High-throughput targeted paleoproteomics sex estimation on medieval Great Moravia individuals using MALDI-CASI-FTICR mass spectrometry https://www.biorxiv.org/content/10.64898/2026.02.17.706309v1
19.02.2026 05:31 — 👍 1 🔁 2 💬 0 📌 0High-throughput targeted paleoproteomics sex estimation on medieval Great Moravia individuals using MALDI-CASI-FTICR mass spectrometry https://www.biorxiv.org/content/10.64898/2026.02.17.706309v1
19.02.2026 05:31 — 👍 1 🔁 2 💬 0 📌 0Thanks a lot!
24.01.2026 07:36 — 👍 0 🔁 0 💬 0 📌 0Dear Ventresca, great job! Have you uploaded your MS Raw files to PRIDE for future re-analysis? :)
23.01.2026 15:19 — 👍 0 🔁 0 💬 1 📌 0I am pleased to share that on December 18, I received the Award for the Best Researcher of the Year at Vilnius University in the Open Science category. This recognition was granted for my work on untargeted #paleoproteomics for the biological sex identification of archaeological remains
19.12.2025 15:03 — 👍 14 🔁 3 💬 0 📌 0
PAASTA WINTER SCHOOL Now is your chance to provide input on our upcoming PAASTA Winter School on tools and topics you’re keen to learn and teach, please give your thoughts on this short form forms.gle/om48gkCKzTt9... before 9th Jan!
05.12.2025 05:32 — 👍 12 🔁 11 💬 0 📌 4I'm hiring!⭐ As part of the DFF Sapere Aude-funded project MiddleEarth, I am looking for 1 postdoctoral researcher and 1 research assistant.
27.10.2025 09:54 — 👍 14 🔁 23 💬 1 📌 3I am honored to have collaborated with him and he has been an excellent mentor to young scientists.
04.09.2025 06:59 — 👍 0 🔁 0 💬 0 📌 0Pr. Jaroslav Brůžek was a notable biological anthropologist based in France and the Czech Republic. He has contributed significantly to the understanding and development of new methods and tools for the sex identification of human remains, including #paleoproteomics
04.09.2025 06:59 — 👍 1 🔁 0 💬 1 📌 0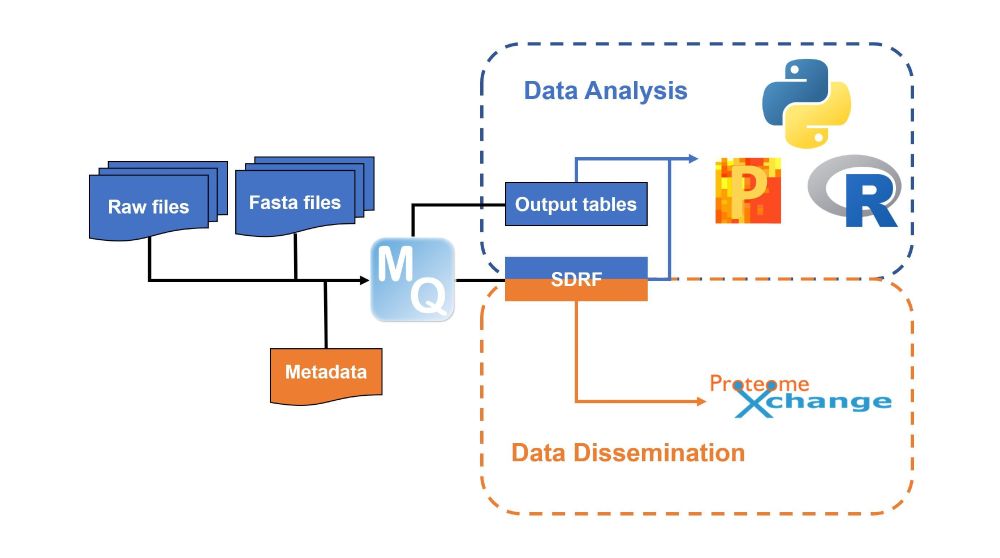
New release! MaxQuant 2.7.0 & Perseus 2.1.5 introduce metadata tools that enhance both public data reusability and personal research workflows:
- Export metadata as SDRF
- Annotate output tables with SDRF
Tutorial: bit.ly/3TO9GyV
Preprint: bit.ly/4eMwqt3
#TeamMassSpec

Good news! My article received enough views to be a Top Viewed Article. Read it here onlinelibrary.wiley.com/doi/full/10.... #TopViewedArticle @wiley.com
17.04.2025 08:37 — 👍 8 🔁 2 💬 0 📌 0
1 of 4 We're hiring a Laboratory Manager for the CODICUM project at @UniCopenhagen to help explore medieval manuscript fragments using cutting-edge biomolecular methods!
Apply by April 13, 2025!
Link: employment.ku.dk/staff?show=1...
#LabJobs #BiomolecularScience #AncientDNA
👇

Human AMELX (position 47 to 66) and AMELY (position 57 to 67) reveal two different SAPS in the alignment, methionine (M) in position 59 and a serine (S) at position 66 instead of a proline (P) in the AMELX corresponding position. Below that are three peptide-spectrum matches from Australopithecus africanus Sts 63 for human AMELY with M and S highlighted in red. Note peptide spectrum graphs plot mass-to-charge (m/z) values of ions on the x-axis, as measured during the mass spectrometric analysis of peptide fragments, and the relative peak abundance (%) on the y-axis. The red peaks represent y-ions, which are generated from fragmentation at the C-terminal side of peptide bonds, and they correspond to red bars between amino acids in the peptide sequence. The blue peaks represent b-ions, which result from fragmentation at the N-terminal side of peptide bonds, and they correspond to blue bars between amino acids in the peptide sequence. These peaks are matched to the theoretical spectrum of the peptide, aiding in the identification of the peptide sequence. This figure was generated using the publicly free site www.proteomicsdb.org/use/ by inputting MS2 mass to charge ratios.
Congratulations to @palaeoprotpalesa.bsky.social!
1st successful #palaeoproteomics analysis of a 3.5-2.01 Ma Australopithecus africanus 🦷 reveals male sex!
Demonstrates huge potential of minimally invasive enamel analysis #teammassspec
Her article in sajs.co.za/article/view...
Thread 🧵

Save 👏 the 👏 date 👏
www.lecturejeunesse.org/evenement/44/

A l’occasion des 50 ans de la découverte, le 24 novembre 1974, de #Lucy squelette partiel d’australopithèque daté de 3,2 millions d’années, Sonia Harmand, archéologue préhistorienne au CNRS, dévoile trois faits méconnus sur cette ancêtre de l’humanité.
www.cnrs.fr/fr/actualite...
I have created a #palaeoproteomics and #ZooMS starterpack - please DM if you want to be added to the list.
22.11.2024 19:23 — 👍 25 🔁 9 💬 1 📌 2