View from the hotel room

Poster session 2024, with Valentina Boeva, Constantin Ahlmann-Eltze and others

Wednesday afternoon hike incl. swim in the mountain river

Another view from the hotel room
Apply for the Ascona workshop "Statistical and AI methods for multi-modal multi-scale modeling of biological systems", 28 Jun-3 Jul 2026 on Monte Verità, Lago Maggiore at the foot of the Swiss Alps.
ascona2026.sciencesconf.org
15.01.2026 16:16 —
👍 20
🔁 16
💬 0
📌 3
Very happy that MuViT, our (CVPR 2026!) work on learning across spatial scales in microscopy w transformers, is out! Check Martin's thread for a nice walkthrough :)
Big thanks to my advisors @maweigert.bsky.social @gioelelamanno.bsky.social!
🖥️: github.com/weigertlab/muvit
📜: arxiv.org/abs/2602.24222
02.03.2026 14:59 —
👍 16
🔁 6
💬 5
📌 0

For pretraining we use multi-resolution MAE: we mask tokens across scales and reconstruct each level from the others (cross-scale inpainting). This yields strong, scale-consistent representations and speeds up downstream training, and lets us leverage large unlabeled microscopy datasets.
02.03.2026 13:42 —
👍 2
🔁 0
💬 1
📌 0

Modern microscopy produces GB images with multi-scale features (cells > tissue > anatomy), yet most models use single res FOV. MuViT jointly encodes multiple physical regions in one transformer with world-coordinate RoPE for coherent cross-scale attention, ie it "sees" local + global context at once
02.03.2026 13:42 —
👍 3
🔁 0
💬 1
📌 0
Excited to share our new paper (CVPR 2026 🚀): "MuViT: Multi-Resolution Vision Transformers for Learning Across Scales in Microscopy" which enables local predictions to use global context.
Great work led by @albertdm.bsky.social & another fun collab w @gioelelamanno.bsky.social! @scadsai.bsky.social
02.03.2026 13:42 —
👍 47
🔁 19
💬 1
📌 2
#Mastodon, the Command Center for Large-Scale Lineage-Tracing Microscopy Datasets, is finally heading for publication (well, let's see). To #cite or not to cite shall no longer be the question 🙃, because the #preprint. Software is available in every #Fiji near you.
www.biorxiv.org/content/10.6...
13.12.2025 18:14 —
👍 113
🔁 48
💬 1
📌 1

Uncertainty quantification for connectomics - Nature Methods
As connectomics datasets grow in size and quantity, future reconstruction methods will have to work with minimal or no human supervision. For that, we will need methods that can quantify data and mode...
Method of the Year 2025: In his Comment, @janfunkey.bsky.social discusses current challenges in connectomics, which is time-consuming and labor-intensive. He focuses on proofreading and discusses how automated models might be further developed to embrace uncertainty.
www.nature.com/articles/s41...
08.12.2025 19:05 —
👍 9
🔁 1
💬 0
📌 0

🧠 The Lipid #Brain Atlas is out now! If you think #lipids are boring and membranes are all the same, prepare to be surprised. Led by @lucafusarbassini.bsky.social with Giovanni D'Angelo's lab, we mapped membrane lipids in the mouse brain at high resolution.
www.biorxiv.org/cgi/content/...
16.10.2025 06:23 —
👍 283
🔁 110
💬 7
📌 11

Large images have to be broken into tiles both for training and inference with neural networks. The tile predictions then need to be merged to produce the final volume prediction.
Segment large images without tiling artifacts: sharing our work that should have been presented at ICCV in 2 weeks - the brilliant first author Elena can’t go because of visa issues.
The paper: arxiv.org/abs/2503.19545 1/🧵
09.10.2025 12:56 —
👍 57
🔁 16
💬 2
📌 0
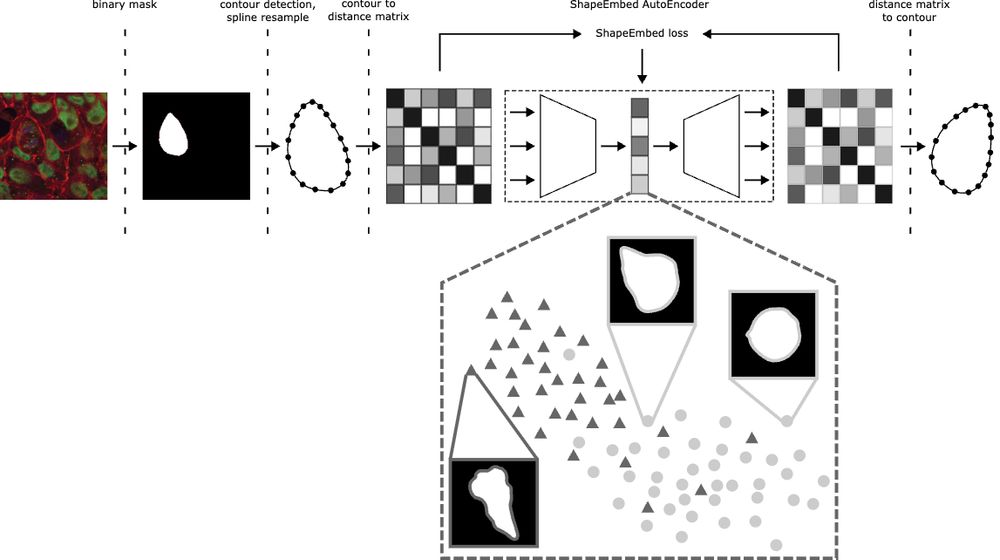
Happy to share that ShapeEmbed has been accepted at @neuripsconf.bsky.social 🎉 SE is self-supervised framework to encode 2D contours from microscopy & natural images into a latent representation invariant to translation, scaling, rotation, reflection & point indexing
📄 arxiv.org/pdf/2507.01009 (1/N)
23.09.2025 08:31 —
👍 71
🔁 26
💬 3
📌 5
🧵1/14 Preprint thread! Can we predict a cell’s fate based on its dynamics? 🔮 Our new study unveils a framework for watching development unfold in real-time, revealing how a cell's shape and movement encode info about its future fate. 🔬📄 Preprint: tinyurl.com/4shf8v4x
22.09.2025 09:37 —
👍 123
🔁 38
💬 9
📌 6
YouTube video by Blender
Microscopy Nodes: handling large biological volumes — Blender Conference 2025
I gave a talk at Blender Conference yesterday, showing how biological volumes work, and how to visualize them in Blender 😋 youtu.be/WPajSWX730o?... #bcon25
19.09.2025 07:35 —
👍 74
🔁 20
💬 5
📌 2
(1/14) I’m happy and proud to introduce: SpinePy – a framework to detect the "spine" of gastruloids and measure biological and physical signals in a local dynamic 3D coordinate system. www.biorxiv.org/content/10.1...
12.09.2025 12:43 —
👍 63
🔁 24
💬 4
📌 2
Thanks Misha for the shout-out!
It has been a truly mind-opening journey. WHOLISTIC is one step towards whole-organism neuroscience and a true physiological observatory - with extensions on the way! I am continually amazed by what the data reveals - so much remains to be discovered and mined.
08.09.2025 16:30 —
👍 48
🔁 11
💬 2
📌 1
Our new preprint is about the airway epithelium microtubule network, cilia, basal body protein composition, averaging of volumetric fluorescence data, and expansion microscopy.
Four years of very hard work from our very talented Emma van Grinsven. www.biorxiv.org/content/10.1... (1/N)
05.09.2025 15:37 —
👍 41
🔁 16
💬 2
📌 1
How is this possible?
This ant can lay eggs of two different species, birthed by the same mother
03.09.2025 15:13 —
👍 160
🔁 64
💬 4
📌 17
Congrats!! Way to go Akanksha!
04.09.2025 10:06 —
👍 1
🔁 0
💬 0
📌 0

Mechanical forces drive evolutionary change
A small tissue fold present in fruit fly embryos buffers mechanical stresses and may have evolved in response to mechanical forces.
Dresden researchers @paveltomancak.bsky.social @bruvellu.bsky.social, Carl Modes, @cuencam15.bsky.social & colleagues published in @nature.com that a tissue fold in fruit fly embryos buffers mechanical stresses & may have evolved in response to mechanical forces. www.mpi-cbg.de/news-outreac...
03.09.2025 15:26 —
👍 81
🔁 30
💬 3
📌 3

EPFL, ETH Zurich & CSCS just released Apertus, Switzerland’s first fully open-source large language model.
Trained on 15T tokens in 1,000+ languages, it’s built for transparency, responsibility & the public good.
Read more: actu.epfl.ch/news/apertus...
02.09.2025 11:48 —
👍 54
🔁 29
💬 1
📌 6

Unified mass imaging maps the lipidome of vertebrate development
Nature Methods - uMAIA is an analytical framework designed to enable the construction of metabolic atlases at high resolution using mass spectrometry imaging data.
Happy to share our work on uMAIA, a framework for building metabolomic atlases from mass spectrometry imaging.
With uMAIA, we mapped the lipidome of zebrafish development, uncovering spatially organized metabolic programs .
www.nature.com/articles/s41...
#developmentalbiology #MSI #lipidtime
03.09.2025 10:40 —
👍 16
🔁 10
💬 2
📌 0

Out today in @nature.com: Together with the Honigmann, Shevchenko, Drobot and Hof labs, we present a general workflow for imaging the localization and transport of individual lipids in cells and mapping their metabolism.
www.nature.com/articles/s41...
21.08.2025 05:18 —
👍 345
🔁 130
💬 31
📌 23

📢We're #hiring Group Leaders!
Apply to lead a lab at Janelia & advance biology using theory, computational modeling & machine learning.
🔹5-year renewable appointment
🔹Pioneer new tools & approaches
🔹Collaborate across disciplines
Apply by Nov. 4👉 https://janelia.link/groupleader
26.08.2025 19:00 —
👍 63
🔁 65
💬 0
📌 2
🧬🔬🧪 🎉 Our team at @czbiohub is thrilled to share TWO companion papers out today in @NatureMethods!
📦 Ultrack — robust, scalable nD cell tracking
🌐 inTRACKtive — a beautiful, open-source web viewer for lineage exploration
Let’s dive in! 👇
www.nature.com/articles/s41...
www.nature.com/articles/s41...
25.08.2025 09:06 —
👍 118
🔁 35
💬 3
📌 3

Our beautiful Hydra made the cover!!! 😍😍😍
An experiment performed by Pavel Mejstrik with a HCR protocol by @giuliaserafini.bsky.social, @paveltomancak.bsky.social lab @mpi-cbg.de
Actin fibers in blue, nuclei in orange.
13.08.2025 08:51 —
👍 154
🔁 25
💬 3
📌 0