The demystification of piRNA clusters
if you wonder how cells generate piRNAs specifically against transposons & you are looking for a weekend read
check out @86dominik.bsky.social's opus magna (or Dominik's great thread)
a shared project with the one and only Rippei Hayashi, lab alumnus & friend
13.02.2026 20:39 —
👍 41
🔁 18
💬 2
📌 1
Do you want to understand how RNA #splicing decisions are made? We‘ve got a PhD position open for you (see 👇👇)!!
Please share #RNAsky and RT!
tinyurl.com/4ztu9cb9
11.02.2026 08:20 —
👍 9
🔁 12
💬 0
📌 0

Ulrich’s research will help advance our understanding of RNA biology & gene regulation, as well as how these mechanisms might be altered during viral infections & other diseases.
@hohmannulrich.bsky.social comes from the @impvienna.bsky.social / @imbavienna.bsky.social
www.imb.de/about-imb/ne...
09.02.2026 11:14 —
👍 40
🔁 16
💬 1
📌 1
Really excited to join IMB for this new scientific endeavour!
06.02.2026 14:49 —
👍 34
🔁 5
💬 1
📌 0

@lorenzoorts.bsky.social previously worked as a postdoctoral fellow at @impvienna.bsky.social in Austria. She will establish a new lab at IMB to investigate how mRNA translation is activated in the early embryo. Welcome to IMB Laura! 💐
Read more here: www.imb.de/about-imb/ne...
06.02.2026 14:27 —
👍 32
🔁 13
💬 1
📌 1
Very cool work! Reminiscent of how target engaged PIWI is selected from a vast pool of PIWI-piRNA complexes. Only target engaged PIWI is recognized by a GTSF1 protein and Maelstrom in the PIWI* complex. www.cell.com/molecular-ce... @juliusbrennecke.bsky.social @julsportell.bsky.social
06.01.2026 10:30 —
👍 4
🔁 1
💬 0
📌 0
When RNA Degradation 🤝 meets 🤝 Protein Degradation! tinyurl.com/E3TDMD In a collaboration of @bartellab.bsky.social and Schulman lab, we show that, in target-directed microRNA degradation (TDMD), 2-RNA-factors recruit an E3 ligase and induce the degradation of not only a protein but also RNA (1/5).
06.01.2026 08:04 —
👍 117
🔁 50
💬 1
📌 4
Very happy to share my postdoc work (on preprint), where we try to understand a long-standing transcription-silencing paradox, and uncovered a hidden RNA decay arm of nuclear piRNA pathway, see detailed threads quoted from @juliusbrennecke.bsky.social
22.12.2025 19:38 —
👍 33
🔁 12
💬 1
📌 0
Looking for a great first read for 2026? Look no further:
Fantastic work spearheaded by my courageous colleague @changweiyu.bsky.social, with surprising findings not only for small RNA but also for exosome and gene expression aficionados
02.01.2026 20:28 —
👍 10
🔁 4
💬 0
📌 0

As 2025 comes to an end, it’s time for me to say good bye to my postdoc home during the past 6 years – the Pauli lab @pauligroup.bsky.social @impvienna.bsky.social
14.12.2025 16:37 —
👍 41
🔁 4
💬 2
📌 1

How does messenger RNA (mRNA) get out of the nucleus to become a protein? Eukaryotic mRNA is packaged, exported, and then translated in the cytoplasm. But how do these steps work? And what are open questions? Check out our new review for our take: www.annualreviews.org/content/jour... (1/3)
21.11.2025 17:36 —
👍 122
🔁 52
💬 1
📌 3

Messenger RNA is made in the nucleus before it is exported to the cytoplasm for translation. But how are only correctly made mRNAs chosen and remodeled in the nucleus for export?
Our new paper investigates the nuclear events leading to human mRNA export. www.nature.com/articles/s41.... (1/4)
21.11.2025 05:02 —
👍 60
🔁 25
💬 2
📌 2
Research
IMB Mainz
Lastly, I’m excited to join @imbmainz.bsky.social in 2026 to start my own lab. We'll explore new mechanisms in eukaryotic gene expression, leveraging ‘evolutionary play’ to uncover how regulation, repurposing, and hijacking shape RNA biology. tinyurl.com/y4x29ctt
Thanks for reading! 20/20
19.11.2025 23:21 —
👍 19
🔁 5
💬 0
📌 1
And of course also a big ‘Thank you!’ to colleagues and support at @imbavienna.bsky.social, @impvienna.bsky.social, @viennabiocenter.bsky.social and for funding to @boehringerglobal.bsky.social, @erc.europa.eu, and my postdoc fellowships from @embo.org and Marie Skłodowska-Curie Actions
19/
19.11.2025 23:21 —
👍 6
🔁 0
💬 1
📌 0
This was a quite a journey, and would not have been possible without my mentors @juliusbrennecke.bsky.social & Clemens, my colleagues in the Brennecke and @plaschkalab.bsky.social labs, and the institutional support at the @viennabiocenter.bsky.social. 18/
19.11.2025 23:21 —
👍 2
🔁 0
💬 1
📌 0

Immunoprecipitations followed by quantitative mass spectrometry of wildtype (wt) or three UAP56 protein mutants probe the mRNA export model. An experiment schematic is shown on the left. The top heatmap (greyscale) shows log2 fold-changes in protein enrichment of wildtype UAP56 versus a control. The bottom heatmaps (blue-white-red scale), show fold-changes of three UAP56 mutants versus wildtype UAP56: UAP56 mutant M1 (D49R, L51D), UAP56–TREX-2 binding mutant; M2 (F336E, R339D), UAP56–THO-binding mutant; M3, combined UAP56 THO- & TREX-2-binding mutant (M1 & M2). The enrichment of GANP in UAP56 mutant M2 is likely due to the binding of free nuclear UAP56 mutant M2 protein to TREX-2.
Finally, we test the linearity of this model through IP-MS of UAP56 mutants. If we perturb TREX-2 binding (UAP56 mutant M1) -> UAP56 accumulates on mRNPs. THO binding (M2) -> UAP56 depletes from mRNPs. THO+TREX-2 binding (M3) -> mirrors loss of THO binding alone. 17/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0
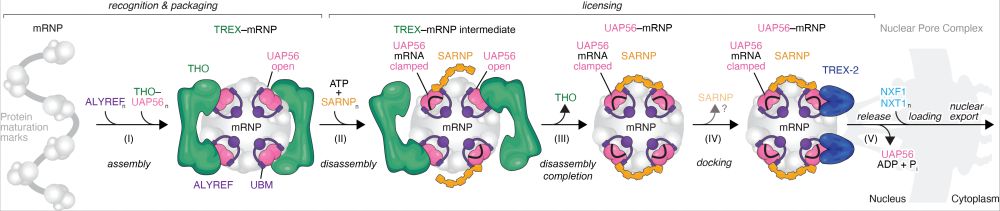
A general model for mRNA nuclear export.
The ATPase and mRNA clampase UAP56 is a central molecular switch that steers human mRNAs through mRNP packaging and multivalent TREX–mRNP globule assembly (I), TREX–mRNP disassembly and the formation of export-competent UAP56–mRNPs, assisted by the multivalent protein SARNP (II and III), mRNP docking (IV), and mRNP release (IV) at the NPC via TREX-2. Loading of the mRNA export factor NXF1–NXT1 onto mRNPs may occur in the nucleoplasm or at the NPC, initiating mRNP nuclear export.
Taken together, we find that UAP56 is an “mRNP packaging mark”, that controls important aspects of the nuclear events of the mRNA export pathway: disassembly of TREX, docking of the mRNP at the NPC, and release into the NPC. 16/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

UAP56 RNA-unclamping assay. Bead-immobilized 15 nucleotide poly-Uridine RNA is incubated with UAP56 and ATP to form UAP56–ADP-Pi–RNA complexes, which are then challenged with recombinant THO or TREX-2M complexes. Bead-remaining UAP56–ADP-Pi–RNA complexes are analyzed by SDS-PAGE (Coomassie-staining).
We can visualize this TREX-2 activity also directly: we bead-immobilize UAP56–ADP-Pi–RNA complexes. Incubation with TREX-2 releases UAP56 from the RNA. In contrast, as expected, the THO complex does not have any effect on clamped UAP56. 15/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

TREX-2 stimulates the apparent ATPase rate of UAP56 in vitro (left). Plotted are the ATPase rates of the RecA2P56–TREX-2 ‘M’ mixtures. The addition of TREX-2 accelerates UAP56’s apparent ATPase rate 57 fold.
The GANP wedge binding mimics the ATP-binding pocket in clamped UAP56 (middle and right). Shown are the UAP56-bound nucleotide and nucleotide base stacking residues in sticks representation, colored by heteroatom. In the UAP56–TREX-2 structure the adenine moiety of the non-hydrolyzable ATP analogue forms stacking interactions with F65 in the UAP56 RecA1 lobe and the GANP wedge residue R678 (middle). In clamped UAP56 (PDB ID 8ENK) (Xie et al., 2023), the adenine base of bound ADP stacks with F65 of UAP56’s RecA1 lobe and F381 of the RecA2 lobe (right).
A highly conserved ‘wedge’ loop of the TREX-2 subunit GANP binds the nucleotide, which is still bound to UAP56’s RecA1 lobe (unusual for an open DExD ATPase). In vitro ATPase assays show a dramatic stimulation of UAP56’s ATPase rate by TREX-2. So TREX-2 promotes RNA release from UAP56! 14/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0
However, to complete export, the mRNP must subsequently be released again from TREX-2. How does this happen? To our surprise, in our cryo-EM structure of the UAP56–TREX-2 complex UAP56 was not in an RNA-bound, closed conformation, but instead open?! 13/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Mutation of the UAP56 NTD–PCID2 interface in PCID2 leads to the accumulation of nuclear poly(A) RNA. Shown are representative cells (left; z-projection) and the ratios of the nucleo-cytoplasmic (N/C) poly(A) RNA FISH signal (right) after the depletion of endogenous PCID2 or GANP for 16 h, or upon the depletion and rescue of endogenous PCID2 with wildtype or mutant PCID2 constructs, M1 and M2. M1, PCID2 NTD-binding mutant (K374D, K388D); M2, PCID2 GANP-binding mutant (D356R, A365F). Scale bar = 10 μm; Four replicates per condition, with >70 cells per replicate. Pairwise significance testing was performed using two-sided Welch t-tests, with FDR correction for multiple testing, where *** P<0.001.
In cells we show: the identified interfaces are critical for cell viability. In addition, perturbing the UAP56–TREX-2 interaction leads to mRNA export defects, as assessed (amongst other assays) through polyA-FISH in human K562 cells with a knockout-rescue setup for the TREX-2 subunit PCID2. 12/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Cartoon representation of a Cryo-EM structure of human UAP56–TREX-2 middle region (TREX-2 ‘M’:GANP residues 602-942, PCID2, SEM1).
Could TREX-2 be a landing pad at the NPC for UAP56-bound, export ready mRNPs? Biochemically we show that UAP56 does indeed bind TREX-2 and solved a cryo-EM strcture of the complex. This revealed a role for the conserved UAP56 NTD in TREX-2 binding. 11/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

AlphaFold2 Multimer in silico protein interaction screen to identify novel UAP56 interactors. All UAP56–candidate predictions are shown (circles) and ranked by the average interface prediction TM score (interface pTM). Known and several novel predicted interactors are highlighted in colors.
Besides known UAP56 interactors we find some novel ones. Among them: GANP and PCID2, two subunits of the TREX-2 complex. A big surprise to us! TREX-2 (sharing only its name with TREX) is involved in mRNA export and bound to the basket of the nuclear pore complex (NPC)… 10/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0
So: TREX disassembly yields mature, export-ready mRNPs, marked by clamped UAP56. Could clamped UAP56 be a molecular mark controlling final steps of nuclear export? To find new UAP56 interactors we used AlphaFold2 Multimer to screen the nuclear UAP56 interactome. 9/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Native TREX–mRNP disassembly assay. Experiment schematic (left) and massspectrometry results (right, heatmap) of bead-retained mRNP-associated proteins after adding the ALYREF N-UBM, SARNP UCM-1, or an UCM-1–N-UBM fusion. The heatmap is colored according to the log2 fold-change compared to the buffer control, after normalizing to mean THO complex subunit levels.
Remarkably, SARNP disassembles THO-UAP56 in the presence of ATP and RNA in vitro. And in endogenous mRNPs? We purified endogenous TREX-bound mRNPs. Again, a minimal SARNP construct (including the UCM) could dissociate the mRNP from THO. 8/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Structural model of a clamped UAP56–ADP-Pi–RNA complex bound to SARNP and ALYREF. The model was obtained by superposing structures of clamped UAP56–ADP-BeF3–RNA (PDB-ID 8ENK) (Xie et al., 2023), open UAP56–ALYREF N- and C-UBMs (Fig. 1), and a UAP56–SARNP UCM-1 AlphaFold2 Multimer prediction.
Is RNA clamping of UAP56 regulated? We identify SARNP as a ‘clamping factor’. It has a motif we named ‘UCM’, which binds adjacent to the newly identified N-UBM binding site on UAP56. N-UBM and UCM cooperatively enhance UAP56’s affinity for RNA, this might aid directionality of mRNP remodelling. 7/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Size exclusion chromatography of UAP56 and ATP with or without 15 nucleotide poly-Uridine RNA. UV-traces at 280 nm (black) and 260 nm (grey) are shown.
Upon RNA clamping UAP56 hydrolyses ATP, and the resulting UAP56–ADP-Pi–RNA complexes are remarkably stable, for example persisting in size exclusion chromatography experiments. This suggests that UAP56 can be ‘deposited’ on the mRNA as a maturation mark. 6/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Representation of open (unclamped) and RNA clamped UAP56. Shown alongside are Grating Coupled Interferometry derived binding curves for open and RNA clamped UAP56 vs. the THO complex. Isolated, open UAP56 binds the tetrameric THO complex with nano molar affinity (top), whereas RNA-clamped, closed UAP56–ATP–RNA has no measurable THO complex affinity (bottom).
Could the conformational change in UAP56 between its open and its RNA-clamped state play a role? Turns out it does: Open UAP56 binds THO, RNA-clamped UAP56 has no measurable affinity! This suggests that clamping of UAP56 releases THO from the mRNP. 5/
19.11.2025 23:21 —
👍 0
🔁 0
💬 1
📌 0

Cartoon representation of the cryoEM structure of a human TREX complex. UAP56 binds ALYREF through an N- and C-UBM at its respective RecA2 and RecA1n domains, and is in an open conformation, primed to clamp onto the associated segment of putative mRNA.
We found that UAP56 binds to the two known UAP56-binding motifs (UBMs) in mRNP export adapters, such as ALYREF, in distinct ways. The C-UBM binds the previously known site on UAP56’s RecA1 lobe, but, unexpectedly, the N-UBM binds a distinct site on the RecA2 lobe. 4/
19.11.2025 23:21 —
👍 1
🔁 0
💬 1
📌 0

Cartoon representation of the cryo-EM structure of the endogenous human TREX complex (THO+UAP56+ALYREF) bound to an mRNP
THO binds to maturing mRNPs (=mRNA+proteins), directing them into the mRNA export pathway. Yet THO must again dissociate, to allow for subsequent steps in export. How? The connection between mRNP and THO is the DExD-box ATPase UAP56, which binds to UBM motifs in mRNP export adapters (like ALYREF).3/
19.11.2025 23:21 —
👍 1
🔁 0
💬 1
📌 0


















