Big congrats to my colleagues upstairs! Great work now published in @natchem.nature.com. Always happy to run #proteomics experiments with you. 🤝
17.09.2025 13:40 — 👍 3 🔁 1 💬 0 📌 0Big congrats to my colleagues upstairs! Great work now published in @natchem.nature.com. Always happy to run #proteomics experiments with you. 🤝
17.09.2025 13:40 — 👍 3 🔁 1 💬 0 📌 0
AIRPred: A Deep Learning Model Predictor for Peptide Intensity Ratios in Cross-Linking Mass Spectrometry Improves Cross-Link Spectrum Matching pubs.acs.org/doi/10....
---
#proteomics #prot-paper

Brace yourselves! Arif from the Fiedler lab discovered+characterized in collaboration with us & other groups within the @fmp-berlin.de a novel PTM: autocatalytic oligophosphorylation (up to 6 Phosphates 😲). Still mysterious in origin, function & regulation.
www.nature.com/articles/s41...
Thank you @nesvilab.bsky.social, it indeed became an invaluable tool throughout this and other projects that we're working on.
20.08.2025 18:36 — 👍 1 🔁 0 💬 1 📌 0A fun little side project from our FMP MS platform just got published! Previously, we struggled with messy Excel sheets to monitor machine performance and allocate machine time to users. Now, Q2C combines those management tasks into one user-friendly, GUI-based software. 💻
05.08.2025 16:34 — 👍 3 🔁 0 💬 0 📌 0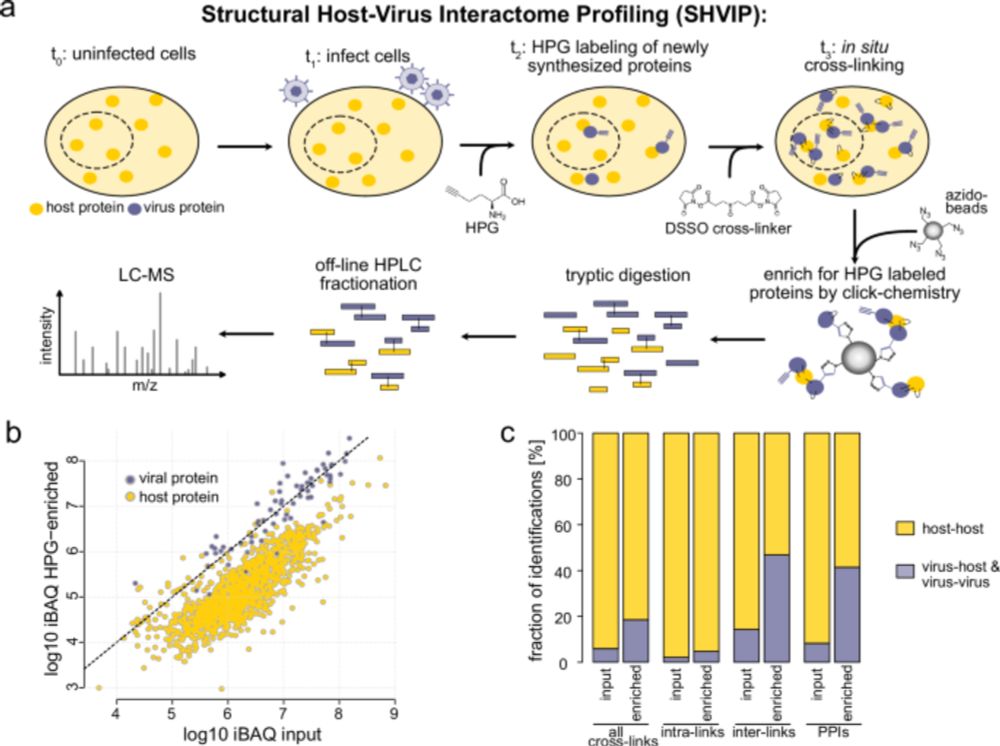
Check out Structural Host–Virus Interactome Profiling (SHVIP) in intact cells infected with Herpes Simplex Virus type 1 (HSV-1)! Now published!
📄 www.nature.com/articles/s41...
#LoveVirology #TeamMassSpec