Square, neon-colored illustration announcing a “Daily Briefing” on mtDNA, showing mitochondria, DNA, AI icons, literature stacks, exercise symbols, and a bar graph, in a futuristic purple-pink style.
🧬 Introducing the #mtDNA Lab Intel Pipeline!
Our system daily scans PubMed and bioRxiv to bring you: 📄 #Gemini generated Daily Dossiers (PDF) and 🎙️ #NotebookLM generated Podcasts with a focus on #mtDNA-QC and #mtDNA-copy-number
📂 Full Archive:
drive.google.com/drive/folder...
07.02.2026 09:33 —
👍 8
🔁 4
💬 0
📌 1

Good morning everyone, below is last issue of #mitophagy news 🪫 🔋 🗑️
biomed.news/bims-tofagi/...
This week you'll find insights on the role of mitophagy in germline cells and in inheritance of mtDNA, in mice and in the male arabidopsis germline
Thanks to @biomednews.bsky.social for pre-sorting
26.01.2026 09:00 —
👍 8
🔁 5
💬 0
📌 0

Figure 1. Rab1A and Rab1B MitoID ident ifies novel Rab1 effectors. (A) Schematic of the Rab1-MitoID approach in which Rab1 is fused to the promiscuous biotin ligase BirA* and a mitochondrial localization signal from monoamine oxidase (depicted in red), resulting in the relocalization of Rab1 to the mitochondrial membrane. Rab1 interactors will be efficiently biotinylated, while other Golgi proteins and Rab interactors are not. (B) Two-dimensional plot comparing the enrichment of proteins biotinylated by Rab1A-MitoID vs control (BirA alone), plotted against the enrichment of proteins biotinylated with GTP-locked Rab1A (QL) vs GDP-locked Rab1A (SN). For the x axis, the value plotted is that for the nucleotide form of Rab1 that gave the greatest fold change over background. For clarity, Rab1A itself is not shown, as it is part of the BirA* construct. Known effectors and regulators are indicated, with known effectors being enriched in the upper right quadrant. Zoomed regions to the right show a region demarcated by well-enriched known effectors with novel proteins of note identified. For all protein identities and enrichment values see Table S1. (C) as for (B) except with Rab1B rather than Rab1A. (D) Overrepresented GO terms for biological process of the proteins within the regions demarcated by known effectors shown in B and C with the known effectors removed. Ranking is by the summed FDR and enrichment rank. (E) Immunoblots of binding to GST-Rab1A–coated beads of the indicated proteins from HEK293A cell lysates. Rab1A was either Q70L (GTP) or S25N (GDP). The known Rab1 effector RABEP1 (rabaptin-5) is included as positive control (Valsdottir et al., 2001). Source data are available for this figure: SourceData F1.
Welcome to another week of mitophagy papers! 🪫🗑️
biomed.news/bims-tofagi/...
highlights:
-A rab1 A Rab1 interactome illuminates a dual role in autophagy and membrane trafficking
-a review on the role of superoxide in mitophagy
-USP15 inhibition ameliorates PD phenotype in mouse models of disease
11.01.2026 10:50 —
👍 6
🔁 1
💬 0
📌 0
Not gonna lie, it does kind of feel like that
28.11.2025 08:55 —
👍 4
🔁 0
💬 0
📌 0

📢Last 2 weeks of #immunometabolism discoveries @biomednews.bsky.social ⬇️⬇️
biomed.news/bims-imicid/...
biomed.news/bims-imicid/...
Exciting to have a paper in the collection!
An inherited mitochondrial DNA mutation remodels inflammatory cytokine responses in macrophages and in vivo in mice
23.11.2025 13:05 —
👍 16
🔁 9
💬 0
📌 0

In this week's mitophagy news:
-a novel screen for endogenous modulators of PINK1-Parkin pathway uncovers unstudied dependencies
-autophagy regulation by phase separation, avidity and wetting
-a bacterial mitophagy receptor to evade inflammation
biomed.news/bims-tofagi/...
23.11.2025 12:04 —
👍 10
🔁 3
💬 0
📌 0

Liu et al. discover that pathogenic bacteria use a shared strategy to exploit the host’s cellular recycling process, #mitophagy. Through an effector protein that acts as an external receptor, bacteria suppress inflammatory responses to promote #infection rupress.org/jcb/article/...
#Autophagy
21.11.2025 17:45 —
👍 5
🔁 4
💬 1
📌 0
Really nice and clear summary :)
22.11.2025 15:29 —
👍 0
🔁 0
💬 1
📌 0
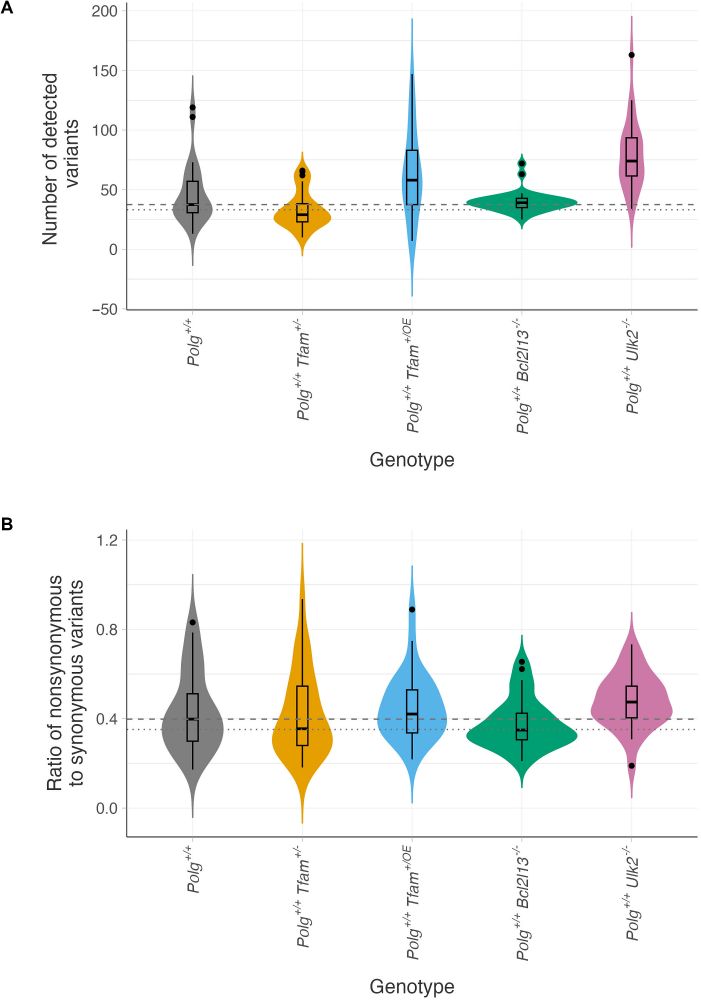
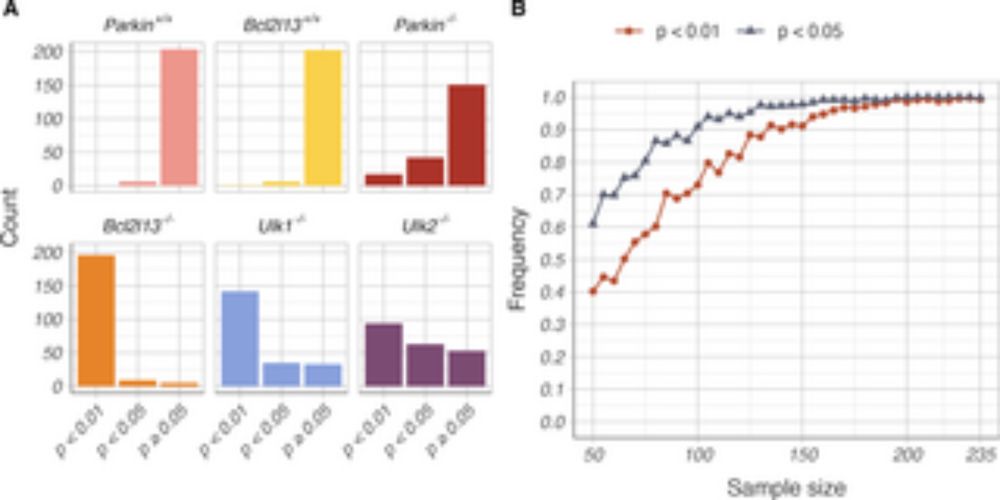
Altered bottleneck and mitochondrial turnover affect mtDNA mutation load and selection efficiency.
(A) Mutational burden was quantified as the total number of mtDNA variants detected per animal. (B) For each animal, the mtDNA N/S ratio was calculated by dividing the number of nonsynonymous variants, normalized to the number of possible nonsynonymous variants in the mitochondrial genome, by the number of synonymous variants, normalized to the number of possible synonymous sites. For (A) and (B): dashed line, median value of the Polg+/+ control group; dotted line, modal value of the Polg+/+ control group; gray, Polg+/+ control group; orange, Polg+/+ Tfam+/− group; blue, Polg+/+ Tfam+/OE group; green, Polg+/+ Bcl2l13−/− group; and pink, Polg+/+ Ulk2−/− group.
Only good papers on mitophagy this week!
biomed.news/bims-tofagi/...
Highlights:
-The bottleneck for maternal transmission of mtDNA is linked to purifying selection by autophagy
-Piecemeal Mitochondrial degradation in plants
-Allophagy?!
Thanks to @biomednews.bsky.social
16.11.2025 10:53 —
👍 9
🔁 5
💬 0
📌 0
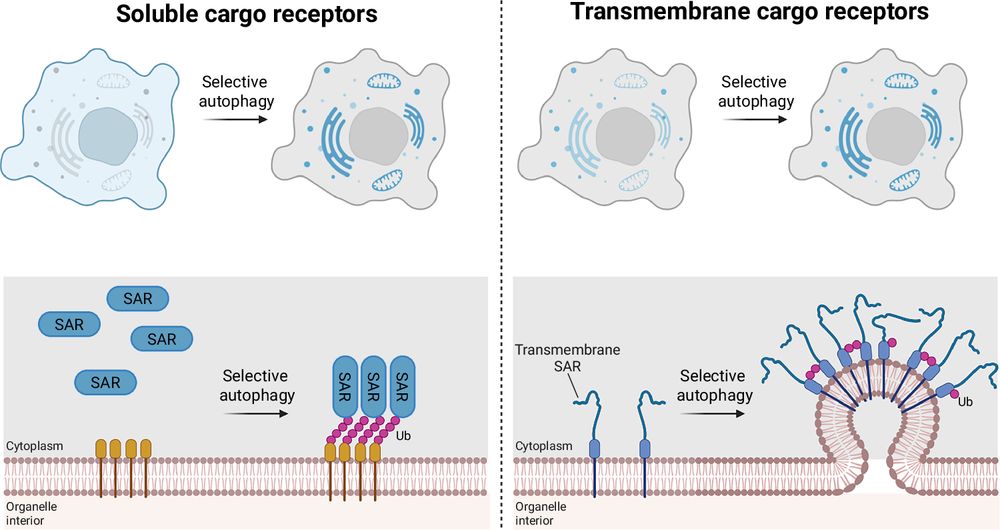
Left, soluble SARs such as SQSTM1/p62, NDP52, OPTN, TAX1BP1, and NBR1 are cytosolic proteins that are recruited to cargo on demand. Their recruitment is commonly driven by ubiquitin (Ub) marks on damaged or misfolded cargo, enabling rapid and flexible targeting of diverse substrates. Upon recruitment, these SARs concentrate autophagy initiation factors at the cargo site to initiate autophagosome biogenesis. Right, transmembrane SARs such as NIX, BNIP3, FAM134B, and CCPG1 are stably integrated within organelle membranes. To prevent unwanted autophagy activation, these SARs require additional layers of control. Selective autophagy can be triggered by transcriptional upregulation (e.g., hypoxia-driven induction of NIX/BNIP3), post-translational modifications such as ubiquitination, or SAR clustering. ER-phagy SARs further promote local membrane bending and scission to excise a portion of the organelle for autophagic removal. While some SARs are transcriptionally induced, others—such as FAM134B—are constitutively present and rely primarily on clustering and post-translational regulation, providing SAR-specific modes of control over autophagosome formation.
Mitophagy updates of last week 🪫🗑️
biomed.news/bims-tofagi/...
Highlights:
-two papers on the beneficial effects of mitophagy in CD8+ T cells
-VPS35 mutations in PD affect PINK1/Parkin mitophagy selectively
-review on selective autophagy initiation
Thanks to @biomednews.bsky.social for pre-sorting
04.11.2025 17:26 —
👍 7
🔁 3
💬 1
📌 0
Wonderful work from @mito-oncogene.bsky.social and @edreznik.bsky.social on the role of mito ribosomal RNA in cancer.
04.11.2025 08:01 —
👍 10
🔁 4
💬 1
📌 0

Assessment of mitophagy in the Malphigian tubules of the dmt-Keima Drosophila kidney
In this week's issue of mitophagy news 🗑️🪫
biomed.news/bims-tofagi/...
-High sugar diet inhbits renal mitophagy and disrupts renal function
-a review by @soleimanpourlab.bsky.social on mitophagy in diabetic ß-cells
-Optineurin is necessary for mitochondrial removal during erythrocyte maturation
26.10.2025 20:23 —
👍 5
🔁 3
💬 0
📌 0
New from us! Had a question about mitophagy but were afraid to ask? 😂
Excited to review the latest advances in beta cell mitophagy research as a key adaptive response in all forms of diabetes in
@cp-trendsendomet.bsky.social. We'd be honored if you have it a read! #T1D #T2D
22.10.2025 18:24 —
👍 27
🔁 9
💬 1
📌 1

q.e.d Science
Critical Thinking AI for constructive criticism and science evaluation
Here's the link to the system, try it! qedscience.com
@qedscience.bsky.social
15.10.2025 12:22 —
👍 203
🔁 66
💬 17
📌 16
🔥 Check out our new paper! @natcellbio.nature.com
Also thanks to @embo.org for their Postdoctoral Fellowship and especially to Mai and Heidi, I learned more than I could ever have imagined in Montreal 🇨🇦 🍁
14.10.2025 13:27 —
👍 17
🔁 3
💬 0
📌 1
Thank you so much 🥹 🗑️🪫❤️🥚
12.10.2025 09:42 —
👍 1
🔁 0
💬 0
📌 0
Thank you so much Cory! Keeping a look out for more great science from your side
11.10.2025 14:18 —
👍 1
🔁 0
💬 0
📌 0

Just out in @science.org: Together with the lab of @wilhelmpalm.bsky.social, we used sequential in-vitro/in-vivo CRISPR screens to decipher metabolic adaptations in tumors. We find that acidosis is a dominant factor that shapes energy metabolism and stress resilience. www.science.org/doi/10.1126/...
09.10.2025 19:36 —
👍 110
🔁 29
💬 1
📌 3
Thank you so much Alessandro! Hopefully see you again sometime soon!
10.10.2025 14:22 —
👍 1
🔁 0
💬 0
📌 0
Thank you Alex, for your help throughout this project and your ubiquitin and mitophagy wisdom! I was really luck to end up working in the same institute as your lab
10.10.2025 14:19 —
👍 1
🔁 0
💬 0
📌 0
Thank you so much Viktor :) Thank god you published your paper on NAD+, autophagy and cell death! The choice of galactose in Figure 4 is all thanks to your lab!! And seeing that the survival experiments followed what you observed was such an important sanity check
10.10.2025 10:58 —
👍 1
🔁 0
💬 1
📌 0
Thank youuu 😍😍 This would have been nothing without your Western blotting teachings
10.10.2025 10:01 —
👍 1
🔁 0
💬 1
📌 0
Thank you so much Dylan🤸 without your encouragement and wishes it would have taken even longer 🤣🤣
10.10.2025 10:00 —
👍 1
🔁 0
💬 1
📌 0