 06.02.2026 06:52 —
👍 0
🔁 0
💬 0
📌 0
06.02.2026 06:52 —
👍 0
🔁 0
💬 0
📌 0

Pleased to be a small part of this collaboration with the @labconti.bsky.social now published at @narjournal.bsky.social.
Here we combine structure prediction, biochem, cryo-EM, and IP-based approaches to explore connections between RNA quality control to transcription termination complexes.
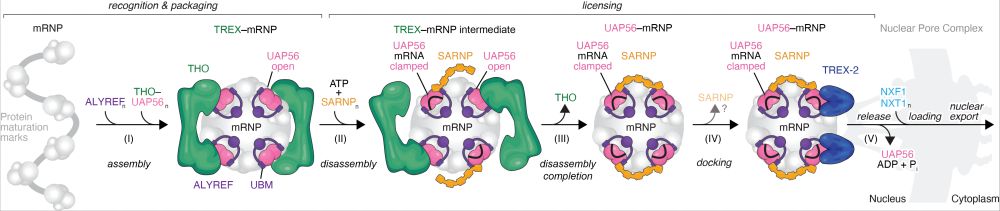
Messenger RNA is made in the nucleus before it is exported to the cytoplasm for translation. But how are only correctly made mRNAs chosen and remodeled in the nucleus for export?
Our new paper investigates the nuclear events leading to human mRNA export. www.nature.com/articles/s41.... (1/4)
RX Bandits - ...and the Battle Begun
21.10.2025 09:10 — 👍 0 🔁 0 💬 0 📌 0
Thrilled to share that I’ll be joining @imbmainz.bsky.social in February 2026 to start my own group!
We will explore new mechanisms in eukaryotic gene expression, leveraging ‘evolutionary play’ to uncover how regulation, repurposing, and hijacking shape RNA biology.
PhD positions available!

How do cells sort which RNAs to keep or destroy? New preprint from THJ, Brenneke and Plaschka labs shows that export and decay machineries (TREX2/PAXT) both recognise UAP56-bound RNAs. Whether they’re exported or degraded depends on where in the nucleus this happens.
www.biorxiv.org/content/10.1...

Registration and abstract submission is open for the 10th annual Danish RNA Society meeting 🇩🇰 - this year in Aarhus!
Deadline is October 1st
danish-rna-society.nemtilmeld.dk/8/
Postdoc positions available in the THJ group in Aarhus, Denmark 🇩🇰
Please share around 🤝

Happy to share our latest collaboration with the Cusack and Kadlec groups—now published in @natsmb.nature.com
Combining cryo-EM, structure-guided mutants & proximity labeling, we show how PHAX, CBC, CRM1 & Ran‑GTP assemble the snRNA export complex
Article: rdcu.be/euC4U
News & Views: rdcu.be/euC5l

Postdoc positions available in the Heick Jensen lab in sunny Aarhus. Sorting of good and bad RNAs in mammalian nuclei. Please get in touch for further information and/or use the following link to apply:
mbg.au.dk/en/news-and-...
Spotlight: shorturl.at/E6je8
Article: shorturl.at/cJsDG

We put together a spotlight for @cp-cellchembiol.bsky.social highlighting the recent work from @baluapuri.bsky.social and @adelmanlab.bsky.social published in @cp-cell.bsky.social.
Here they show how a breakdown in nuclear RNA regulation sends ripples of stress throughout the cell.

Happy to have played a small part in this new preprint on DGCR8, retrotransposons, and IFN regulation. Nice collaboration with @saramaciasrna.bsky.social @herassr.bsky.social
www.biorxiv.org/content/10.1...
I've been wanting to try something like this. Awesome to see that it works 👍
28.04.2025 13:27 — 👍 1 🔁 0 💬 1 📌 0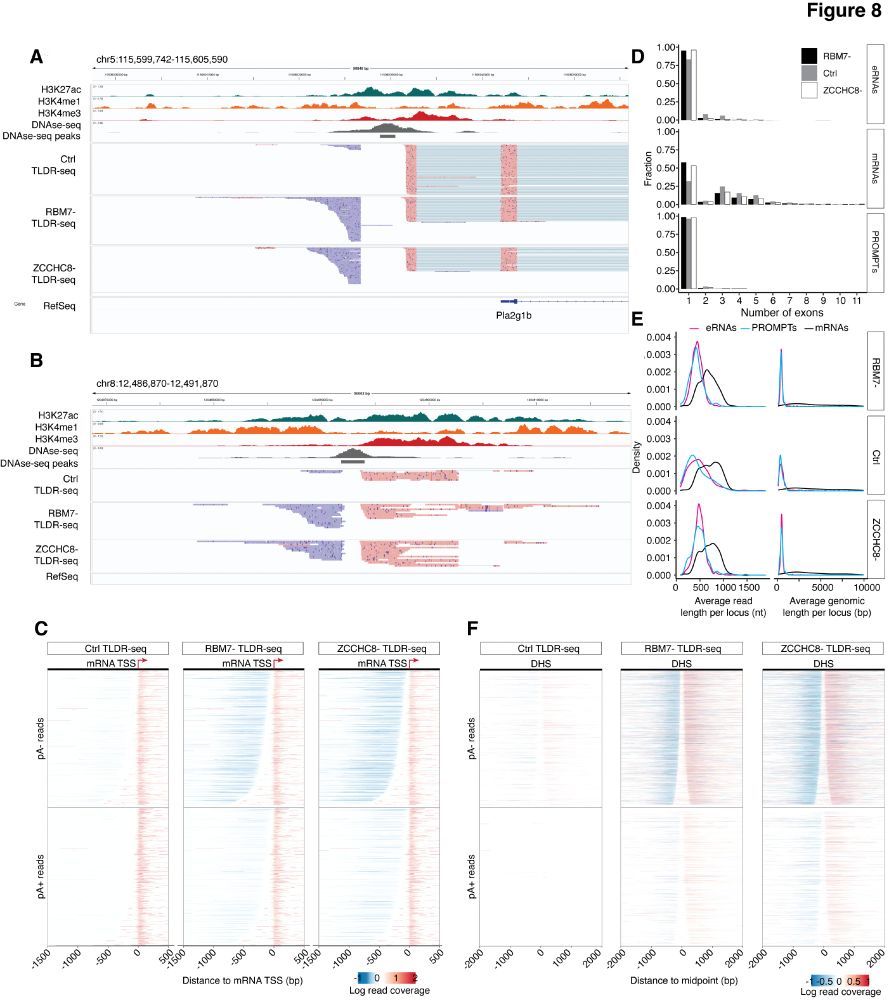
Very proud to share this work just out in NAR: spearheaded by @jamieauxillos.bsky.social and Arnauld Stigliani: TLDR-seq, a method for 5’ to 3’ end long-read sequencing of capped RNAs regardless of 3’ end polyadenylation, based on the @nanoporetech.com platform. (1/4) tinyurl.com/3c2ksdmr
07.04.2025 08:15 — 👍 28 🔁 11 💬 1 📌 1PhD position vacant in the lab. Please spread the news. Study topic: Mammalian nuclear RNA production and turnover systems. Learn more about the lab here: www.mRNP.au.dk
20.02.2025 09:53 — 👍 8 🔁 12 💬 0 📌 0
“Today Google is launching an AI co-scientist, a new AI system built on Gemini 2.0 designed to aid scientists in creating novel hypotheses and research plans.
AI co-scientist is a collaborative tool to help experts gather research and refine their work”
blog.google/feed/google-...
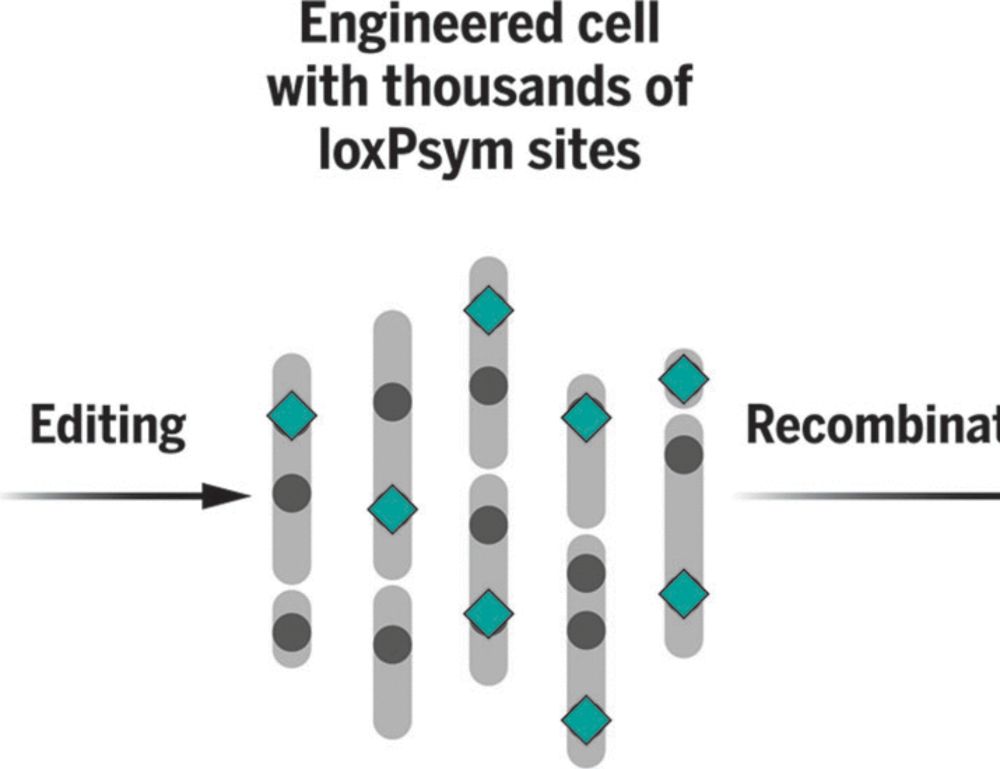
We're delighted to share our work on scrambling the human genome using prime editing, repetitive elements, and recombinases in @science.org , led by @jonaskoeppel.bsky.social , @f-raphael.bsky.social , with @proftomellis.bsky.social and George Church.
www.science.org/doi/10.1126/...

New postdoc opportunity in the THJ lab! Please share to anyone interested in nuclear RNA biology
@molbiolau.bsky.social
mbg.au.dk/en/news-and-...

We are pleased to announce registration for RNA 2025 - the 30th Annual Meeting of the RNA Society - is now open!
www2.rnasociety.org
This year’s meeting will be held at the beautiful Town and Country Resort & Convention Center in sunny San Diego, California, USA from May 27th - June 1st, 2025.
Hello BlueSky! Follow us to stay up to date on the inspiring science published in our journals and more.
To get started, check out our #starterpack to meet all the Cell Press journals currently on the platform, with more to come🧪🔬🤖🧬🌳🪲
go.bsky.app/TdKYuS7
Resident colour blind member of the lab here 🙋♂️
Please make your figures friendly to us.
Restrictor slows early transcription elongation to render RNA polymerase II susceptible to termination at non-coding RNA loci https://www.biorxiv.org/content/10.1101/2025.01.08.631787v1
10.01.2025 18:33 — 👍 4 🔁 4 💬 0 📌 0
Is there a common contaminant database for proximity labelling-MS data? I see that the CRAPome has a section for proximity-dependent biotinylation but it is limited to human cells.
I keep seeing the same nonsense proteins enriched in mTurbo-experiments and would love to know if this is common crap.

40 years ago all of Genbank was published in print form by NAR. The same format today would require over 4 light seconds of shelf space. To a year of progress in 2025.
29.12.2024 09:50 — 👍 564 🔁 216 💬 15 📌 25
Cell lines are frozen, chrome tabs, spreadsheets and documents have been closed. All done for this year ✌️
Back in Derbyshire and busy making sausage rolls.
It's been a productive year, but i'm in dire need of a break.

This project was speaheaded by Guifen Wu and is now available online at @naturecomms.bsky.social
doi.org/10.1038/s414...
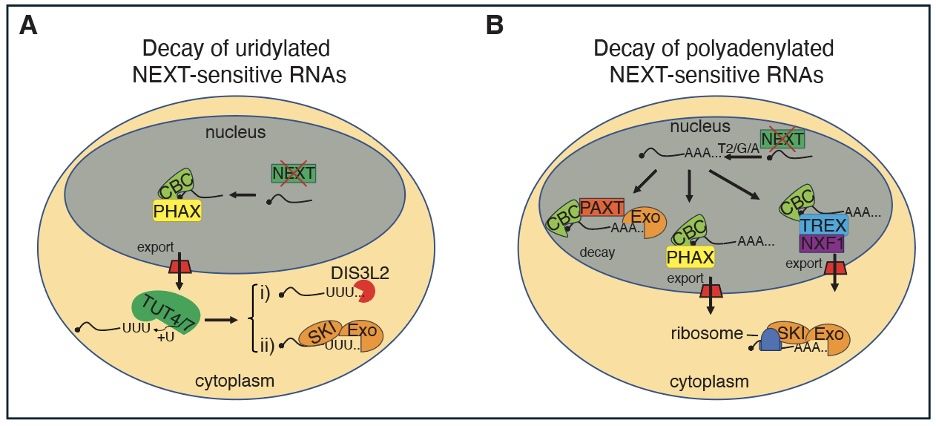
📢 Latest publication from the lab! We explore how cells manage the excess pA− RNAs generated by pervasive transcription. While the NEXT complex usually takes charge, in its absence, a network of tailing enzymes, export factors, and exonucleases steps in to modify and degrade these RNAs.
13.12.2024 08:25 — 👍 9 🔁 4 💬 1 📌 0
Don't use red and green data lines/surfaces in the same panel please #chemsky. It can be difficult for some colorblind readers to differentiate them. I've accepted (in principle) 2 papers today, and both sets of authors were asked to remove red/green colour contrasts www.nature.com/articles/d41...
10.12.2024 15:13 — 👍 113 🔁 47 💬 4 📌 3