
New paper alert: Paralog interference contributes to the preservation of genetic redundancy www.cell.com/current-biol...
28.02.2026 02:00 — 👍 45 🔁 28 💬 0 📌 0
New paper alert: Paralog interference contributes to the preservation of genetic redundancy www.cell.com/current-biol...
28.02.2026 02:00 — 👍 45 🔁 28 💬 0 📌 0Now in Nature Comms w/ @ritatewari.bsky.social, Pushkar Sharma & @ryanase.bsky.social (thanks!). Aurora kinases fascinate me: single ancestor - parallel duplications in eukaryotes - paralogs with distinct functions. ARK1 is the CPC Aurora in the malaria parasite. rdcu.be/e5NRT #plasmodium #mitosis
26.02.2026 13:02 — 👍 13 🔁 4 💬 0 📌 0We're on a roll here. Check out this cool paper by @scienceleah.bsky.social et al. on not one, but two types of sperm (!) in the silk worm Bombyx mori. Happy to have contributed. #meiosis4ever
26.02.2026 12:57 — 👍 7 🔁 3 💬 0 📌 0As a cell biology lab, we acknowledge the decades-long impressive efforts to uncover evolutionary relationships using advanced phylogenomics methods. These approaches undergo continuous improvements that lead to adjustments of data interpretation, as is the case in every scientific field. (1/3)
23.02.2026 15:11 — 👍 6 🔁 2 💬 1 📌 0Before he passed away in 2021, Tom Cavalier-Smith had drafted parts of his autobiography. It's now 'published' because Gáspár Jékely put a lot of effort in! Please enjoy and share: doi.org/10.5281/zeno...
23.02.2026 14:37 — 👍 28 🔁 13 💬 1 📌 1Hamer + Spijker. Tijd voor verandering van de PhD stelsel op Nederlandse Universiteiten. Lees dit opiniestuk van @erikvansebille.bsky.social in het Digitaal Universiteitsblad van @utrechtuniversity.bsky.social
16.02.2026 10:10 — 👍 6 🔁 3 💬 2 📌 0Perpetually in awe of the astonishing creative capacity of retrotransposons in eukaryotic cell biology — makes you wonder what remarkable, still-hidden origin stories centromeres across the tree of life carry.
18.02.2026 19:49 — 👍 12 🔁 2 💬 0 📌 0
A 🆕 #Ichthyosporea story
Led by@margaridaaraujo.bsky.social, in Co. @gautamdey.bsky.social & @hiralshah.bsky.social+🔥team across @embl.org & @sciencesunige.bsky.social
We finally have a look at what #microtubules are doing during #cellularization in #Sphaeroforma
▶️ tinyurl.com/MTSARC
#ProtistsOnSky

Recently, a Hypothesis was posed in @jcellsci.bsky.social in which the root of eukaryotes was placed between kinetoplastids and all other eukaryotes. From this, it was implied that LECA did not have a kinetochore. We argue this is highly unlikely. A 🧵(1/12)
Read our reply here: tinyurl.com/n87myhpr
Direct binding of chromosome axis and cohesin complexes underlies meiotic chromosome architecture in fungi and plants https://www.biorxiv.org/content/10.64898/2026.01.19.700424v1
20.01.2026 17:46 — 👍 1 🔁 1 💬 0 📌 0Congrats on this achievement (despite the lockdown start..) - may there be many more years to come.
07.01.2026 10:34 — 👍 1 🔁 0 💬 0 📌 0
Another #notTHECover unfortunately.
But this gorgeous, Tron-like vibe, drawn by the amazing @munafomarzia.bsky.social for our recent #ExM work with @gautamdey.bsky.social & @centriolelab.bsky.social will still be printed out in the lab.
Read here: www.cell.com/cell/fulltex...
Some light reading for the holidays :). Let us know what you think.
22.12.2025 18:55 — 👍 15 🔁 4 💬 0 📌 0
New preprint🚨
Imagine (re)designing a protein via inverse folding. AF2 predicts the designed sequence to a structure with pLDDT 94 & you get 1.8 Å RMSD to the input. Perfect design?
What if I told u that the structure has 4 solvent-exposed Trp and 3 Pro where a Gly should be?
Why to be wary🧵👇
#EvoChromo ciliate kinetochores @maxraas.bsky.social @kopslab.bsky.social . Previous work w\ @jvhooff.bsky.social : tinyurl.com/y5h4ehnt showed a lack of Tetrahymena kinetochore complexity. Replacement, simplification or missed homologies? All of the above? - check out our pre-print tiny.cc/hp8w001
10.12.2025 12:24 — 👍 11 🔁 0 💬 0 📌 0#protistsonsky
02.12.2025 19:13 — 👍 1 🔁 0 💬 0 📌 0
Our story on the kinetochore composition of the ciliate Tetrahymena thermophila is out now on bioRxiv! We find surprisingly many orthologs of conventional kinetochore components, but also components that have very different evolutionary origins. A 🧵 (1/11)
Check it out here: tinyurl.com/4ectm9x4

How do new centromeres evolve while staying compatible with the division machinery?
Discover it in our new Nature paper! We show centromeres transition gradually via a mix of drift, selection, and sex, reaching new states that still work with the kinetochore.
👉 doi.org/10.1038/s41586-025-09779-1

Our new preprint describes Plasmodium NEK4 as a key regulator coupling meiotic initiation and morphogenesis—a critical step for malaria transmission. This involves MTOC-driven nuclear movement strikingly analogous to "horsetail" movement in fission yeast.
www.biorxiv.org/content/10.1...
New work on ARK1 (AURKB?) in Plasmodium (malaria parasite) with @ritatewari.bsky.social and Pushkar Sharma. CPC in Plasmodium is (ofcourse..) different compared to conventional models: two INCENPs, no borealin/survivin - mainly at spindle microtubules/MTOCs. Special thanks to @ryanase.bsky.social!
31.08.2025 12:26 — 👍 7 🔁 2 💬 0 📌 1
Pleased to share our new preprint: "Plasmodium ARK1 regulates spindle formation during atypical mitosis and forms a divergent chromosomal passenger complex". Many thanks to our collaborators.
@ritatewari.bsky.social @eelcotromer.bsky.social @davidguttery.bsky.social
www.biorxiv.org/content/10.1...
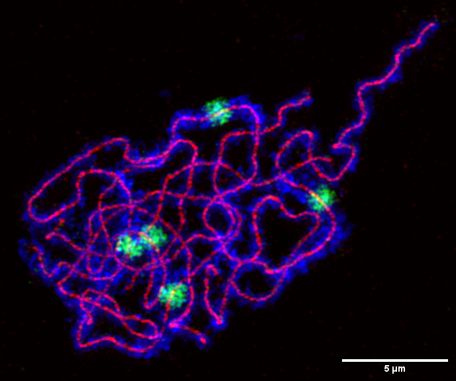
Happy to share the first story from my postdoc in @schnittger-lab.bsky.social
We studied TITAN9, a Csm1-like protein in Arabidopsis.
Even though I now work with soil, chromosome segregation and meiosis still have a special place in my heart.
#meiosis4eva #meiosis #chromosomesegregation

So excited to share this preprint from the Rosin lab in collaboration with the Hawley lab and @eelcotromer.bsky.social on moth spermatogenesis! We investigate the meiotic errors that occur during the formation of apyrene sperm (that have no DNA) in silkworms!
www.biorxiv.org/content/10.1...
Postdoc position currently available in our group studying meiosis in arabidopsis and wheat:
www.jic.ac.uk/vacancies/po...

We are growing! A new @zeiss-microscopy.bsky.social widefield tailored for ExM- supported by @moorefound.bsky.social is settling! Another one is on its way at @gautamdey.bsky.social Lab !
These two will become the powerhorses for high-througput imaging of the ExM Atlas of #Microbial Eukaryote !
Happy to see our work published and glad to have contributed together with @maxraas.bsky.social ! Looking forward to all the projects that will come out of this work!
23.06.2025 11:18 — 👍 19 🔁 6 💬 0 📌 0ps. will you bring back taxonomy selection (at least prokaryote/viral/eukaryote?)
20.05.2025 12:40 — 👍 0 🔁 0 💬 0 📌 0much appreciated!
20.05.2025 12:31 — 👍 0 🔁 0 💬 1 📌 0great to see the new version of the webserver - will you bring back jackhmmer? that was such a useful tool for quick checks :)
20.05.2025 11:51 — 👍 1 🔁 0 💬 1 📌 0
It’s out! The evolutionary origins of yeast point centromeres uncovered!
“Ancient co-option of LTR retrotransposons as yeast centromeres”
www.biorxiv.org/content/10.1...