
CCN Neural Data Analysis Workshop @ FENS 2026 www.simonsfoundation.org/event/ccn_fe...
03.03.2026 16:39 — 👍 3 🔁 1 💬 0 📌 0@wulfdewolf.bsky.social
PhD student @edinburgh-uni.bsky.social NeuroRSE intern @flatironinstitute.org NeuroAI intern @cshlnews.bsky.social Trying to understand spatial computation and memory in biological neural networks. I run a lot. wulfdewolf.github.io

CCN Neural Data Analysis Workshop @ FENS 2026 www.simonsfoundation.org/event/ccn_fe...
03.03.2026 16:39 — 👍 3 🔁 1 💬 0 📌 0Come analyse some neural data!! 🍍🍍🍍
03.03.2026 22:15 — 👍 3 🔁 0 💬 0 📌 0space cells
27.02.2026 23:50 — 👍 1 🔁 0 💬 0 📌 0
Mission: record hippocampal place cells in zero gravity
Crew: 3 rats with electrode arrays
Vehicle: Space Shuttle Columbia
Status: data now publicly available on DANDI, 28 years later
The ratstronauts' mission is finally complete. 🐀🚀 h/t NASA
about.dandiarchive.org/blog/2026/02...

Applications are now open for the summer school: 𝐌𝐚𝐭𝐡𝐞𝐦𝐚𝐭𝐢𝐜𝐚𝐥 𝐌𝐞𝐭𝐡𝐨𝐝𝐬 𝐢𝐧 𝐂𝐨𝐦𝐩𝐮𝐭𝐚𝐭𝐢𝐨𝐧𝐚𝐥 𝐍𝐞𝐮𝐫𝐨𝐬𝐜𝐢𝐞𝐧𝐜𝐞
🧠 Apply before March 15: www.compneuronrsn.org
📍 Located in beautiful Eresfjord 🇳🇴
🗓️ Between July 6-24
Supported by the @kavlifoundation.org
In collaboration with @kavlintnu.bsky.social
Go fit some neural models!!
JAX, Pynapple, Poisson GLMs; what more can you ever wish for...
Yes, fair. Though, if the LLM knows about the standardised format (NWB here), it does allow some handy prompts like: "Can you find datasets recorded from the hippocampus that provide both raw recordings and spike-sorted cells?"
It would take me a lot longer to list all of those.
chat.neurosift.app/chat
does this reasonably well for DANDI/EBRAINS/OpenNeuro datasets

Pre-COSYNE Brainhack 2026 – Join Us in Lisbon!
March 10–11, 2026 • Lisbon, Portugal
pre-cosyne-brainhack.github.io/hackathon2026/
Kick off COSYNE week with two days of hands-on, team-based hacking around real electrophysiology data, open tools, and reproducible workflows. #brainhack #cosyne2026

An overview of the Open Software Summer School 2026 schedule. There are two tracks in the first week, "Animals in Motion" and "Large Array Data". "BrainGlobe" and "Extracellular Electrophysiology" will be run in the second week.
Three weeks left to apply for the Neuroinformatics Unit Open Software Summer School, August 17-28 2026 in London, UK!
Bringing together researchers and open source developers of ephys, behaviour and image analysis tools.
neuroinformatics.dev/open-softwar...
Deadline January 31st. Apply now!

Can humans & animals really use internal maps to take shortcuts?
Tolman famously said yes - based largely on his Sunburst maze.
Our new review & meta-analysis suggests evidence is far weaker than you might think.
🧵👇 doi.org/10.1111/ejn....
@uofgpsychneuro.bsky.social @ejneuroscience.bsky.social

Just published my review of neuroscience in 2025, on The Spike.
The 10th of these, would you believe?
This year we have foundation models, breakthroughs in using light to understand the brain, a gene therapy, and more
Enjoy!
medium.com/the-spike/20...

A 🍍 spotted in the wild at @edinburgh-uni.bsky.social!
Great crash course on neuro data processing with @pynapple.bsky.social and @spikeinterface.bsky.social delivered by @matthiashennig6.bsky.social, @wulfdewolf.bsky.social and Chris Halcrow. So nice to see both libraries working so well together

Really excited to share this Opinion piece we've been working on with fellow head-direction cell geeks @apeyrache.bsky.social @desdemonafricker.bsky.social and (bsky-less?) Andrea Burgalossi! While head-direction cells pop up in many cortical regions, we think that one of them is quite unique (1/8)
15.10.2025 20:10 — 👍 46 🔁 18 💬 2 📌 1
@benjocowley.bsky.social @david-klindt.bsky.social
@dinanthos.bsky.social
Would love to hear what you think!
We present our preprint on ViV1T, a transformer for dynamic mouse V1 response prediction. We reveal novel response properties and confirm them in vivo.
With @wulfdewolf.bsky.social, Danai Katsanevaki, @arnoonken.bsky.social, @rochefortlab.bsky.social.
Paper and code at the end of the thread!
🧵1/7
Finally preprinted! TL;DR: thalamic head-direction neurons can shape their activity regardless of input. Big thanks to @apeyrache.bsky.social and master experimentalist @sskromne.bsky.social .
16.09.2025 20:25 — 👍 27 🔁 6 💬 0 📌 0
New workshop! Just before SFN, join the Flatiron Institute Center for Computational Neuroscience of the Simons Foundation for a workshop on pynapple and NeMoS. Learn how to use these open-source packages to analyze and model neural data! Accommodation & meals provided. Link ⬇️
25.08.2025 18:29 — 👍 14 🔁 9 💬 1 📌 0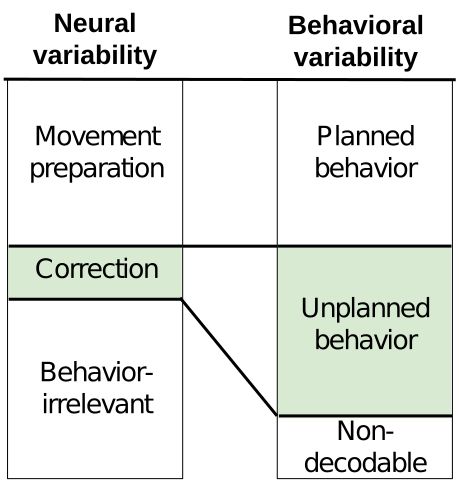
Excited to share our new pre-print on bioRxiv, in which we reveal that feedback-driven motor corrections are encoded in small, previously missed neural signals.
07.04.2025 14:54 — 👍 26 🔁 16 💬 1 📌 1
any chance I can still get the call link?
registrations are closed...

New paper from the lab: feedback from piriform cortex to the olfactory bulb carries identity and reward contingency signals in a multimodal (odor and sound) rule-reversal task. Feedback is re-formatted within seconds and reflects changes in the perceived rules of engagement.
doi.org/10.1038/s414...

We are pleased to be partnering with The UKRI AI CDT in Biomedical Innovation at the School of Informatics (University of Edinburgh). These PhD projects have a strong translational focus and seek to develop & apply AI approaches to biomedical domains ✨.
Apply by 20 Jan www.ai4biomed.io/how-to-apply/

Network model with excitatory feedforward connections (W) and recurrent inhibitory connections (M).
Check our latest work towards understanding learning and local plasticity in the brain with @matthiashennig6.bsky.social at @edinburgh-uni.bsky.social 🧠
Published in a @neuripsconf.bsky.social workshop this year. Now out on arxiv - arxiv.org/abs/2501.02402
Both the people and the environment at CSHL made my summer there the best I've ever had.
If you want to learn about neuroscience<->AI and meet interesting people, it would be a waste not to apply!
for better visibility: video of the Olney house woodpecker that made sure I was up on time every day
PhD project!
For those interested in NeuroAI: "Using large scale neural network models to bridge the translational gap from animal models to human insights in autism spectrum and neurodevelopmental disorders"
Apply here: www.ai4biomed.io/research/pro...
Now!
Or more likely, people should change what they think place cells mean.
Will give it a try! Thanks!!
22.11.2024 17:50 — 👍 1 🔁 0 💬 0 📌 0
Can't such much smart on CA2/3, will have to read up!
My only thought was that if (CA1) place cells "respond to the sequence of sensory inputs leading to a location",
then similar snapshots of activity will most often encode different things.
PCA only sees the instantaneous patterns, no sequences.
The paper doesn't look at place cells explicitly, but assuming everything transfers: at what point do we decide not to call it a place cell anymore?
22.11.2024 17:22 — 👍 0 🔁 1 💬 1 📌 0
I was going to comment about a biologically plausible implementation, but it seems you've heard that one enough.
As someone that looks at recordings all day, I'm constantly thinking about what signatures these models would leave behind.
I wonder if there's anything particular for CSCG.