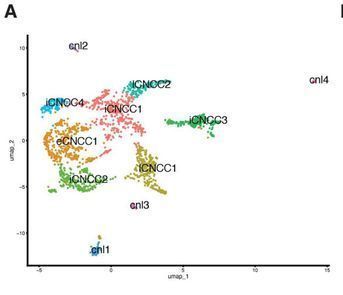
Great collaboration between current and former members of my lab including Nagham Khouri-Farah and @EmmaWentWin, the Skene Lab @n_skene , and the Leslie lab at Emory. Stay tuned for accessiblity data revealing cell type specific enhancers!
21.01.2025 21:38 —
👍 4
🔁 0
💬 0
📌 0

We also integrate with human genetic data revealing cell types that contribute to both normal facial variation and craniofacial disease risk. There's even some evo devo tidbits including cell types that are likely to contribute to differences in human and neanderthal morphology
21.01.2025 21:38 —
👍 4
🔁 0
💬 2
📌 0
Impressive study from Alex Stark lab identifying 3 new types of silencer elements in Drosophila
New TF binds a particular isolated motif in non-accessible sites to recruit G9a. Deletion of these elements can lead to upregulation of nearby gene
www.sciencedirect.com/science/arti...
20.11.2024 21:34 —
👍 51
🔁 16
💬 1
📌 1