Always good to start the year with a new preprint. We charted DNA methylation in Salmonella enterica: www.biorxiv.org/content/10.64898/2026.01.27.702048v1.
30.01.2026 11:53 — 👍 5 🔁 3 💬 0 📌 0Always good to start the year with a new preprint. We charted DNA methylation in Salmonella enterica: www.biorxiv.org/content/10.64898/2026.01.27.702048v1.
30.01.2026 11:53 — 👍 5 🔁 3 💬 0 📌 0🔊 Interested in doing a PhD on RNA-binding proteins in an abundant microbiota species and becoming a member of the German Priority Programme “Illuminating Gene Functions in the Human Gut Microbiome”? Apply here: www.uni-wuerzburg.de/karriere/sin...
01.12.2025 09:20 — 👍 7 🔁 15 💬 0 📌 1
This work is now out in Microbiology! www.microbiologyresearch.org/content/jour.... Congrats to first author Aalap Mogre!
Publish with the Microbiology Society! A membership charity and not-for-profit publisher: @microbiologysociety.org !
Please RT:
We have an opening for a junior group leader position in „Phage Biology & Biotechnology“.
www.fz-juelich.de/de/karriere/...
Interested candidates are encouraged to contact me via email for further details.
@spp2330.bsky.social; @mibinet.bsky.social

Happy International Microorganism Day! 🎉
Today, we are celebrating the tiny powerhouses that have a massive impact on our world. From the bacteria in our gut to the fungi that give us antibiotics, these incredible organisms are essential to life.
Let's make some noise for the microscopic!

Poster quoting from a reviewer saying "I have been supported by the Microbiology Society so much, so I will always support them."
Publish with us and invest in the research community. Supported by income from our journals, the Microbiology Society awards hundreds of grants every year, providing opportunities and experience for microbiologists at all career levels.
29.08.2025 15:05 — 👍 5 🔁 3 💬 2 📌 0Cool stuff! Many congrats Lauren and Team!
11.08.2025 11:55 — 👍 2 🔁 0 💬 1 📌 0New preprint! We characterise the small, regulatory RNA Arp which controls DNA uptake and twitching motilty in A. baumannii. Led by our Dr Fergal Hamrock @hamrockfergal.bsky.social and in collaboration with @westermannlab.bsky.social and Mike Gebhardt's lab!
21.07.2025 14:56 — 👍 3 🔁 6 💬 0 📌 1New preprint! We characterise the small, regulatory RNA Arp which controls DNA uptake and twitching motilty in A. baumannii. Led by our Dr Fergal Hamrock @hamrockfergal.bsky.social and in collaboration with @westermannlab.bsky.social and Mike Gebhardt's lab!
21.07.2025 14:56 — 👍 3 🔁 6 💬 0 📌 1Woohoo! Many, many congratulations Charline! Well deserved!🥳
21.07.2025 14:50 — 👍 0 🔁 0 💬 0 📌 0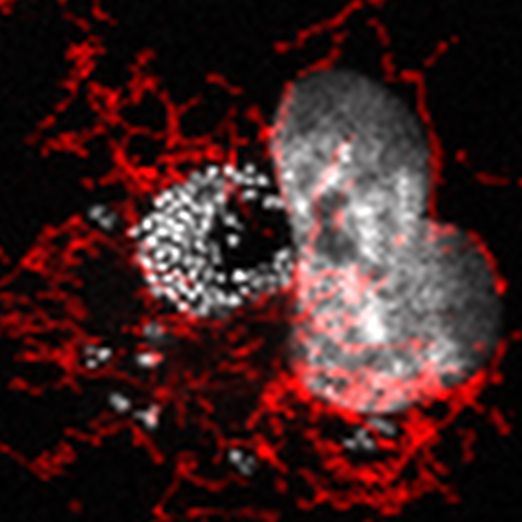
Human endothelial cell infected with Acinetobacter baumannii strain ABC141. Bacteria are visible with Dapi in white along with the nucleus of the cell. Bacteria are mostly organized in a large cluster with a few scattered ones. Mitochondria are visible in red.
Charline studied how some clinical strains of Acinetobacter baumannii create a transient intracellular niche before infecting neighboring cells.
Great collaboration with @kroegerlab.bsky.social
If you are interested, check out her preprint!
www.biorxiv.org/content/10.1...
Great to see that our pgtE story continues! Congrats to the cool work!
18.07.2025 16:00 — 👍 1 🔁 0 💬 0 📌 0Latest preprint from our lab describing a new 2-plasmid system to investigate sRNAs-target molecule interactions in A. baumannii.
14.07.2025 07:26 — 👍 8 🔁 3 💬 0 📌 1Transposon-directed insertion-site sequencing (TraDIS) analysis of Enterococcus faecium using nanopore sequencing and a WebAssembly analysis platform https://www.biorxiv.org/content/10.1101/2025.03.03.641140v1
05.03.2025 03:20 — 👍 5 🔁 1 💬 0 📌 0
Our lab contributed to this including @hamrockfergal.bsky.social : CsrA-mediated regulation of a virulence switch in Acinetobacter baumannii. Great work led by Phil Rather's group.
journals.asm.org/doi/10.1128/...
A work that started 10y ago with a simple experiment with a curious result. Great to see it finally published. Massive effort by @nic-ler.bsky.social and Andrew Cameron! Happy to have contributed to this!
24.02.2025 11:21 — 👍 7 🔁 3 💬 0 📌 0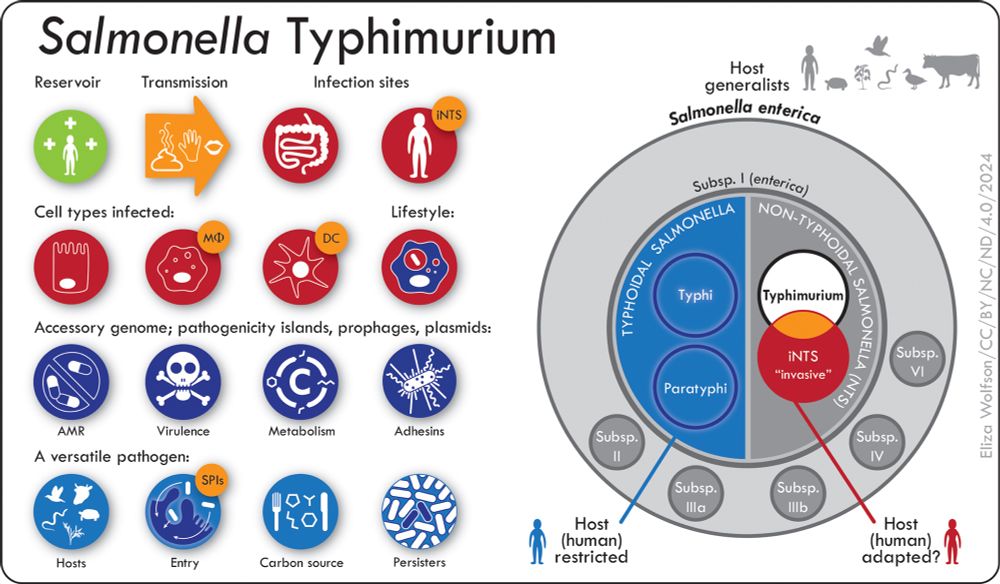
The taxonomy & lifestyle of Salmonella Typhimurium
www.microbiologyresearch.org/content/jour...
23.01.2025 22:56 — 👍 22 🔁 9 💬 0 📌 0Hi Matt. Could you add me please? Thank you :-)
17.01.2025 11:24 — 👍 1 🔁 0 💬 1 📌 0
Delighted to share our work led by IRC GoI PG Scholar Fergal Hamrock:
Global analysis of the RNA–RNA interactome in Acinetobacter baumannii AB5075 uncovers a small regulatory RNA repressing the virulence-related outer membrane protein CarO.
academic.oup.com/nar/advance-...