
In a recent Nature paper, @djcalebc.bsky.social et al. explore how PIKfyve regulates lysosomal lipid recycling in PDAC. This opens up a therapeutic window: co-inhibition of PIKfyve and the KRAS–MAPK pathway leads to synthetic lethality & tumor regression.
View it here >>> nature.com/articles/s41...
19.06.2025 12:40 —
👍 1
🔁 1
💬 0
📌 0
My team and I are at #AACR25 and have posters to discuss parts of the project (and some future directions) tomorrow! Rüya Pakkan will present on Tuesday AM Abstract #4158 Sec 13 board 8. I will present on Tuesday PM #5384 Sec 12 board 12. Come visit!
28.04.2025 23:58 —
👍 0
🔁 0
💬 0
📌 0
Thank you!! And thank you for all your help and support along the way 😊
26.04.2025 21:49 —
👍 1
🔁 0
💬 0
📌 0
Thanks @bryantkl.bsky.social for the shoutout!! Look forward to seeing your future work including target PIKfyve as well as RAS in PDAC 💪
26.04.2025 21:49 —
👍 1
🔁 0
💬 0
📌 0
Thanks so much Siva!!! Look forward to seeing yours out soon 🫡
26.04.2025 21:46 —
👍 1
🔁 0
💬 0
📌 0
🫶
26.04.2025 21:45 —
👍 0
🔁 0
💬 0
📌 0
Thanks for this writeup and for highlighting our work!!
26.04.2025 21:45 —
👍 3
🔁 0
💬 0
📌 0
Thanks so much!!
26.04.2025 21:44 —
👍 1
🔁 0
💬 0
📌 0
🔥Smoking hot new paper and tweetorial from @djcalebc.bsky.social (collab with Chinnaiyan Lab)! 🔥
We are over the moon about this new work and its therapeutic implications for pancreatic cancer. Please have a read and share your thoughts/feedback! 🙏🏼
24.04.2025 20:20 —
👍 12
🔁 6
💬 0
📌 0
Excited to be part of this #AACR25 (@theaacr.bsky.social @rickbuck.bsky.social) Social Media Ambassadorship!!
Look out for updates on the #Tumor-microenvironment ( #TME )-relevant events and highlights. I’ll be posting from Sunday afternoon to midday Wednesday 😎
24.04.2025 21:28 —
👍 32
🔁 4
💬 0
📌 0
Thank you for the highlight!!
24.04.2025 15:22 —
👍 0
🔁 0
💬 0
📌 0
Thanks for the highlight!!!
24.04.2025 15:21 —
👍 1
🔁 0
💬 1
📌 0
13/ Finally thanks to the many friends for your support from both @lyssiotislab.bsky.social and the Chinnaiyan lab @brandontwchen.bsky.social @radykm.bsky.social @juliaugras.bsky.social @ja-nietocarrion.bsky.social @mruckert.bsky.social @shiva251993.bsky.social @federicadanzi.bsky.social 💜💜
24.04.2025 15:19 —
👍 7
🔁 0
💬 2
📌 0

12/ Also very grateful to officially have my first paper out in the pancreatic cancer field! This community has been so welcoming, and I look forward to working on PDAC throughout my training and career! @pancan.bsky.social
24.04.2025 15:19 —
👍 2
🔁 0
💬 1
📌 0

11/ THANK YOU to all the many contributors to this project, collaborators, mentors, and leaders. I am so honored to have been a part of this exciting study @awaddomi.bsky.social @lyssiotislab.bsky.social Arul Chinnaiyan and Yuanyuan Qiao.
24.04.2025 15:19 —
👍 3
🔁 0
💬 2
📌 0

We show this concept in efficacy studies using xenografts, orthotopic allografts, and KPC models in which the combo of RAS-MAPK and PIKfyve inhibitors resulted in regression/cures! Thanks especially to @jenmorton.bsky.social @macleanoncology.bsky.social @cruk-si.bsky.social @uofglasgow.bsky.social
24.04.2025 15:19 —
👍 3
🔁 1
💬 1
📌 0

a man in a leather jacket is making a funny face while looking at the camera .
ALT: a man in a leather jacket is making a funny face while looking at the camera .
8/ These suggest that co-targeting KRAS-MAPK and PIKfyve may result in synergistic effects through opposing effects on lipid metabolism!
24.04.2025 15:19 —
👍 0
🔁 0
💬 1
📌 0

7/ We then found that KRAS can regulate de novo lipid synthesis through regulating genes such as FASN and ACC1 through MAPK signaling, and that KRAS-MAPK inhibition partially reverses the transcriptional and metabolic adaptation that PIKfyve inhibition elicits on PDAC cells.
24.04.2025 15:19 —
👍 1
🔁 0
💬 1
📌 0
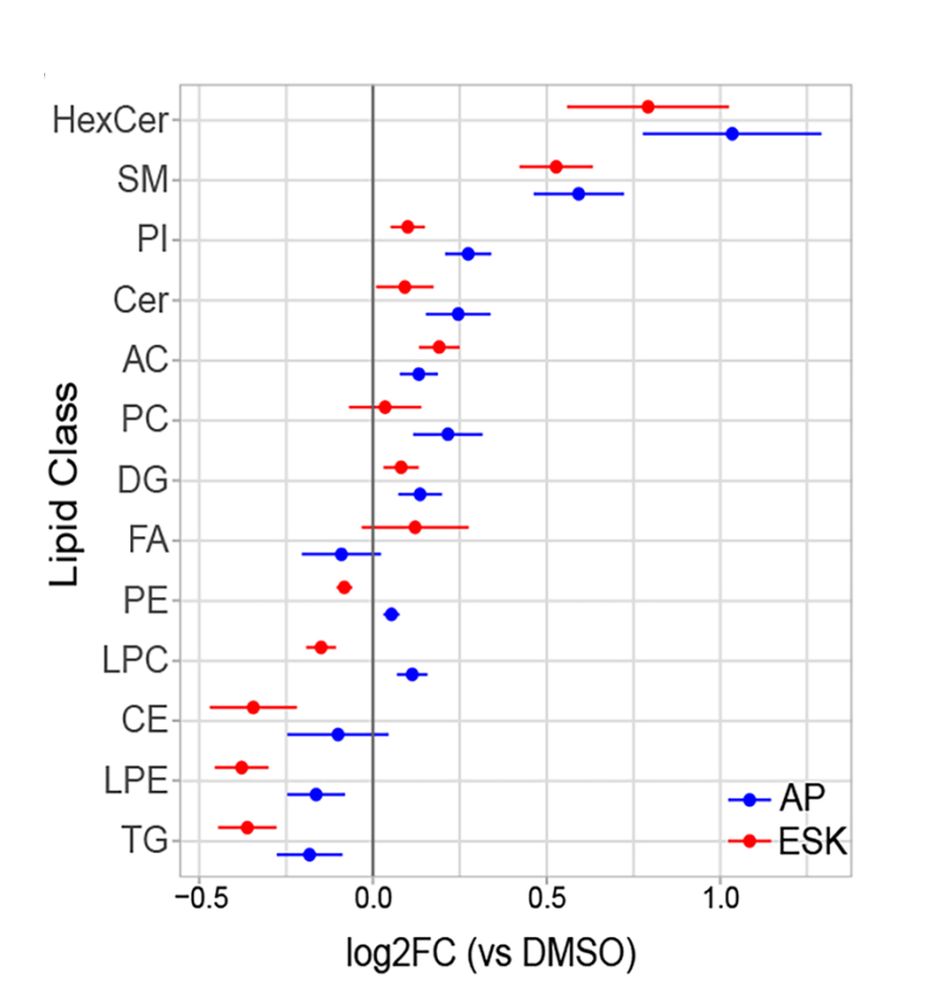
6/ RNA-seq, metabolomics, lipidomics, and stable isotope tracing strongly corroborated these results and suggested that PIKfyve inhibition triggers an adaptive lipogenic transcriptional and metabolic program. Special thanks to Dr. Pietro Morlacchi from Agilent for their help with the lipidomics!
24.04.2025 15:19 —
👍 1
🔁 0
💬 1
📌 0

5/ To assess the metabolic effect of inhibiting PIKfyve in PDAC, we ran a metabolism-focused CRISPR screen using PIKfyve inhibition as a selective pressure. The screen strongly demonstrated that PIKfyve inhibition forces PDAC cells rely on de novo lipid biosynthesis.
24.04.2025 15:19 —
👍 2
🔁 0
💬 1
📌 0

4/ We started with a GEMM developed by Yuanyuan Qiao demonstrating that loss of Pikfyve slows PDAC development in KC/KPC models while preserving pancreas function. Using a phase 1 cleared inhibitor ESK981, we also found that PIKfyve inhibition slows PDAC development and growth.
24.04.2025 15:19 —
👍 0
🔁 0
💬 1
📌 0

3/ Here, we show that inhibiting PIKfyve slows PDAC growth and induces an adaptive lipogenesis that can be blocked by KRAS inhibition. Follow along below for more in-depth tweetorial!
24.04.2025 15:19 —
👍 0
🔁 0
💬 1
📌 0
Beyond grateful to share that my main PhD thesis project, Targeting PIKfyve-driven lipid metabolism in pancreatic cancer (@lyssiotislab.bsky.social + Arul Chinnaiyan lab), has just been published in @nature.com!
24.04.2025 15:19 —
👍 12
🔁 6
💬 1
📌 2
Thank you!!!!
23.04.2025 20:05 —
👍 1
🔁 0
💬 0
📌 0

Mitochondria-organelle crosstalk in establishing compartmentalized metabolic homeostasis
Mitochondria serve as central hubs in cellular metabolism by sensing, integrating, and responding to metabolic demands. This integrative function is a…
Happy to share our most recent review on mitochondria-organelle metabolic communication with Yatrik Shah and Costas Lyssiotis now online at Molecular Cell!
We explored aspect of lipid transfer, organelle membrane contacts, ROS, metabolites, and metals!
www.sciencedirect.com/science/arti...
17.04.2025 21:24 —
👍 49
🔁 18
💬 1
📌 0
Excited for this!
30.03.2025 02:42 —
👍 1
🔁 0
💬 0
📌 0













