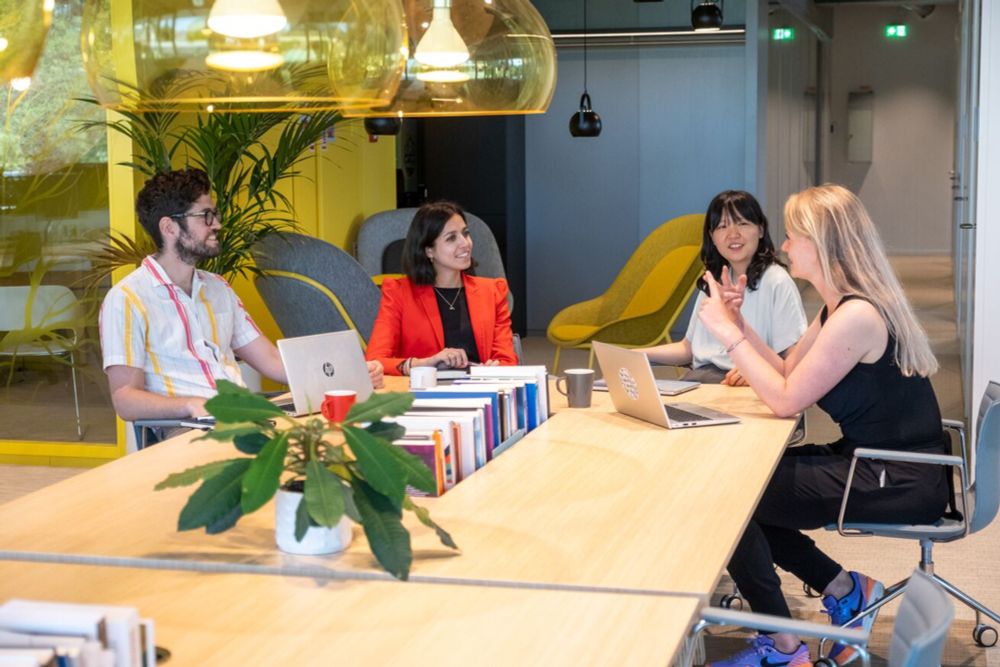

On both synthetic and realistic data LOAD
🏎️is more computationally efficient than global methods, performing close to local methods,
💎recovers high-quality, statistically efficient adjustment sets,
🔮thus enables reliable causal effect estimation even at scale
7/8
23.10.2025 15:22 —
👍 0
🔁 0
💬 1
📌 0
LOAD follows 5 steps:
➡Learn causal relations between targets
✅Test identifiability of the effect
🐣Find explicit descendants of treatment
🧩Find mediators
🎯Collect optimal adjustment set
For unidentifiable effects, LOAD exits early and returns locally valid adjustments
6/8
23.10.2025 15:22 —
👍 0
🔁 0
💬 1
📌 0

To do this, we develop a sufficient and necessary test for the identifiability of the causal effect of a treatment on an outcome using only local information around the treatment and its siblings, no matter how far the treatment and the outcome are in the causal graph 🔭
5/8
23.10.2025 15:22 —
👍 0
🔁 0
💬 1
📌 0
🎯 Local Optimal Adjustments Discovery (LOAD) does exactly that! It provably finds the same ✨optimal adjustments✨ as global methods, but using much more ⚡computationally efficient⚡ local causal discovery around variables
4/8
23.10.2025 15:22 —
👍 0
🔁 0
💬 1
📌 0

🌐 Global discovery methods can find optimal adjustment sets, but at a huge computational cost.
📍 Local discovery methods are fast, but can only find sub-optimal adjustment sets.
Can we get the best of both worlds and find optimal adjustment sets from local information?
3/8
23.10.2025 15:21 —
👍 1
🔁 0
💬 1
📌 0

While all valid adjustment sets enable unbiased estimation of causal effects, using the optimal adjustment set in terms of asymptotic variance is crucial for reliable causal effect estimation!⚠️
But how to find the optimal adjustment set if the causal graph is not available?
2/8
23.10.2025 15:21 —
👍 0
🔁 0
💬 1
📌 0
Estimating causal effects efficiently doesn’t have to mean discovering the entire causal graph! Now you can find the optimal adjustment from only local information using LOAD!
📜 Preprint: arxiv.org/abs/2510.14582
👾 Code: github.com/Matyasch/load
🧵 1/8
23.10.2025 15:21 —
👍 13
🔁 2
💬 1
📌 0
Carlos Cinelli, Avi Feller, Guido Imbens, Edward Kennedy, Sara Magliacane, Jose Zubizarreta
Challenges in Statistics: A Dozen Challenges in Causality and Causal Inference
https://arxiv.org/abs/2508.17099
26.08.2025 05:56 —
👍 10
🔁 4
💬 0
📌 0
Are you interested in improving the #interpretability, #robustness and #safety of current AI systems with #causality and #RL?
Apply to our PhD position in Amsterdam 🚲🌷🇳🇱
Deadline: June 15
26.05.2025 08:32 —
👍 11
🔁 6
💬 1
📌 2

CAR
Causal Abstractions and Representations��Workshop @ UAI 2025�July 25th 2025, Rio de Janeiro 🇧🇷
🔥 Got a great work on causal representation learning, abstraction, high-dimensional discovery, or other hot topics in causality?
🇧🇷 Don’t miss your chance to present in Rio at the CAR Workshop at #UAI2025!
⏰ Deadline is in 1 week – May 26!
🌐 sites.google.com/view/car-25/
19.05.2025 19:52 —
👍 15
🔁 7
💬 0
📌 1


A little over a week ago, I had the chance to attend #AISTATS and present our poster on SNAP (matyasch.github.io/snap)! Three days of brilliant invited talks and a stream of fascinating papers left me with a much longer reading list about ideas to explore.
13.05.2025 10:25 —
👍 2
🔁 1
💬 0
📌 0
Just arrived in Phuket for #AISTATS2025. Can't wait to present our poster (in tube) about SNAP 🫰 on day 2, Sunday! Come check it out and let's chat about scalable causal discovery!
02.05.2025 11:47 —
👍 9
🔁 2
💬 1
📌 0
⏰ Don't miss Mátyás talk today at 3PM!
🎥 See you online meet.google.com/cqt-ufji-xfz
🤌 ...or live in the CS Department of the University of Pisa!
07.03.2025 11:15 —
👍 7
🔁 3
💬 0
📌 0
A few weeks ago, I presented SNAP at the wonderful #Bellairs Workshop on Causality in Barbados🐢
This Friday 🫰meets 🤌 as I will get to present SNAP again at the kick-off of the newest season of @causalclub.di.unipi.it! Check out this, and their other amazing upcoming talks at causalclub.di.unipi.it
04.03.2025 14:03 —
👍 10
🔁 1
💬 0
📌 1

Sequential Non-Ancestor Pruning | Matyas Schubert
10/10 SNAP is joint work with a fantastic team of Tom Claassen and @smaglia.bsky.social. Visit our project page on matyasch.github.io/snap/, run SNAP using our publicly available code at github.com/matyasch/snap, and visit to our poster at #aistats2025! 🏖️
13.02.2025 14:03 —
👍 3
🔁 0
💬 0
📌 0
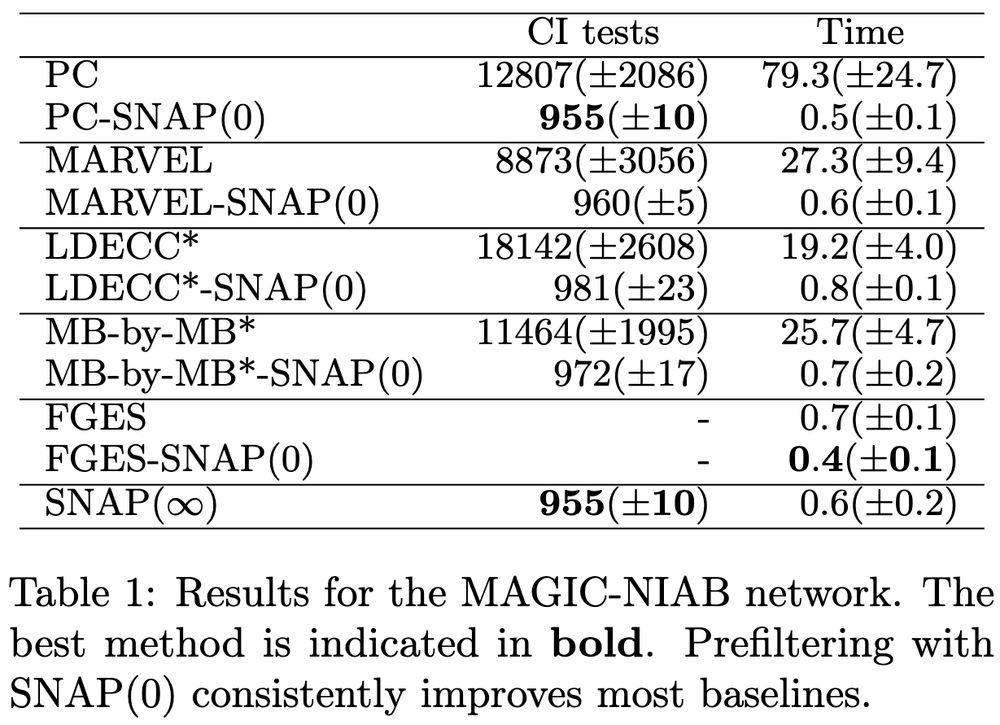
9/10 We also evaluate SNAP on semi-synthetic settings including data generated from the MAGIC-NIAB network, which captures genetic effects and phenotypic interactions 🧬 We see that SNAP greatly reduces the number of CI tests and execution time compared to most baselines.
13.02.2025 14:03 —
👍 3
🔁 0
💬 1
📌 0

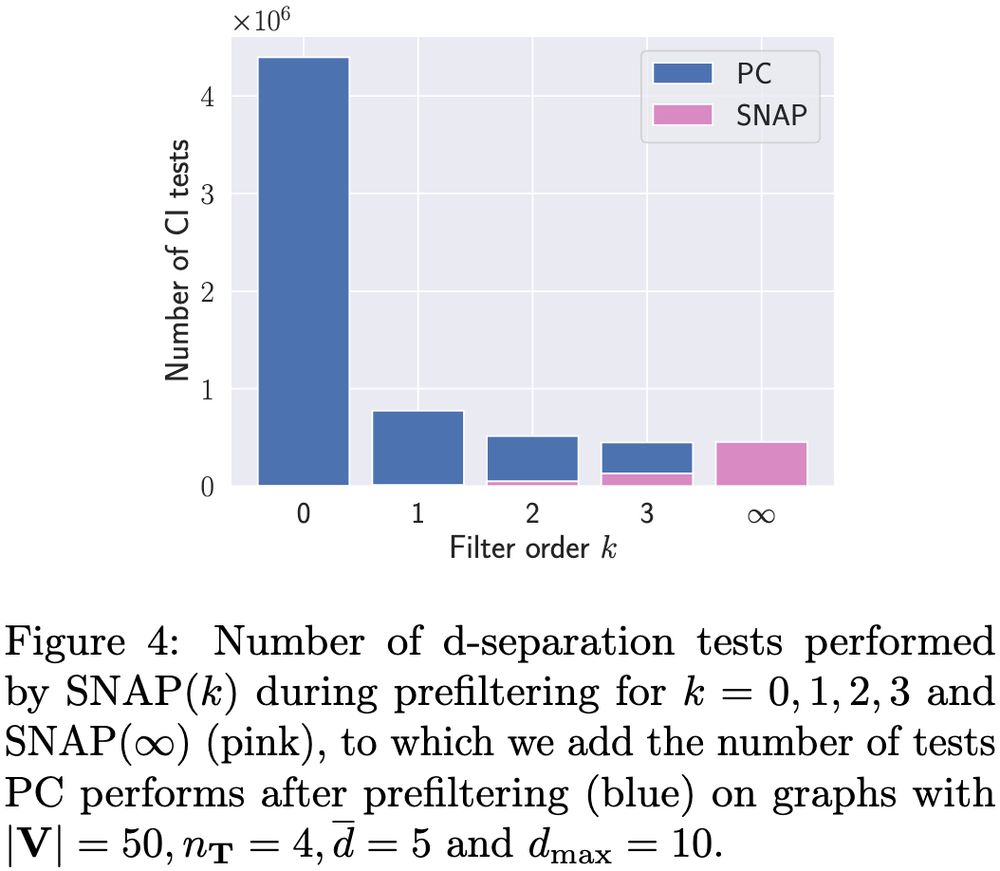
8/10 Many non-ancestors are already identified by marginal tests, enabling prefiltering with SNAP(0) to significantly speed up computation time. Increasing the number of prefiltering iterations k further reduces the number of CI tests needed, especially in dense graphs 🧶
13.02.2025 14:02 —
👍 1
🔁 0
💬 1
📌 0
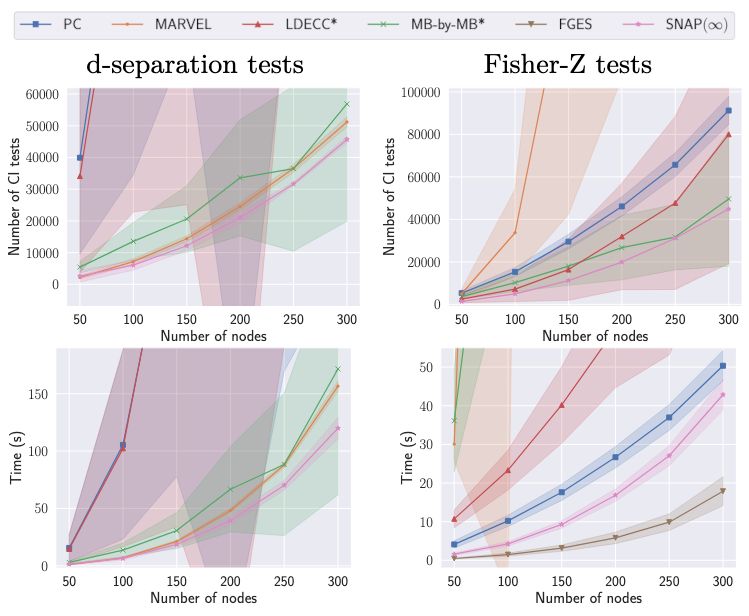

7/10 SNAP(∞) consistently ranks among the best in the number of CI tests and computation time across all domains, while maintaining a comparable intervention distance. In contrast, other methods vary in performance depending on the setting 🚀
13.02.2025 14:02 —
👍 2
🔁 1
💬 1
📌 0
6/10 We can also run SNAP until completion, to obtain a stand-alone causal discovery algorithm, called SNAP(∞). SNAP(∞) is sound and complete over the possible ancestors of targets ✅ Thus, unlike previous work on local causal discovery, it finds efficient adjustment sets.
13.02.2025 14:01 —
👍 1
🔁 0
💬 1
📌 0
5/10 SNAP is straightforward to combine with readily available causal discovery algorithms 🧩 We can simply stop it at any maximum iteration k and run another algorithm on the remaining variables. We refer to this approach as prefiltering with SNAP(k).
13.02.2025 14:01 —
👍 1
🔁 0
💬 1
📌 0
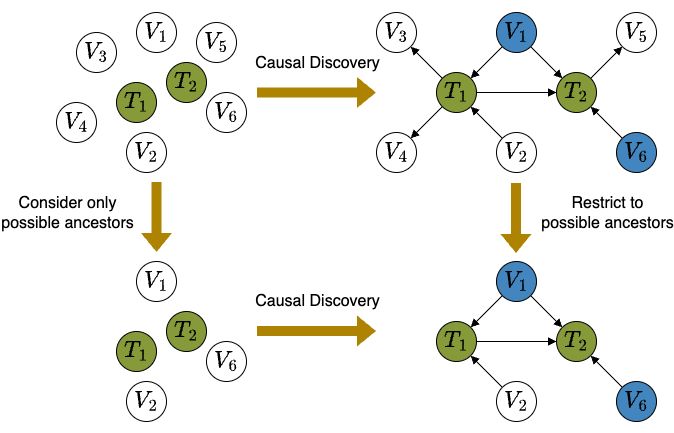
4/10 To solve this task, we show that only possible ancestors of the targets are required to identify their causal relationships and efficient adjustment sets💡 Driven by this, we propose SNAP to progressively prune non-ancestors, leading to much fewer higher order CI tests.
13.02.2025 14:00 —
👍 3
🔁 1
💬 1
📌 0
3/10 We formalize this as the task of “targeted causal effect estimation with an unknown graph”, which focuses on identifying causal effects between a small set of target variables in a ✨computationally and statistically efficient way✨
13.02.2025 14:00 —
👍 1
🔁 0
💬 1
📌 0
2/10 Discovering causal relations can help us estimate causal effects, but it is expensive 📈 If we are only interested in estimating the causal effects between a few target variables, can we instead only discover a subgraph that includes these targets and their adjustment sets?
13.02.2025 14:00 —
👍 1
🔁 0
💬 1
📌 0
Do you want to estimate causal effects for a small set of target variables without knowing the causal graph, but discovering it takes too long? Now you can get adjustment sets in a SNAP🫰accepted at #aistats2025!
📜 arxiv.org/abs/2502.07857
🧩 matyasch.github.io/snap/
🧵 1/10
13.02.2025 13:59 —
👍 19
🔁 5
💬 1
📌 4

Professor Imbens also had a mentoring session with our PhD students actively working on causality, discussing their ideas and the potential impact of their applications! 👨🔬👩🔬
@matyasch.bsky.social @roelhulsman.bsky.social @rmassidda.it @danruxu.bsky.social 🔥
13.12.2024 08:47 —
👍 13
🔁 4
💬 1
📌 1
Congrats to @smaglia.bsky.social for now being an ELLIS Scholar! 🤩🥳🎉
09.12.2024 16:02 —
👍 19
🔁 3
💬 0
📌 0
if you ever wondered how to concisely represent causal models, don’t miss my talk on causal abstraction tomorrow at @ellisamsterdam.bsky.social
03.12.2024 22:43 —
👍 13
🔁 4
💬 2
📌 0




















