
Very happy to share our paper rdcu.be/eUImj out today in @natcellbio.nature.com 🎉🎉🎉
We uncover an unexpected role for endogenous Xist RNA in regulating X-linked genes that escape X-inactivation.

Very happy to share our paper rdcu.be/eUImj out today in @natcellbio.nature.com 🎉🎉🎉
We uncover an unexpected role for endogenous Xist RNA in regulating X-linked genes that escape X-inactivation.
📣 Yesterday our study uncovering the ancestry bias in gene annotations was finally published in @natcomms.nature.com!
www.nature.com/articles/s41...
If you work in human genetics or transcriptomics, do not miss our tweetorial!👇
@guigolab.bsky.social @bsc-cns.bsky.social @crg.eu

Giving a seminar today @ethz.ch on our new X-reactivation findings during aging—back at the institute where I started my PhD 10 years ago. Can’t wait to see some familiar faces!
25.11.2025 09:25 — 👍 2 🔁 1 💬 0 📌 0
Poster advertising the 6th X-inactivation meeting from Oct 19-23 2026 in Sapporo, Japan. The organizers are Asato Kuriowa, Edda Schulz, Ikuhiro Okamoto, Rafael Galupa, Takashi Sado, Mitinori Saitou.
📣 SAVE THE DATE
Next X-inactivation meeting in Sapporo, Japan, 19-23 October 2026. Visit x-inactivation-meeting.org to join our mailing list. 🧬 speakers @dandergassen.bsky.social @marnieblewitt.bsky.social @heard65.bsky.social @crougeulle.bsky.social @sexchrlab.bsky.social @zhouqi1982.bsky.social

This work, led by Lison Lemoine and co-authors @sarahhoelzl.bsky.social, @hasenbeint.bsky.social and Elisabeth Graf @tum.de, has been published today in Scientific Reports: www.nature.com/articles/s41...
11.11.2025 19:00 — 👍 2 🔁 1 💬 0 📌 0This workflow provides a powerful and accessible framework for studying allele-specific transcript diversity, highlighting the mechanistic insights made possible by long-read transcriptomic data.
11.11.2025 19:00 — 👍 0 🔁 0 💬 1 📌 0We explored imprinted loci, known for allele-specific coding and non-coding isoforms, and successfully benchmarked historical findings. Our approach also uncovered isoforms expressed from both the active and inactive X chromosomes of escape genes in females.
11.11.2025 19:00 — 👍 1 🔁 0 💬 1 📌 0
New paper from the lab: we’re using long-read sequencing to disentangle isoform complexity at allele-specific loci 🧬💡
Here, we combine the PacBio Iso-Seq workflow with the established WhatsHap phasing approach to assign long reads to the correct allele in polymorphic F1 mouse hybrids.
Thank you 🔥
25.10.2025 09:57 — 👍 0 🔁 0 💬 0 📌 0
All mouse (6 major organs) and human (54 GTEx tissues) lncRNA-to-target predictions are easily accessible for visual inspection, allowing researchers to select candidates based on target and mechanism predictions
GitHub IGV resource link: github.com/AndergassenL...
Importantly, as more individual sequencing data and risk variants become available, we anticipate that this strategy will continue to elucidate targets and mechanisms, ultimately decoding the entire cis-acting landscape of the non-coding genome.
24.10.2025 02:43 — 👍 0 🔁 0 💬 1 📌 0We applied the strategy to mouse organs and to the largest allele-specific human dataset @gtexportal.bsky.social, including 54 tissues from nearly 1000 individuals. Given the high genetic diversity in humans, each individual allows for the discovery of new allelic correlation events.
24.10.2025 02:43 — 👍 0 🔁 0 💬 1 📌 0By further integrating H3K27ac data, we showed that the same principle can be used to link enhancers and other cis-acting DNA regulatory elements to their corresponding targets!
24.10.2025 02:43 — 👍 1 🔁 0 💬 1 📌 0The strategy is simple: A repressive lncRNA-to-target prediction occurs when the lncRNA and nearby protein-coding genes show opposite allele-specific expression biases, while an enhancing interaction occurs when both show allelic expression bias toward the same allele.
24.10.2025 02:43 — 👍 1 🔁 0 💬 1 📌 0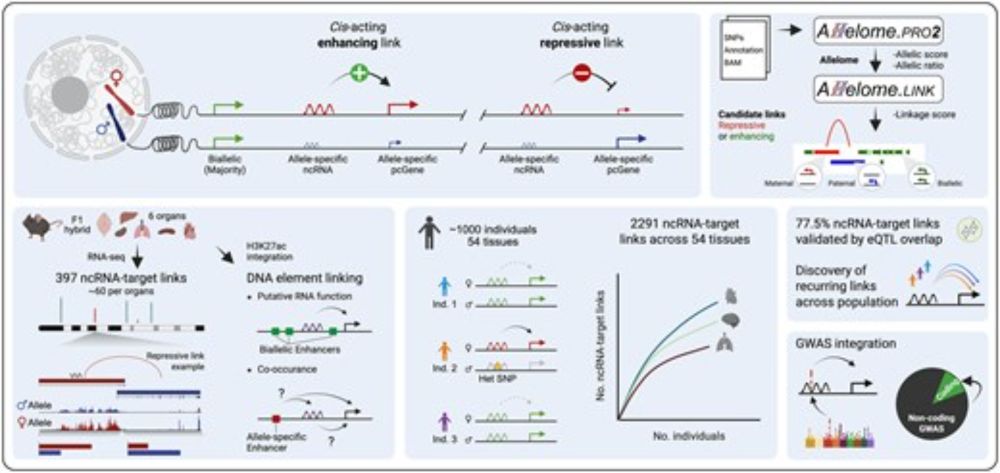
This work, led by @hasenbeint.bsky.social and co-authors @sarahhoelzl.bsky.social, Stefan Engelhardt at the @tum.de #TRR267, has been published in @narjournal.bsky.social
link: doi.org/10.1093/nar/...
Yes! The Allelome.LINK framework integrates allele-specific transcriptomics and/or epigenomics to connect cis-acting lncRNAs and enhancers with their nearby protein-coding targets, thereby linking overlapping non-coding disease variants to the genes they affect.
24.10.2025 02:43 — 👍 2 🔁 0 💬 1 📌 0
Cracking the code of the non-coding genome via allele-specific genomics?
Can we link non-coding elements—like lncRNAs and enhancers—to their protein-coding target genes, and in doing so, connect overlapping non-coding disease variants to their protein-coding counterparts?

📢 Paper Alert: tinyurl.com/yzy2d864. We characterized tRNA-overlapping lncRNA loci = tROLs! tROL perturbations silence
codon-biased genes in inter-chromosomal proximity. tROLs bridge the non-coding
and coding genomes. @sickkidsto.bsky.social @uoftpress.bsky.social
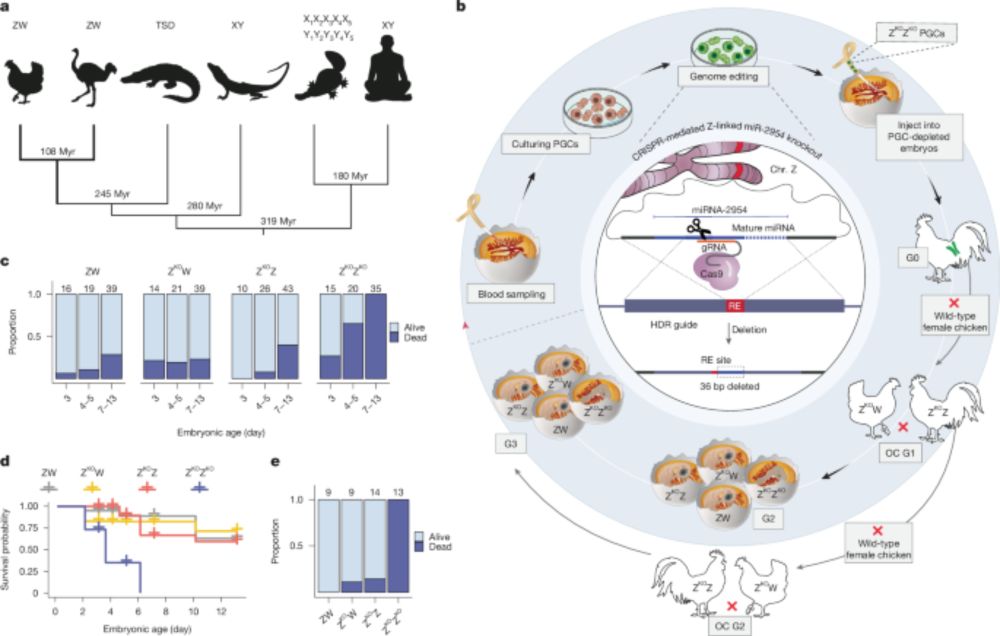
Our study on a male-essential microRNA and the evolution of other dosage compensation mechanisms in birds is now out in Nature! www.nature.com/articles/s41...
16.07.2025 15:23 — 👍 123 🔁 58 💬 10 📌 6
🗞️Our June issue is live!📷 This month, we're featuring work on editing epigenetic age, somatic mutation, senescence, Alzheimer’s biomarkers and much more. Read it all here: nature.com/nataging/vol...
18.06.2025 14:36 — 👍 12 🔁 4 💬 0 📌 0
We are so happy to be on the cover @sarahhoelzl.bsky.social 🎉 The cover depicts the Three Fates, who manipulate the threads of life and death. The Fates are shown as three older women, unraveling the threads of the inactive X chromosome during aging. www.nature.com/nataging/vol...
18.06.2025 17:20 — 👍 5 🔁 1 💬 1 📌 0
Why do women experience aging-related diseases differently than men? A new study shows that with age, genes on the inactive X chromosome can reactivate – potentially influencing conditions like #dementia and #autoimmunity: go.tum.de/987255
#genetics #aging
📷 @dandergassen.bsky.social

We're excited to publish our latest study led by Bryony Leeke @bryonyleeke.bsky.social and Wazeer Varsally, now out in @nature.com 🍾This study focusses on the epigenome of marsupial embryos 🦘 mapping DNA methylation in embryo development to specific embryo events www.nature.com/articles/s41... 🧵
14.05.2025 16:01 — 👍 98 🔁 38 💬 9 📌 4
Our new contribution to the quest to find causal GWAS genes! Sam Ghatan from my lab at @nygenome.org led a systematic comparison of eQTLs and CRISPRi+scRNA-seq screens. TL;DR: they provide highly complementary insights, with ortogonal pros and cons. 🧵👇
www.biorxiv.org/content/10.1...

Do you work in 💫human genetics💫?
Have you ever worried about what’s inside your gene annotation GTF⁉️
WELL, YOU SHOULD! 😱
Especially when studying a genetically diverse 🌍 cohort!
🔴We discover that gene annotations are European-biased 👉 impacting downstream analyses!
Don't miss this thread🧵⬇️
1/13

Finally, we would like to thank all the reviewers for their valuable comments, the @nataging.nature.com editors, as well as Anton Wutz for highlighting our research in the News & Views section of Nature Aging: rdcu.be/eknAp
02.05.2025 19:57 — 👍 3 🔁 1 💬 0 📌 0
Overall, the escape landscape shows a high degree of organ and cell type specificity, with strong evidence of cluster organization. Explore the full Escape Atlas in the IGV browser for each organ and age time point: github.com/AndergassenL...
02.05.2025 19:57 — 👍 1 🔁 0 💬 1 📌 0Since we find that several age-specific escapees are associated with human diseases, we propose that their elevated expression in females may contribute to sex-biased disease progression with age, offering a new mechanism for age-related sex differences beyond hormonal influence.
02.05.2025 19:57 — 👍 0 🔁 0 💬 1 📌 0
In addition, allele-specific single-cell analysis revealed that age-specific escape manifests within distinct cell types, further providing evidence that age-related epigenetic changes promote gene escape.
02.05.2025 19:57 — 👍 1 🔁 0 💬 1 📌 0
An intriguing example is Reps2, where a specific isoform escapes in an age-dependent manner. While the long isoform of Reps2 is expressed from the Xa, the promoter of the short isoform shows age-dependent biallelic activity, leading to Xi-specific expression of the short isoform.
02.05.2025 19:57 — 👍 0 🔁 0 💬 1 📌 0