Hello everyone, I am looking for my next role in the computational chemistry/biophysics space 🛫
I have 12 years of research experience in protein simulations/SBDD, looking for national lab/industry scientist positions anywhere in the world. Any connections/leads welcomed - thanks in advance! 🙏🏻
05.03.2026 20:59 —
👍 0
🔁 0
💬 0
📌 0
Paper was rejected today! Scanning through the reviews, there were some factual/scientific errors that need to be corrected before it can be published 😱 Happy that peer review caught this, would much rather publish a better paper later than rush one out with errors
24.11.2025 22:10 —
👍 2
🔁 0
💬 0
📌 0
Congrats Isabelle, that's great!
24.11.2025 22:01 —
👍 0
🔁 0
💬 0
📌 0
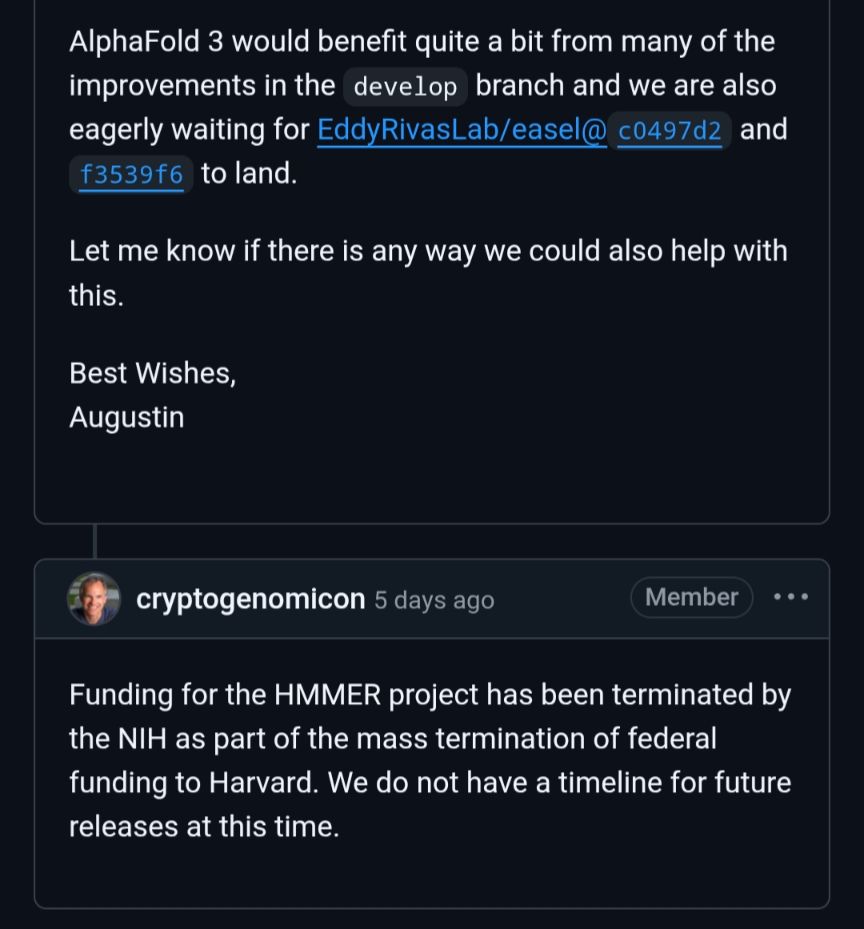
The war on science in the US is already having an effect on private sector research like AlphaFold. Bears repeating but the private sector builds on top of things created by academic research for the public good. This hurts everyone.
28.05.2025 10:18 —
👍 239
🔁 105
💬 2
📌 12
Are you a grad student or postdoc at the bench in the UC system? If so, please DM.
(for the record, this is a question about laboratory procedure, not any individual PI)
#chemsky ⚗️
22.05.2025 18:38 —
👍 8
🔁 4
💬 0
📌 0
congrats Kai + co! 🥳
13.05.2025 05:15 —
👍 0
🔁 0
💬 0
📌 0
Here at the BV-BRC workshop - refreshing to be at a workshop without a set problem to solve. Learning just for the sake of learning 😉
05.05.2025 21:53 —
👍 1
🔁 0
💬 0
📌 0

Examples of alternate conformations with environmental explanations.
(A) a protein in two states (open and closed) mediated by allosteric effects; (B) a population of two discrete rotamer states of a sidechain; (C) bound and unbound states due to ligand interaction; (D) different oxidation states of a cysteine pair. Furthermore, some regions in many protein structures exhibit such a degree of flexibility, that no accurate modelling is possible (dashed lines, E). The backbone conformation cannot be determined with crystallographic methods in such cases.

Failure of ensemble prediction algorithms in altloc segments.
Although structure and ensemble prediction techniques based on amino acid sequences are capable of accurately reconstructing the overall backbone structure (left column), they consistently fail to produce viable samples corresponding to one of the two altloc conformations (as shown in the magnified regions in the second through fourth columns).
Some crystal structures show evidence of multiple stable conformations, such as loops or side chains with two distinct regions of density. These can be correctly modeled by MD, but not modern protein ensemble prediction tools like AlphaFlow or DiG (BioEmu not tested)
www.biorxiv.org/content/10.1...
23.04.2025 14:32 —
👍 45
🔁 12
💬 0
📌 0
every time I'm off blue sky for more than a day or two and return, my feed (I only follow scientists) morphs into a tsunami of political clickbait, worse than twitter. Blocking left and right but it never gets any better. Spending less and less time here because of all the political junk smh
11.04.2025 21:02 —
👍 0
🔁 0
💬 0
📌 0
not attending but met up with @leftymiller.bsky.social at #ACSSpring2025! I feel people are always excited about the science at conferences but I'm always excited about the connections and the people behind the science
25.03.2025 22:51 —
👍 3
🔁 0
💬 0
📌 0
Water-mediated interactions between glycans are weakly repulsive and unexpectedly long-ranged https://www.biorxiv.org/content/10.1101/2025.02.16.638483v1
21.02.2025 01:48 —
👍 2
🔁 1
💬 0
📌 0

Figure 1 from arXiv preprint https://doi.org/10.1101/2025.01.06.631610
Fig. 1 Espaloma is an end-to-end differentiable molecular mechanics parameter assignment scheme for arbitrary organic molecules. Espaloma (extensible surrogate potential optimized by message-passing) is a modular approach for directly computing molecular mechanics force field parameters FFF from a chemical graph G such as a small molecule or biopolymer via a process that is fully differentiable in the model parameters FNN. In Stage 1, a graph neural network is used to generate continuous latent atom embeddings describing local chemical environments from the chemical graph. In Stage 2, these atom embeddings are transformed into feature vectors that preserve appropriate symmetries for atom, bond, angle, and proper/improper torsion inference via Janossy pooling.54 In Stage 3, molecular mechanics parameters are directly predicted from these feature vectors using feed-forward neural networks. This parameter assignment process is performed once per molecular species, allowing the potential energy to be rapidly computed using standard molecular mechanics or molecular dynamics frameworks thereafter. The collection of parameters FNN describing the espaloma model can be considered as the equivalent complete specification of a traditional molecular mechanics force field such as GAFF38,39/AM1-BCC55,56 in that it encodes the equivalent of traditional typing rules, parameter assignment tables, and even partial charge models. Reproduced from ref. 49 with permission from the Royal Society of Chemistry.
Everything is chaos, but I wanted to share some awesome recent science from the lab that hints at where the future of biomolecular simulation is headed:
Foundation simulation models that can be fine-tuned to experimental free energy data to produce systematically more accurate predictions.
19.02.2025 19:30 —
👍 107
🔁 30
💬 3
📌 1
exciting times! Will definitely be trying this out
17.02.2025 19:06 —
👍 0
🔁 0
💬 0
📌 0

🚨Preprint Alert!🚨
#ProteinDesign is advancing rapidly—wouldn't it be great to seamlessly combine design tools to achieve more than what each can do alone?🤔
Here, we introduce AI.zymes: A modular platform for evolutionary #EnzymeDesign.♻️🖥️
biorxiv.org/content/10.1...
1/🧵
12.02.2025 19:01 —
👍 30
🔁 7
💬 1
📌 1

We’re thrilled to announce the launch of Bind Research, the UK’s first not-for-profit Focused Research Organisation, with long-term support from the Department for Science, Innovation and Technology’s Research Ventures Catalyst Programme, industry, and charities.
11.02.2025 09:24 —
👍 25
🔁 13
💬 1
📌 2
We developed a computationally efficient method to predict hot spots driving conformational changes in small molecule sensing proteins. It can easily be generalized to explore allosteric couplings in a wide range of proteins. Code available in the preprint, check it out ! #compchem #compbio
11.02.2025 19:12 —
👍 2
🔁 1
💬 0
📌 0

Nanobodies with extended CDR3 conformations are correctly predicted the majority of the time, whereas kinked CDR3 conformations are rarely correctly predicted
CDR3 loop conformation, specifically whether the C-terminus is kinked or not, is a major determinants of whether a nanobody-antigen complex is correctly predicted using SOTA protein structure prediction software doi.org/10.1021/acs....
13.02.2025 07:06 —
👍 13
🔁 4
💬 0
📌 1
congrats Steffen + co!
01.02.2025 05:32 —
👍 1
🔁 0
💬 1
📌 0
Molecular Biophysics Database
Our colleagues in Prague have released a database for molecular biophysics data - mbdb-data.org! 🌍📊 A great step forwards FAIR data in biophysics. 🙌✨ @mosbri.bsky.social
31.01.2025 10:24 —
👍 15
🔁 6
💬 1
📌 1

There's just one week until the application period ends for the Frontera Fellowship! Collaborate with experts at the Texas Advanced Computing Center and use the powerful Frontera supercomputer to aid your research. Applications close February 7.
Learn more: bit.ly/425QQc7
31.01.2025 21:25 —
👍 1
🔁 2
💬 0
📌 0
It was helpful for me when I was writing my thesis, great job
31.01.2025 05:27 —
👍 0
🔁 0
💬 0
📌 0
Why LiveCoMS matters
| Living Journal of Computational Molecular Science
The Living Journal of Computational Molecular Science (LiveCoMS) is on BlueSky! Check out our recent blog post on why @livecomsjournal.bsky.social matters to find out more about us: livecomsjournal.org/index.php/li... #compchem #chemsky
30.01.2025 21:11 —
👍 21
🔁 9
💬 0
📌 0
Let’s stop treating "leaving academia" as negative. Scientists train for academia, industry, or what suits them. Industry isn’t an alternative; it’s a choice. In fields like engineering, most PhDs go to industry, and no one calls it "leaving academia." We should train scientists for diverse paths.
26.01.2025 11:03 —
👍 658
🔁 133
💬 30
📌 23
Our newest preprint is out. A PyMOL plugin giving access to a broad suite of high-performance, fast, and easy to use methods for virtual screening (50K molecules/h), docking, analysis of molecular interactions, dynamics, engineering permitting complex workflows.
24.01.2025 14:38 —
👍 9
🔁 3
💬 1
📌 1











