
Comparative evaluation of analytical methods for CSF proteomics link.springer.com/ar...
---
#proteomics #prot-paper

Comparative evaluation of analytical methods for CSF proteomics link.springer.com/ar...
---
#proteomics #prot-paper

(ACA) prepIMS: a Robust Data Preprocessing Workflow for Ion Mobility Mass Spectrometry Imaging: Publication date: Available online 28 November 2025
Source: Analytica Chimica Acta
Author(s): Linlin Wang, Chengyi Xie, Lei Guo, Thomas Ka Yam Lam, Chris Kong Chu Wong, Mudassir… #AChimActa #MassSpecRSS

Rapid assay development for low-input targeted proteomics using a linear ion trap — a promising step toward accessible, high-sensitivity proteomics workflows.
🔗 www.nature.com/articles/s41...
Looking to master Skyline but lack time? Skyline Online returns!
Join our accelerated Small Molecule course Oct 8-10, 9am-5pm EST, covering comprehensive Skyline applications in metabolomics, lipidomics, and beyond for MS data from any major vendor.
Just $175 USD to go from intro to expert.

We’re excited to invite you to a career panel discussion featuring three outstanding speakers: Rudolf Aebersold, Marcus Bantschoff, Pascal Gaudet
📅 Date: November 27
⏰ Time: 2:00 PM
Don't miss it out and hear the inspiration from leaders in the field!
👉 Register here: docs.google.com/forms/d/e/1F...

NEW PUBLICATION FROM THE GROUP IN #JEV!
A compendium of bona fide reference markers for plant #EVs🌿
isevjournals.onlinelibrary.wiley.com/doi/10.1002/...
🙌 Impressive work by: Miriam M.R. de Lope, Ibone Rubio, @educhicano.bsky.social and more!
@isev.org @snevresearch.bsky.social
Highlights here!👇

It's now properly published. If you want to easily check important characteristics of your data before diving into complicated statistics, check out PSManalyst.
PSManalyst: A Dashboard for Visual Quality Control of FragPipe Results | Journal of Proteome Research pubs.acs.org/doi/10.1021/...
The three horsemen of the apocalypse
… hahahahahaha

(Proteomics) Enhancing Lipidomics With High‐Resolution Ion Mobility‐Mass Spectrometry: PROTEOMICS, EarlyView. #MassSpecRSS
14.08.2025 14:07 — 👍 1 🔁 1 💬 0 📌 0
This week’s #THEProteomicsShow continues to explore #AltProteomics, but this time more about the data. Ben @proteomicsnews.bsky.social and I sat down with Yasset @ypriverol.bsky.social and talked about non-mass spec data and related aspects. Find it wherever you find fine podcasts and enjoy!
15.08.2025 10:21 — 👍 8 🔁 2 💬 1 📌 2
(ACS Anal Chem) [ASAP] Subcellular-Resolution Molecular Pathology by Laser Ablation–Rapid Evaporative Ionization Mass Spectrometry: Analytical ChemistryDOI: 10.1021/acs.analchem.5c02013 #MassSpecRSS #ACSAChem
07.08.2025 20:05 — 👍 1 🔁 1 💬 0 📌 0
Happy to share our latest preprint doing low cell number (mini-bulk) and single cell #proteomics on tumour associated neutrophils from human glioblastoma where we find multiple functional states that would be invisible to scRNAseq, some showing pro-tumoural states with potential therapeutic value
27.07.2025 12:28 — 👍 64 🔁 18 💬 3 📌 3
Want to watch our timsTOF HT and Evosep acquire 100 bat plasma samples in real time? I know you do!! Will the column clog or things break? Tune in to find out!! I'm obviously procrastinating btw 😉
www.youtube.com/live/Mb2pJpQ...



We visited the IMIBIC Mass Spectrometry and Molecular Imaging Unit (IMSMI lab) in #Córdoba, headed by @educhicano.bsky.social. Edu’s team and the Advanced Microscopy unit (Gema García Jurado and Álvaro Carrasco Carmona) warmly welcomed us and shared their spatial omics knowledge.
04.07.2025 14:40 — 👍 1 🔁 1 💬 1 📌 0
If you asked me 5 years ago if it would be possible to use a de novo tool on DIA data, I would have thought it would only exist in science fiction. Love being proved wrong. Great work from Justin Sanders. #proteomics #massspectrometry
www.nature.com/articles/s41...

SPROUTS_DB: an implemented database of contaminants for extracellular vesicle proteomics studies www.biorxiv.org/cont...
---
#proteomics #prot-preprint

In-depth and high-throughput spatial proteomics for whole-tissue slice profiling by deep learning-facilitated sparse sampling strategy www.nature.com/artic...
---
#proteomics #prot-paper

Advances in total glycomic analysis including sialylated sub-glycan isomers by SALSA method #BBAAdv #MassSpec www.sciencedirect.com/science/arti...
17.03.2025 20:39 — 👍 2 🔁 2 💬 0 📌 0(BioRxiv All) Proteomic profiling uncovers sexual dimorphism in the muscle response to wheel running exercise in the FLExDUX4 murine model of facioscapulohumeral muscular dystrophy: FLExDUX4 is a murine experimental model of facioscapulohumeral muscular… http://dlvr.it/TJb0SP #BioRxiv #MassSpecRSS
17.03.2025 22:10 — 👍 0 🔁 1 💬 0 📌 0
🌟 Life Science Tools (Single-Cell, Spatial Biology, Proteomics, Next-Gen Sequencing)
Strongest growth drivers:
Single-cell and Spatial Omics:
10X Genomics ($TXG), NanoString (part of Bruker $BRKR), Akoya (part of Quanterix $QRTX), and emerging players like Vizgen.

DDM (n-dodecyl β-D-maltoside). This seems too good to be true?! It's giving us a huge boost in hydrophobic peptide signal.
07.03.2025 18:15 — 👍 36 🔁 4 💬 13 📌 0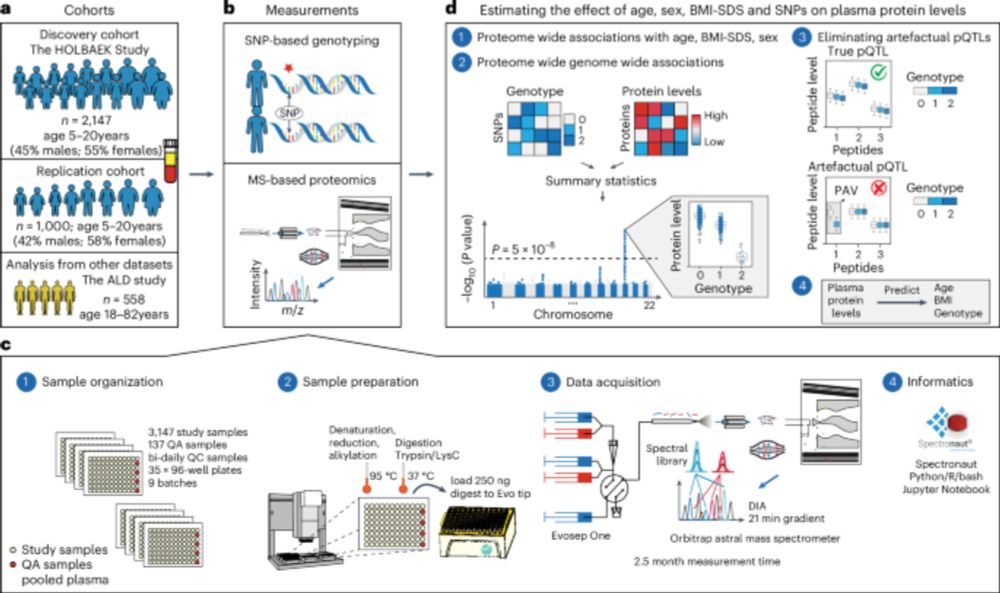
Out today in Nature Genetics: Using MS-based proteomics, we mapped 1,200+ plasma proteins in 2,100+ children, showing how genetics & development shape blood protein levels during childhood.
nature.com/articles/s41588-025-02089-2
#PediatricProteomics #pQTL
First author @liliniu.bsky.social explains ⬇️
Mass calibration algorithm in DIA-NN 2.0 can successfully correct even for the below 🙄😊. Apparently this kind of things can happen to some instruments.
If you spot sth odd in your data - please always let us know, sometimes troubleshooting is really quick and interesting things are discovered.

Courses at EMBL-EBI Data science for life scientists. Hinxton. Closes 2 March. Data visualisation for biology. Hinxton. Closes 9 March. Systems biology: from large datasets to biological insight. Hinxton. Closes 16 March. Proteomics bioinformatics. Hinxton. Closes 6 April.
We have several courses open for applications - whether you're a #lifescientist hoping to get to grips with #datascience, or you want to improve your #dataviz skills or you're hoping to learn about #omics and #bioinformatics then take a look!
More info: ebi.ac.uk/training/liv...
🧪🧬🖥️ #GeneSky
❗The Deadline for our EuPA Awards is extended to 16th of March❗
Send in now your application and join us at the #EuPA-FPS2025. We are looking forward to you EuPA Awards Application!
Hi all, the Finnish Proteomics Society is now on Bluesky! We just elected a new board and are ready to dive into new adventures.
#Finland #Proteomics #MassSpec #education @eupaproteomics.bsky.social
❗Only 3 days left to register ❗Join our 2nd YPIC Annual Proteomics Gathering in Athens. One day with fruitful discussions, amazing presentations, and a lot of networking! Here is the link to register: docs.google.com/forms/d/e/1F...
18.02.2025 15:16 — 👍 3 🔁 4 💬 0 📌 0
icbl2025.at
From today, registration for the #icbl2025 is open!
Don't miss this fantastic meeting and a gorgeous venue!

News in Proteomics Research blog post | Immunopeptidomics on low cell numbers without crappy antibodies! proteomicsnews.blogs...
---
#proteomics #prot-other

DIA-NN 2.0 is released! We consider it the biggest step forward in the history of DIA-NN. On modern LC-MS almost all identifications are now peptidoform-confident, with major improvements e.g. for phospho. Some other cool things too: github.com/vdemichev/Di...
29.01.2025 09:04 — 👍 150 🔁 39 💬 6 📌 2