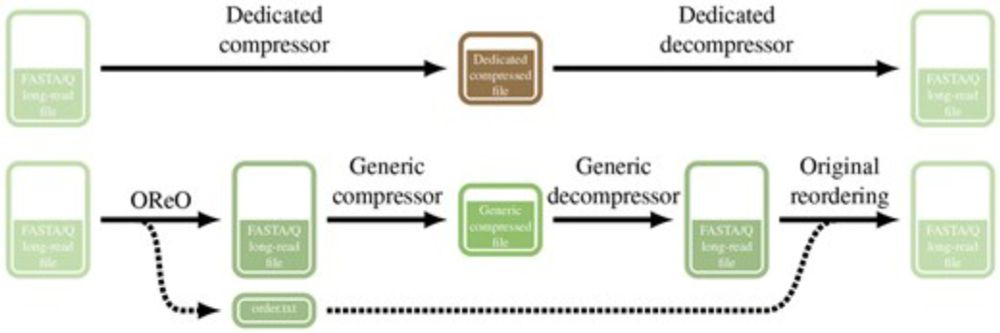
OReO: optimizing read order for practical compression
AbstractMotivation. Recent advances in high-throughput and third-generation sequencing technologies have created significant challenges in storing and mana
Paper alert!
We present Oreo a tools that reorder long reads datasets in a way to compress them efficiently with ANY universal compressor like gz, zstd, xz ...
TLDR: You can get state of the art compression WITHOUT a dedicated compressor/decompressor!
academic.oup.com/bioinformati...
A thread!
03.07.2025 10:52 —
👍 23
🔁 18
💬 1
📌 1
I worked with Thomas during a three months research visit during his PhD, and it resulted in a paper in NAR. I highly recommend him. doi.org/10.1093/nar/...
02.07.2025 11:48 —
👍 9
🔁 8
💬 0
📌 0

Thomas Baudeau defended his thesis on Studying the properties of viral long reads mapping methods - congrats docteur Baudeau you'll be deeply missed in the team. I'm very glad I got the chance to work with you. Thomas is also on the lookout for a postdoc 👀
30.06.2025 16:35 —
👍 7
🔁 3
💬 0
📌 2
🧵1/n
Estimating mutation rates using k-mers is fast—but what happens when repeats dominate the genome?
In a new preprint, Haonan Wu, Antonio Blanca, and myself propose a *repeat-aware* estimator that's accurate even in centromeres.
25.06.2025 13:19 —
👍 30
🔁 14
💬 1
📌 0

Hey yeast lovers. Do you like pangenomes?
O'Donnel et al. 2023 produced T2T assemblies of different strains, including phased haplotypes for yeast.
Here I selected 10 phased haplotypes and the S288C reference,
and looked for the MST28 / YAR033W gene reported to contain SVs such as indels.
👇🏻👇🏻
11.06.2025 14:46 —
👍 9
🔁 7
💬 2
📌 0
Congrats 👏👏
09.05.2025 22:37 —
👍 1
🔁 0
💬 1
📌 0
IMO it matters a lot as a 'first impression'
08.05.2025 13:34 —
👍 1
🔁 0
💬 0
📌 0
I did only very minor impl. contributions, but from my (non-expert) view, I like that (1) it installs easily (also on a MacBook) and (2) no header files. Felt much easier to get started with than, e.g., C++. I never truly learned good .h/.cpp practices, and I could never get OpenMP/g++ working well
08.05.2025 13:33 —
👍 2
🔁 0
💬 1
📌 0
As for results, isONclust3 handles a 37M reads PacBio dataset from a revio machine in under 10h while other algorithms fail (>256Gb mem or >120h runtime). On the other datasets, isONclust3 has comparable or better accuracy than the other benchmarked tools.
08.05.2025 13:04 —
👍 0
🔁 0
💬 1
📌 0
The algorithm follows isONclust's algorithm in the general structure (greedy minimizer matching) but adds three key concepts: high confidence minimizers, on-the-fly cluster information update, and iterative (post-)cluster merging.
08.05.2025 13:04 —
👍 0
🔁 0
💬 1
📌 0
The motivation to develop this algorithm came from the inability of other algorithms to process recent large datasets (10-100M reads) from Revio or PromethION machines.
08.05.2025 13:04 —
👍 0
🔁 0
💬 1
📌 0

@tolyan.bsky.social is our very last speaker, on randstrobes ( high sensitivity seeds ) and their evolution the multi context seeds
25.04.2025 07:39 —
👍 1
🔁 2
💬 0
📌 0