Client Challenge
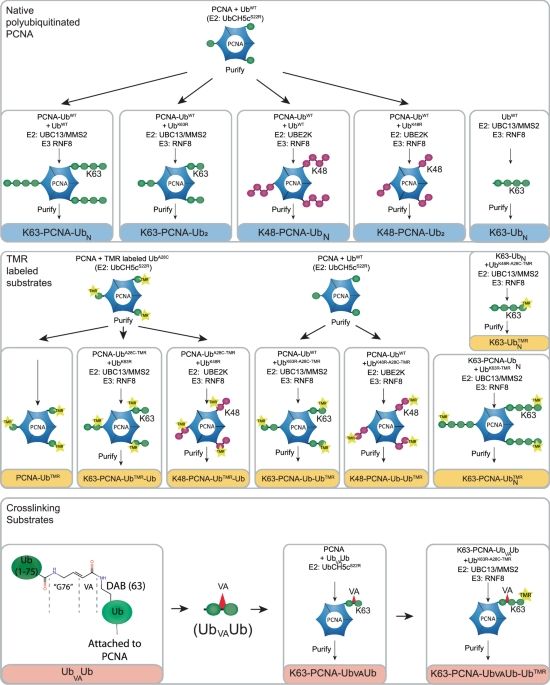
We are excited to share that our latest work from the lab aimed at understanding how the tRNA nuclease SLFN11 is activated in response to DNA damage and replication stress has just been published in @natcellbio.nature.com!
Open access link: www.nature.com/articles/s41...
(1/9)
09.01.2026 14:42 —
👍 36
🔁 12
💬 1
📌 2
Likewise! Happy holidays.
23.12.2025 09:05 —
👍 1
🔁 0
💬 1
📌 0

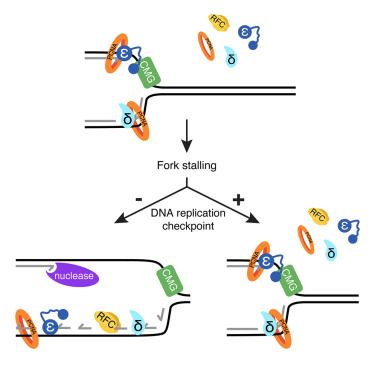
a model for TIMELESS positions at the active fork: one complex sits at the leading edge of the fork, while the other one is on the lagging strand ssDNA/RPA, between the Okazaki fragments
Ever wondered how TIMELESS/TIPIN can sit at the leading edge of the replication fork regulating its speed, but also bind polymerases and ssDNA/RPA in replication stress? We have an idea how it might work! Check out our new preprint, any feedback is highly appreciated!
www.biorxiv.org/content/10.6...
18.12.2025 15:03 —
👍 24
🔁 11
💬 0
📌 0

How DNA secondary structures drive replication fork instability
DNA secondary structures, such as hairpins, cruciforms, triplexes, G-quadruplexes and iMotifs, are common, dynamic features that replication forks rou…
Our @costerlab.bsky.social review is out!
It’s a great read on the impact of DNA secondary structures on eukaryotic replication fork progression - a totally unbiased opinion, of course!
www.sciencedirect.com/science/arti...
@adityasethi.bsky.social @billiedelpino.bsky.social
10.12.2025 11:53 —
👍 16
🔁 8
💬 0
📌 1

Among the anti-recombinases, FIGNL1 rules them all. So much that inactivating it brings BRCA2-deficient cells to life. Who is responsible for RAD51 loading without BRCA2/FIGNL1, check out the paper to find out! Great collaboration with @raychaudhurilab.bsky.social
www.science.org/doi/10.1126/...
30.10.2025 20:41 —
👍 53
🔁 22
💬 6
📌 1
Happy to have contributed to this interesting study from penengolab.bsky.social! another interesting example of how chromatin modifications and their readers orchestrate replication fork plasticity and protection 👏👇
30.10.2025 13:19 —
👍 11
🔁 4
💬 0
📌 0

What specific targets & roles are there for all distinct polyubiquitin chain types?
Niels Mailand, Robert Shearer et al profile a panel of #ubiquitin replacement cell lines, implicating K29 chains in chromatin regulation via SUV39H1 destabilization
www.embopress.org/doi/full/10....
27.10.2025 11:51 —
👍 12
🔁 6
💬 0
📌 0

See our paper “Mechanism of trinucleotide repeat expansion by MutSβ-MutLγ and contraction by FAN1”. Using biochemistry, we show how DNA incisions by the MutLγ nuclease can lead to expansions, and how expansion is prevented by FAN1. Well done Issam and Valentina!
www.nature.com/articles/s41...
27.10.2025 13:12 —
👍 39
🔁 12
💬 1
📌 1
Well deserved!
01.08.2025 09:33 —
👍 1
🔁 0
💬 1
📌 0
Well done Ivan. Beautiful work!
11.07.2025 18:16 —
👍 0
🔁 0
💬 0
📌 0
Home - Birmingham Centre for Genome Biology 2025
BCGB 2025, Birmingham Centre for Genome Biology Conference 2025
Birmingham Centre for Genome Biology 2025 Anniversary Conference, 11-12 September! This symposium covers DDR and gene regulation. Register here:
uobevents.eventsair.com/birmingham-c...
24.06.2025 16:20 —
👍 8
🔁 7
💬 0
📌 0
Postdoctoral Research Associate in the Blackford Group
A position as Postdoctoral Research Associate is available at the Department of Cellular and Molecular Medicine, in the group headed by Dr. Andrew Blackford. Th
We’re #hiring, please share! A fully funded #postdoc position is available in my lab to work on DNA damage response mechanisms.
Please get in touch with your CV if you are interested; closing date: 27th June 2025.
More details here: candidate.hr-manager.net/ApplicationI...
29.05.2025 11:26 —
👍 23
🔁 35
💬 0
📌 0
What a legend. Such a shame.
12.06.2025 16:16 —
👍 0
🔁 0
💬 0
📌 0
We are looking to recruit three posts at the @crick.ac.uk in structural biology, cell biology, and pharmacology to join a 20+ strong multidisciplinary team focused on delivering the first precision medicines for the treatment of ALT cancers. Interested? Please check out the advertised jobs below:
27.05.2025 21:52 —
👍 3
🔁 4
💬 1
📌 0

Parp1 deletion rescues cerebellar hypotrophy in xrcc1 mutant zebrafish - Scientific Reports
Scientific Reports - Parp1 deletion rescues cerebellar hypotrophy in xrcc1 mutant zebrafish
Loss of XRCC1 disrupts cerebellar development in zebrafish due to toxic PARP1 accumulation. Strikingly, parp1 knockdown rescues the XRCC1 phenotype, supporting PARP1 inhibition as a potential therapy in recessive XRCC1-related neurodegenerative disorders with ataxia. www.nature.com/articles/s41...
18.05.2025 16:23 —
👍 10
🔁 5
💬 0
📌 0