
📣 I'm excited to share our latest preprint!
We adapt and characterise a neurosphere-based CNCC differentiation protocol, and demonstrate utility for quantitative phenotyping and craniofacial disease modelling! 🧫
Read about Array-CNCC here:
www.biorxiv.org/content/10.6...
@uoe-igc.bsky.social
28.01.2026 14:36 —
👍 48
🔁 13
💬 2
📌 1
We're also super excited to point you towards a wonderful, highly complimentary study from our colleagues Ana Banito and @nsbenab.bsky.social
www.biorxiv.org/content/10.6...
28.01.2026 09:54 —
👍 4
🔁 2
💬 0
📌 0
Combining degraders targeting p300 and SWI/SNF:
🧬 Strong synergy in SS18–SSX–positive cells
🧬 Profound loss of SS18–SSX chromatin binding
🧬 Broad collapse of oncogenic gene expression
🧬 Dramatic loss of cell viability
28.01.2026 09:54 —
👍 2
🔁 0
💬 1
📌 0
SWI/SNF and p300 impact different subsets of SS18-SSX binding sites
This gave us an idea...
👉 What if we therapeutically target both?
28.01.2026 09:54 —
👍 0
🔁 0
💬 1
📌 0
To recap:
Remove SWI/SNF ➡️ Modest effects on SS18-SSX binding
Remove p300 ➡️ Major loss of SS18-SSX occupancy and transcription
❗ P300, not SWI/SNF, sustains the oncogenic program
28.01.2026 09:54 —
👍 1
🔁 0
💬 1
📌 0
We found that p300 is physically and functionally coupled to SS18-SSX
🧬 Co-localises with SS18-SSX on chromatin
🧬 Supports SS18-SSX binding at target sites
🧬 Required for SS18-SSX driven transcription
➡️ Remove p300 and SS18-SSX falls of chromatin
28.01.2026 09:54 —
👍 1
🔁 0
💬 1
📌 0
So, what is the engine?
️A focused CRISPR screen revealed a striking dependency on p300
🧬 Not its paralog CBP
🧬 Specific to SS18-SSX positive sarcoma lines
28.01.2026 09:54 —
👍 1
🔁 0
💬 1
📌 0
Even when SWI/SNF activity is broadly eliminated:
🧬 Oncogenic transcription does not shut down
🧬 SS18-SSX transcriptional activity persists
➡️ Conclusion: SWI/SNF is not the main engine of SS18-SSX driven transcription
28.01.2026 09:54 —
👍 2
🔁 0
💬 1
📌 0
We used degraders to dismantle SWI/SNF complexes entirely, including SS18-SSX containing assemblies.
Result:
🧬 SWI/SNF complexes collapse
🧬 SS18-SSX stays bound at many target loci
🧬 Gene expression stays largely intact
28.01.2026 09:54 —
👍 3
🔁 0
💬 1
📌 0
Clinical efforts to target SWI/SNF (e.g. BRD9 degraders, based on our previous discoveries) have shown limited and transient benefits in synovial sarcoma patients.
So, we asked a simple question:
👉 Does SS18-SSX actually need SWI/SNF to drive oncogenic transcription?
28.01.2026 09:54 —
👍 1
🔁 0
💬 1
📌 0
Just a few days after international DMG/DIPG day, our paper showing the importance of CBX4/PCGF4 containing cPRC1 complexes in glioma is out in @cp-molcell.bsky.social. Below is a nice synopsis of the paper by joint first author @EimearLagan.bsky.social
21.05.2025 16:33 —
👍 8
🔁 1
💬 0
📌 0
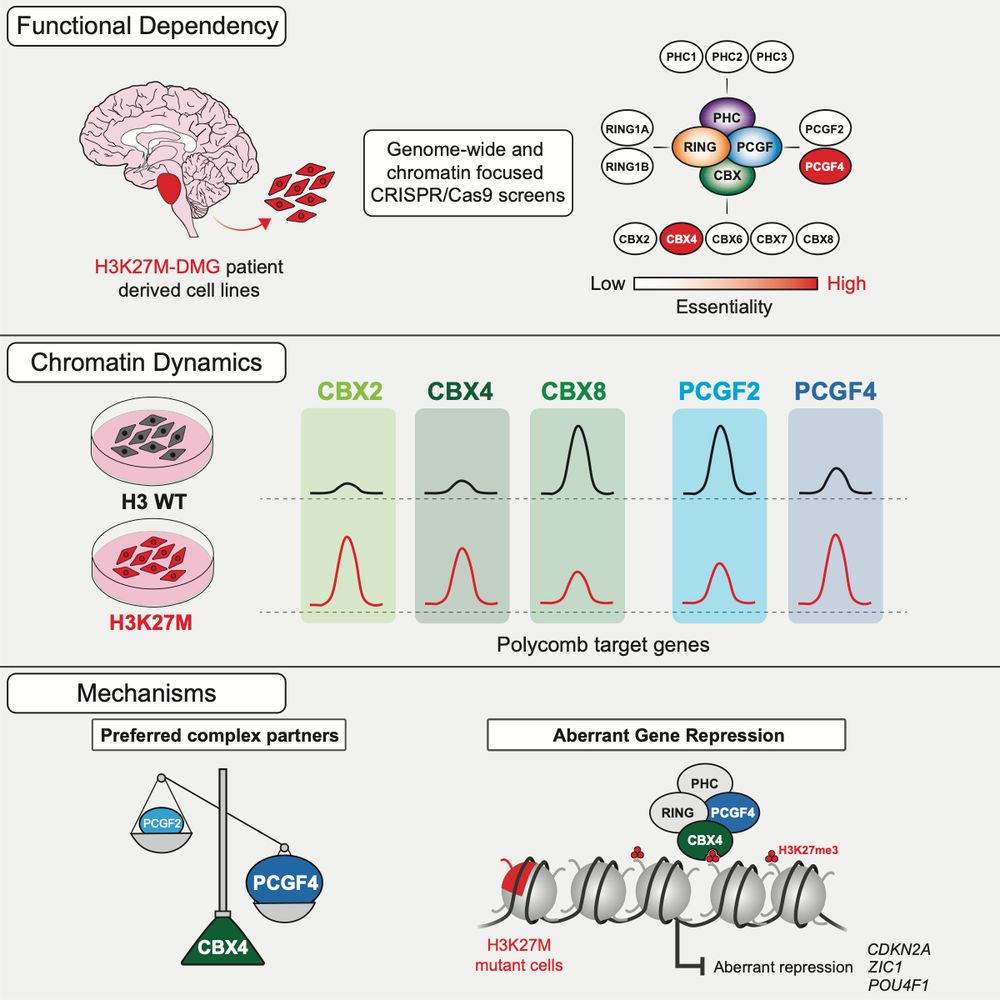
Excited to share our new paper out today in @cp-molcell.bsky.social! We show that the H3K27M oncohistone rewires cPRC1, creating a unique dependency on CBX4/PCGF4-containing complexes, and also reveal a previously unknown function of CBX4. Highlights below (1/11).
21.05.2025 16:12 —
👍 23
🔁 10
💬 5
📌 2

