That's great! Thank you!
27.06.2025 21:47 — 👍 1 🔁 0 💬 0 📌 0That's great! Thank you!
27.06.2025 21:47 — 👍 1 🔁 0 💬 0 📌 0Yes you're right - I think I need to look into our data a bit more and see how this is playing out. Even for the larger clusters, the PC is just a linear combination of the expression values so it would just be a QTL on a weighted sum/difference over more variables
10.06.2025 16:28 — 👍 1 🔁 0 💬 1 📌 0so if something is a PC2 QTL, then it would be pulling expression away from that main axis of correlated variation, and then to a larger difference in expression levels for any given individual.
09.06.2025 22:11 — 👍 1 🔁 0 💬 1 📌 0
Thank you, I think you are right - I've been conceptualizing (at least for a correlated 2 gene cluster) PC1 capturing the shared main axis of variation, and PC2 capturing the spread away from that main axis (so for positively correlated variables I think you're right that would be the difference)
09.06.2025 22:11 — 👍 2 🔁 0 💬 1 📌 0Just brainstorming, but do you think multi-ancestry GWAS/QTLs or comparing between ancestries could help to at least give us different LD confounders and tease apart those relationships? I'm not as familiar with the literature in that space though
09.06.2025 21:58 — 👍 0 🔁 0 💬 0 📌 0
I think establishing a gold-standard set of single-gene effect variants would be difficult - there are just so many strange mechanisms in biology! For instance, NMD of mutated mRNAs can cause up-regulation of paralogs. So even a PTVs can have multi-gene effects www.nature.com/articles/s41...
09.06.2025 21:55 — 👍 2 🔁 0 💬 0 📌 0
I think also the omnigenic model has influenced how I think about this. If one of the regulated genes is a "core gene" then maybe that one is the most important but the other regulated genes still have some impact pmc.ncbi.nlm.nih.gov/articles/PMC...
09.06.2025 21:44 — 👍 1 🔁 0 💬 0 📌 0How to get at this in an unbiased way is further complicated by the fact that its fairly common for paralogs or functionally related genes to be near each other, like those IFNa/b and OAS1/OAS2/OASL loci
09.06.2025 21:39 — 👍 2 🔁 0 💬 0 📌 0I don't, I think it would be really interesting to think of a way to systematically access how often it is one or the other. I like @kauralasoo.bsky.social 's point about comparing putative proxitropic regulatory QTLs with missense or splicing QTLs which are more likely to be single-gene effects.
09.06.2025 21:39 — 👍 1 🔁 0 💬 1 📌 0Awesome! I will DM you
09.06.2025 21:29 — 👍 1 🔁 0 💬 0 📌 0It was a delight to work with you on this Tami! 🫶 hope for more collaboration in the future :)
09.06.2025 17:54 — 👍 1 🔁 0 💬 0 📌 0
This MPRA work from our lab suggests that this multiple-causal variant senario is fairly likely (17% of eQTLs tested) www.science.org/doi/10.1126/...
09.06.2025 17:48 — 👍 3 🔁 0 💬 0 📌 0
That's an interesting point and definitely worth thinking about. I think a case with multiple causal variants in LD with different marginal effects on each of the genes near them would require orthogonal approaches (perhaps MPRAs?) to disentangle.
09.06.2025 17:48 — 👍 2 🔁 0 💬 1 📌 0
www.medrxiv.org/content/10.1... this paper @mykmal.xyz recommended elsewhere in this thread may be of interest to you as a framework to think about co-expression effects
09.06.2025 17:42 — 👍 2 🔁 0 💬 0 📌 0I hadn't seen your paper, thank you for sharing! I was wondering, in your analysis, did you see if cis-eQTLs did worse in the prioritization task when the pQTL gene they attempted to pick shared regulation with nearby genes or overlapped nearby genes?
09.06.2025 17:40 — 👍 1 🔁 0 💬 1 📌 0
Ribeiro et. al. has a nice analysis showing eSNPs shared between multiple eGenes are more indeed pleiotropic. We also see this in our data www.nature.com/articles/s41...
09.06.2025 17:23 — 👍 2 🔁 0 💬 0 📌 0I think that both of these occur. Sometimes maybe a pcQTL effects multiple genes and we need the power boost from looking at all the genes to detect it, but only one gene actually impacts the trait, but I believe sometimes it is a multi-gene or even second order gene co-expression effect
09.06.2025 17:14 — 👍 3 🔁 0 💬 2 📌 0Yes I think you are exactly right! They tend to be weaker effects. We also have some preliminary results not in this preprint indicating that as you increase sample size, you end up discovering many of the loci as eQTLs that were only pcQTLs at the smaller sample size
09.06.2025 17:10 — 👍 3 🔁 0 💬 1 📌 0Very exciting you'll be coming to Stanford, I would love to chat when you're here! Do you think extending COWAS to larger groups of proteins with PCs would be feasible?
09.06.2025 04:09 — 👍 1 🔁 0 💬 1 📌 0Thank you! No worries, I only just made the account so it would have been impossible to find!
08.06.2025 18:30 — 👍 1 🔁 0 💬 1 📌 0If you're interested in a bit more detail about what we did, check out our tweetorial here! bsky.app/profile/itsk...
08.06.2025 18:20 — 👍 0 🔁 0 💬 0 📌 0If you're interested in a bit more detail about what we did, check out our tweetorial here! bsky.app/profile/itsk...
08.06.2025 18:18 — 👍 0 🔁 0 💬 0 📌 0Thank you for the shoutout! I just posted a tweetorial (bsky.app/profile/itsk...) if anyone is interested in a bit more about what we did!
08.06.2025 18:16 — 👍 0 🔁 0 💬 1 📌 0
🧵5/5 Make allelic proxitropy work for you! Even this simple approach substantially increased colocalizations. We’re excited to see future refinements as we move beyond the “one variant, one gene” paradigm to get a more complete picture of the complexities of local gene regulation 🔬🧬✨
08.06.2025 17:38 — 👍 4 🔁 0 💬 1 📌 0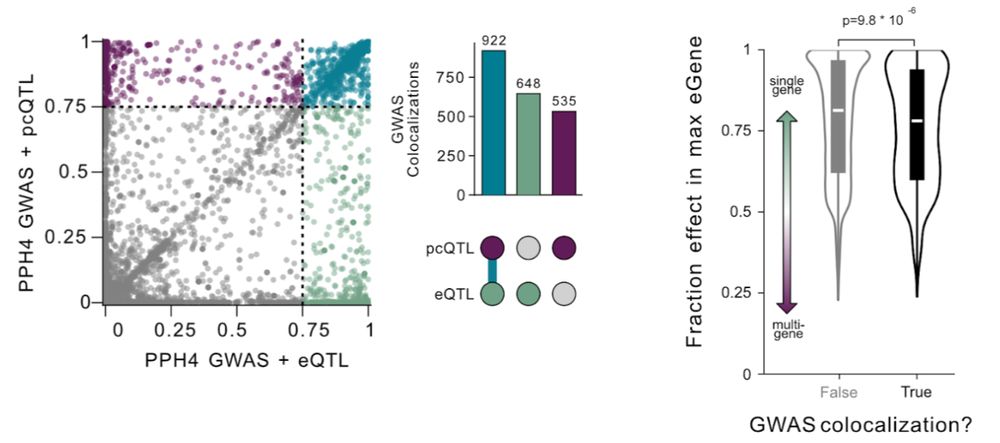
🧵4/5 But are new pcQTLs relevant? We examined associations with GWAS for complex traits and diseases and found over 500 new colocalizations – a 34% increase over single-gene-based analyses! In fact, variants with distributed effects are more likely to be GWAS hits than those affecting just one gene
08.06.2025 17:38 — 👍 5 🔁 2 💬 1 📌 0
🧵3/5 In 13 human tissues (using GTEx data), we identified > 12,000 clusters nearby, co-expressed genes, then mapped pcQTLs, finding ~1,400 pcQTLs/tissue. 27% of pcQTL were missed by single-gene QTL approaches! pcQTLs often are smaller distributed effects that we can detect when combined 🤝
08.06.2025 17:38 — 👍 4 🔁 0 💬 1 📌 0
🧵2/5 We use principal components (PCs) to summarize nearby genes and map pcQTLs. How do pcQTLs work? 🔍 For correlated genes (A, B, C), standard methods test genotype associations independently along each axis. pcQTLs test for association on a PC axis instead to capture shared regulatory effects 🔗
08.06.2025 17:38 — 👍 5 🔁 1 💬 1 📌 0
Standard methods are equivalent to a flashlight, looking at each gene independently. We combine signals from multiple genes, turning a floodlight onto the genome.
Excited to share my first PhD paper in the @sbmontgom.bsky.social lab with @tamigj.bsky.social (www.biorxiv.org/content/10.1...)! Standard QTL methods treat each gene independently. But what if a single variant regulates multiple nearby genes at once - what we call “allelic proxitropy”? 🧵 ⬇️
08.06.2025 17:38 — 👍 91 🔁 33 💬 6 📌 4wow looks like a sick trail behind there
28.03.2025 18:26 — 👍 0 🔁 0 💬 0 📌 0