Redirecting
Our most recent work on the “function and evolution” of #nuclear-speckles is now online at Cell @cp-cell.bsky.social
doi.org/10.1016/j.ce...
Read the thread👇 for the highlights of our findings.
25.02.2026 16:01 —
👍 120
🔁 58
💬 9
📌 5
🚨 1/ Preprint Alert!
Sex determination outcome is conserved across vertebrates (i.e. generating 2 compatible sexes) ♀️♂️
But are the cell types and gene programs behind them conserved too? 🧬
Spoiler: not really 👀
Find out in our new preprint ⬇️
www.biorxiv.org/content/10.6...
03.02.2026 10:02 —
👍 23
🔁 9
💬 1
📌 1
Job offer 2026.pdf
🧬 We’re #Hiring a #Bioinformatician/ Computational Biologist!
#Single-cell genomics, #Epigenomics & #3Dgenome biology in #Zebrafish #DevBio 🐟
📍 @cabd-upo-csic.bsky.social, CABD, Seville (Spain)
🕒 Full-time, 3-year position (start March 2026)
📅 Apply by Feb 15, 2026
👉 drive.google.com/file/d/1M8rp...
12.01.2026 15:57 —
👍 17
🔁 22
💬 1
📌 2
Woohoo! Congratulations, Fany!
02.12.2025 20:04 —
👍 2
🔁 0
💬 1
📌 0

📢 We are hiring!
👩🏾⚕️Research technician/Lab manager
📍 @cabd-upo-csic.bsky.social , Sevilla, Spain
🧬 Gene regulation in animal development & evolution
Check the details 👇 & join us!
drive.google.com/file/d/1cZJi...
#ResearchTechnician #Zebrafish #GeneRegulation #DevelopmentalBiology
15.10.2025 15:00 —
👍 8
🔁 9
💬 0
📌 2
We’re looking for a research technician to join our teams with @christinapaliou.bsky.social
Passionate about Zebrafish and gene regulation? 🐟🧬
Apply now and dive into exciting research with us! ✨
See the job offer below 👇
17.10.2025 14:04 —
👍 3
🔁 0
💬 0
📌 0

Out today. 🙏 again to everyone for this wonderful piece of work, in particular to Aurelie @aurhin.bsky.social Chase @chasebolt.bsky.social and Brent @homeobox.bsky.social. 🙏 also to the Harris lab @fish4walking.bsky.social and @neilshubin.bsky.social @biology-unige.bsky.social @college-de-france.fr
17.09.2025 17:30 —
👍 96
🔁 41
💬 2
📌 1

hoxd13a reporter in a zebrafish larva
Are digits modified fins, or evolutionary innovations? Read how we tackled this old question from a new angle🧪
A story with @chasebolt.bsky.social, @homeobox.bsky.social and myself, coordinated by @denisduboule.bsky.social from @college-de-france.fr and published in @nature.com today!
#InHoxWeTrust
17.09.2025 17:26 —
👍 113
🔁 44
💬 4
📌 6

Great work from the #CABD researchers in this call of @ageinves.bsky.social for the ‘Proyectos de generación de conocimientos’!
The provisional resolution proposes funding for:
🟢 13 projects
🟢 3 PhD contracts
🟢 ~ 3,5 million €
Way to go!!!
29.07.2025 16:51 —
👍 11
🔁 3
💬 0
📌 2
No new genes needed to fly - just rewire what you have! 🦇🧬
Great new paper from the labs of @fany-real.bsky.social @stemundi.bsky.social @dariloops.bsky.social
doi.org/10.1038/s415...
#EvoDevo #SingleCell #BatWings
16.07.2025 13:49 —
👍 118
🔁 44
💬 3
📌 1
Congratulations, Julian @julianeg.bsky.social! It was an exciting project, and I am happy to have contributed. Wonderful to see the paper in its final form.
10.07.2025 12:10 —
👍 2
🔁 0
💬 1
📌 0

POSTRE: a tool to predict the pathological effects of human structural variants
Abstract. Understanding the pathological impact of non-coding genetic variation is a major challenge in medical genetics. Accumulating evidences indicate t
3/3
Fantastic teamwork with Víctor Sánchez Gaya @sanchezgaya.bsky.social, Álvaro Rada Iglesia, and long-time collaborators Adrian Daly and Giampaolo Trivellin @gp83.bsky.social. Grateful to be part of it! 🙌
Original POSTRE paper - a powerful tool for clinical genetics!
🔗 tinyurl.com/3ys78kvc
09.07.2025 13:47 —
👍 2
🔁 0
💬 1
📌 0
2/3
In collaboration with @radaiglesiaslab.bsky.social, we applied and expanded POSTRE, a bioinformatics tool that rapidly classifies structural variants in the pituitary, distinguishing pathogenic duplications from benign ones in diagnostic settings.
#SVs #GPR101 #enhancers #TADs #TADopathies
09.07.2025 13:47 —
👍 3
🔁 0
💬 1
📌 0
I am very proud of you, Marta. You did outstanding work in the lab, and your defence was spotless. I am also very happy to now know an expert in gene compensation between paralogous genes!
02.07.2025 09:32 —
👍 2
🔁 2
💬 0
📌 0

Reconfiguration of genome-lamina interactions marks the commissioning of limb cell-fates
Diverse forms of heterochromatin block inappropriate transcription and safeguard differentiation and cell identity. Yet, how and when heterochromatin is reconfigured to facilitate changes in cell-fate...
🚨 Preprint 🚨
Ever wondered how cells prepare their genomes to enable new cell-fates? In this team up with the Kind lab, we show that genes are repositioned in the nucleus to get ready for future activation and tissue formation. Read 🧵👇 to find out how and when this happens!
doi.org/10.1101/2025...
08.05.2025 09:42 —
👍 36
🔁 23
💬 1
📌 3

Our BAT paper is accepted in Nature Ecology & Evolution! Single cell, gene regulation, TADs and key TFs that shape the bat wing. Genomics goes Evodevo 😍
06.05.2025 08:34 —
👍 64
🔁 16
💬 1
📌 0
😱 A very hungry #PacManoid ate the original post 🍬🍬🍬
Here’s the 🧵 on our latest in @cp-cellstemcell.bsky.social again!
Gastruloids w/ a sweet tooth better model natural embryos: doi.org/10.1016/j.st...
Spearheaded by @cryaaa.bsky.social & Alba Villaronga-Luque @mpi-cbg.de @poldresden.bsky.social
19.04.2025 08:48 —
👍 78
🔁 17
💬 6
📌 12

We’ve just launched our new website! 🚀
Check it out and stay tuned for updates! 🖥️ 🧬 🧫 🐁
fmreallab.github.io/website/
19.03.2025 14:08 —
👍 9
🔁 4
💬 0
📌 0
 27.02.2025 12:07 —
👍 0
🔁 0
💬 0
📌 0
27.02.2025 12:07 —
👍 0
🔁 0
💬 0
📌 0

Wow, Google... #GeographyFun or just going the extra mile to keep someone happy?
13.02.2025 12:44 —
👍 3
🔁 1
💬 0
📌 0

Implantation of engineered adipocytes suppresses tumor progression in cancer models - Nature Biotechnology
Adipose manipulation transplantation can reduce tumor growth and proliferation in vitro and in mouse models.
Tumors need a lot of nutrients to grow. By engineering and implanting fat cells to outcompete cancers for nutrients we can suppress them. Novel cancer therapy called Adipose Manipulation Transplantation (AMT). AMAZING work led by and dedicated to Hai Nguyen.
www.nature.com/articles/s41...
04.02.2025 15:49 —
👍 27
🔁 7
💬 2
📌 3
X account deactivated. Rest in peace, Twitter.
"Ich kann nicht so viel fressen, wie ich kotzen möchte." Max Liebermann, 1933
22.01.2025 07:50 —
👍 0
🔁 0
💬 0
📌 0
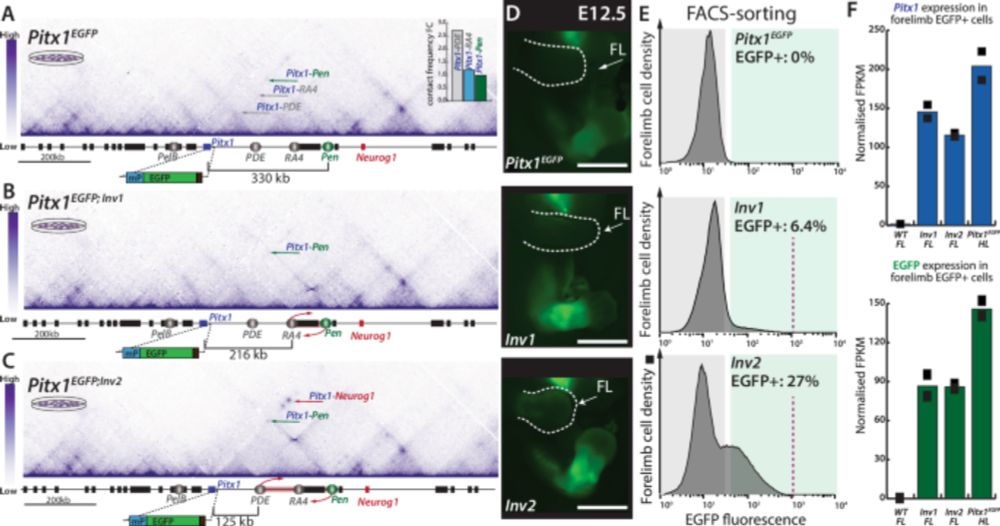
Ever wondered how 3D chromatin rewires during differentiation? We wanted to understand this process better & distinguish if different processes or elements drive this rewiring. To do so, we used mESC=>NPC differentiation and focused on a single locus. biorxiv.org/content/10.110… 1/17
01.12.2024 14:52 —
👍 25
🔁 10
💬 3
📌 0












 27.02.2025 12:07 —
👍 0
🔁 0
💬 0
📌 0
27.02.2025 12:07 —
👍 0
🔁 0
💬 0
📌 0

