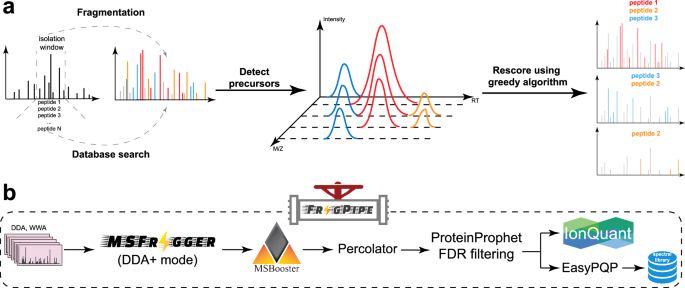
This project started 5 years ago. It led us to add isotope-labeling support to #FragPipe/#IonQuant. Since then, the tools have grown so much and are now widely used in #Chemoproteomics.
Huge thanks to everyone, and special thanks to @stephanhacker2.bsky.social and @pzanon.bsky.social
30.10.2025 14:15 —
👍 15
🔁 5
💬 0
📌 0
Start Page: /home/software/Skyline/events/2025 Webinars/Webinar 26
Proteomics Webinar: DIA with FragPipe, DIA-NN, and Skyline
Presenters: Eduard Sabidó and Brendan MacLean
When: Tuesday, September 16, 8am (Pacific Time)
Register Now ... skyline.ms/project/home...
#massspec #proteomics
15.09.2025 08:26 —
👍 33
🔁 16
💬 0
📌 0


It was a great pleasure to teach #FragPipe at the Biological Proteomics for Beginners workshop at #UCSD, sponsored by Thermo Fisher Scientific. We had a fantastic group of grad students, postdocs, and professors. Yes, I even got to teach UCSD professors how to analyze bottom-up proteomics data 😁
10.09.2025 14:52 —
👍 18
🔁 1
💬 1
📌 0
Interesting idea. May I ask two questions?
1. Is the precursor detection performed before or after the database search?
2. If it is performed before the search, how can one identify the complementary fragment ions without knowing the precursor mass or peptide sequence?
Thanks
14.08.2025 11:52 —
👍 0
🔁 0
💬 1
📌 0

Profiling the proteome-wide selectivity of diverse electrophiles
Targeted covalent inhibitors are powerful entities in drug discovery, but their application has so far mainly been limited to addressing cysteine residues. The development of cysteine-directed covalen...
Interested in the proteome-wide selectivity of diverse electrophiles?
The proteomics data for our study on this headed by @pzanon.bsky.social are now public on @pride-ebi.bsky.social: www.ebi.ac.uk/pride/archiv...
Full story: chemrxiv.org/engage/chemr...
#ChemBio #ChemSky #ChemicalProteomics
12.08.2025 09:50 —
👍 21
🔁 8
💬 0
📌 0

Conventional proteomics searches struggle with many modifications and fully open searches may be difficult to interpret. We introduce a "detailed" mass offset search in #MSFragger boosting interpretability and localization especially in complex cases (e.g. FPOP data): www.biorxiv.org/content/10.1...
01.08.2025 21:33 —
👍 34
🔁 6
💬 0
📌 0
Don't tell PD, but I have quietly switched all of my analyses to Fragpipe. What an extremely powerful software. I'm often amazed at the sheer quantity of information I can get using Fragpipe. Thanks to everyone who recommended it.
11.07.2025 20:49 —
👍 12
🔁 2
💬 2
📌 0
Excited to be one of the guest editors! We’re calling for papers on cutting-edge #proteomics and #metabolomics in #multiomics research. Looking forward to your submissions!
15.07.2025 06:21 —
👍 15
🔁 3
💬 0
📌 0
In addition to these exciting events, I’ll also be one of the panel members at the single-cell proteomics Evening Workshop on Wednesday. Looking forward to the discussions ahead!
30.05.2025 21:42 —
👍 7
🔁 1
💬 0
📌 0

Wow, a record breaking number of the Nesvizhskii lab members attending #ASMS2025! 9 posters, 3 evening workshops, and one Bioinformatics Hub on #FragPipe. Plus multiple collaborative posters with other groups. See you in Baltimore! PS. Below is our recent group photo, including all those attending
30.05.2025 19:45 —
👍 35
🔁 8
💬 0
📌 1
The power of #MSFragger open search! “we applied the mass-tolerant search engine MSfragger and found that phosphorylation as well as ubiquitination were well preserved after XDNAX. To our great interest, we found an additional modification of 321 Da occurring only in the irradiated SILAC channel”
23.05.2025 12:41 —
👍 14
🔁 2
💬 0
📌 0
If there any #Sciex decision makers here on Bluesky - I urge you to reconsider. Skyline/Proteowizard support is not only important for your customers using these tools, but it also benefits other bioinformatics efforts that depend on these tools.
15.05.2025 01:24 —
👍 25
🔁 9
💬 0
📌 1

We didn’t cherry-pick the dataset for the benchmark—FragPipe also delivers excellent performance on another dataset (PXD028735).
Questions are always welcome! We're always happy to hear your feedback. (3/3)
08.05.2025 01:22 —
👍 1
🔁 0
💬 0
📌 0

Peptide-level LFQbench-style plot using the PXD003881 (IonStar) dataset. (2/3)
08.05.2025 01:21 —
👍 1
🔁 0
💬 1
📌 0

We also benchmarked LFQ precision and accuracy against the latest versions of other popular tools. Here's an protein-level LFQbench-style plot using the PXD003881 (IonStar) dataset. (1/3)
08.05.2025 01:21 —
👍 1
🔁 0
💬 1
📌 0
In this release, one of the major improvements is LFQ using IonQuant. Thanks to this excellent preprint (www.biorxiv.org/content/10.1...), we identified and fixed a suboptimal step in the XIC. We're always eager to listen to feedback from the community!
07.05.2025 23:40 —
👍 25
🔁 4
💬 1
📌 0
Feature request well received🫡
04.03.2025 20:05 —
👍 5
🔁 0
💬 0
📌 0

One of the most significant and challenging projects of my career so far. PepCentric: a scalable computational platform utilizing novel 2-D fragment indexing for rapid peptide-centric searches, enabling proteogenomics searches against billions of spectra in seconds. www.biorxiv.org/content/10.1...
04.03.2025 09:09 —
👍 72
🔁 23
💬 3
📌 2

diaTracer enables spectrum-centric analysis of diaPASEF proteomics data www.nature.com/artic...
---
#proteomics #prot-paper
03.01.2025 17:20 —
👍 9
🔁 1
💬 1
📌 0
Looks interesting, and they already coupled it with MSFragger/FragPipe to search MS data using their fasta. I recommend using the group FDR option (available in FragPipe; requires annotating the headers in fasta in certain way) or a two-pass search option (canonical DB->unassigned mzML->custom DB).
10.12.2024 17:31 —
👍 8
🔁 1
💬 1
📌 0
No, we didn't exclude those shifts when looking for decoys. As you can see, there is room for us to improve. And I would like to do so if time permits.
06.12.2024 20:19 —
👍 1
🔁 0
💬 0
📌 0