The small #GTPase Rab1 is a master regulator of #Golgi traffic and #autophagy. van Vliet, Munro et al. @mrclmb.bsky.social performed a proteomic screen to identify novel Rab1 effectors, including a #dynein motor adaptor and cargo receptors for autophagy. rupress.org/jcb/article/...
#Trafficking
07.01.2026 17:45 —
👍 17
🔁 12
💬 0
📌 1
#KANK1 #LC8 #IntrinsicallyDisorderedProteins #AlphaFold #EM
08.01.2026 15:07 —
👍 1
🔁 1
💬 0
📌 0

Disordered linker of Kank1 forms rigid protein rod upon binding to 15 LC8 molecules and bridge the gap between membrane and microtubule.
New collaborative preprint!
A striking result: a disordered linker in KANK1 binds as many as 15 LC8 dimers, forming a protein rod that bridges cell membranes and microtubules!
www.biorxiv.org/content/10.1...
07.01.2026 18:09 —
👍 18
🔁 11
💬 0
📌 2

Final version is out! Our large-scale cryo-ET dataset 🔬 of Chlamydomonas reinhardtii 🦠 is now published in @cp-molcell.bsky.social
Huge collaborative effort! So glad to see the community already using it to develop new resources & tools.
Check it out here: shorturl.at/z4i4c
#CryoEM #CryoET
19.12.2025 16:45 —
👍 137
🔁 47
💬 3
📌 1
Weak Motifs, Strong Complex: KANK1 Uses Cooperative Multivalency with the Hub-Protein LC8 to Bridge Cytoskeletal Complexes https://www.biorxiv.org/content/10.1101/2025.07.16.665182v1
21.07.2025 11:47 —
👍 1
🔁 1
💬 0
📌 0

Dynein-2 prefers to bind the retrograde-targeted A-tubule in intraflagellar transport - likely because of its tyrosination, suggest cryo-ET data and MD simulations by @ichikawa-lab.bsky.social and coworkers
www.embopress.org/doi/full/10....
28.11.2025 16:10 —
👍 9
🔁 3
💬 0
📌 0

Volume 138 Issue 20 | Journal of Cell Science | The Company of Biologists
Journal of Cell Science publishes cutting-edge science encompassing all aspects of cell biology. It is published by The Company of Biologists, a not-for-profit organisation.
Our special issue on ‘Cilia and Flagella’ is now out! Huge thanks to all (authors + reviewers) who contributed, and to our wonderful Guest Editors (@cilialab.bsky.social + @lottepedersen.bsky.social) for pulling it together.
Happy reading!
#cilia #flagella
journals.biologists.com/jcs/issue/13...
03.11.2025 17:32 —
👍 50
🔁 23
💬 1
📌 1

Advances in ciliary proteomics – towards cracking the hidden proteome code of cilia
Summary: This Review explores proteomic strategies for investigating cilia, tracing the evolution from early discoveries to recent advancements enabled by proximity labeling techniques.
📣 Here it is — our overview on ciliary proteomics! Great discussions and writing with @mick-lab.bsky.social and Ronald Roepman 🙏🙏
Hope we didn’t miss anything important - apologies if we did!
#cilia #ciliopathies #proteomics
@theracilproject.bsky.social @for5547.bsky.social
tinyurl.com/yc8ztn5v
01.11.2025 16:34 —
👍 18
🔁 8
💬 1
📌 1

a man stands in front of a white board with the words keep that in mind written on it
ALT: a man stands in front of a white board with the words keep that in mind written on it
Miao Gui and I wrote a Review for the @jcellsci.bsky.social special issue “Cilia and Flagella: From Basic Biology to Disease”. It provides a comprehensive and up-to-date overview of MIP structure, function, and the latest tools used to study them, crafted to serve both newcomers and experts.
03.11.2025 05:18 —
👍 8
🔁 5
💬 1
📌 0

1000 citation alert by Google Scholar
A milestone reached! 1000 citations on Google Scholar!
30.10.2025 16:39 —
👍 5
🔁 0
💬 0
📌 0
Happy to share our lab's recent #cryoEM studies that show how a SPIN90 dimer activates Arp2/3 complex to form either uni or bidirectional actin filaments. Congrats🎉👏 @justusfrancis.bsky.social @achyutha-kp.bsky.social @tejashyamk.bsky.social @sridharskulkarni.bsky.social @kiranvyshnav.bsky.social
15.09.2025 12:05 —
👍 15
🔁 6
💬 1
📌 0
Excited to share our latest work with @simonbullock11.bsky.social! We looked at how diverse mRNAs get selected for subcellular localization and it turns out that a single protein can recognize different RNA elements using shared features that weren’t apparent before.
www.biorxiv.org/content/10.1...
05.08.2025 10:43 —
👍 143
🔁 45
💬 4
📌 9
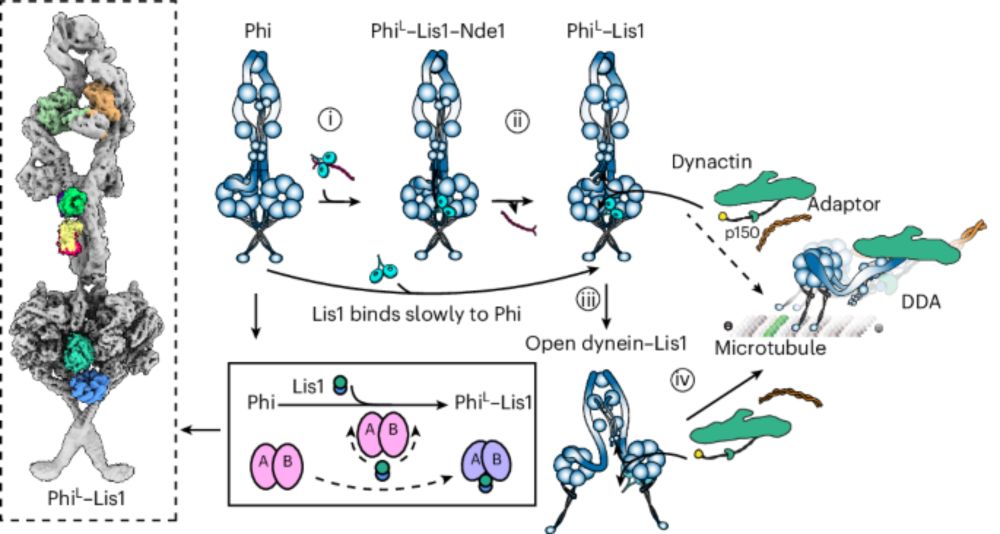
Thrilled to see our study on how kinesin-2 motors are switched on and off published in @natsmb.nature.com ⚛️
➡️ www.nature.com/articles/s41...
Congrats to all authors from me and Anthony 🎉 @dunnschool.bsky.social Check out this animation made by talented PhD student @matthew-batisio.bsky.social 😆
29.07.2025 09:04 —
👍 63
🔁 26
💬 1
📌 3
Congratulations!!! 🎉🎉🎉
01.08.2025 15:43 —
👍 1
🔁 0
💬 0
📌 0
Come work at Nature Methods! A dream job. I know from experience 😄
10.07.2025 12:35 —
👍 37
🔁 22
💬 0
📌 1
Microtubule end stabilisation by cooperative oligomers of Ska and Ndc80 complexes
During mitosis, properly aligned chromosomes stabilise microtubule ends with the help of kinetochores to ensure timely segregation of chromosomes. Microtubule-binding components of the human outer kinetochore, such as Ndc80 and Ska complexes, are present in multiple copies and together bind several microtubule ends, creating a highly multivalent binding interface. Whereas Ndc80:Ndc80 and Ndc80:microtubule binding is crucial for interface stability, Ndc80 alone in absence of Ska is unable to support stable kinetochore-attachments. Using cryoET, we demonstrate that oligomeric Ndc80:Ska assemblies stabilise microtubule ends against shortening by strengthening lateral contacts between tubulin protofilaments at microtubule plus-ends. We further identify a point mutation within the SKA1 microtubule-binding domain that does not affect microtubule-binding of individual Ska molecules, but does abolish Ska:Ska interactions. Finally, we report that oligomerisation of Ska, in a cooperative fashion together with the Ndc80, is necessary to maintain stable microtubule attachments both in vivo and in vitro. ### Competing Interest Statement The authors have declared no competing interest. BBSRC, BB/X014975/1, BB/W019698/1 Wellcome Trust, https://ror.org/029chgv08, 308895/Z/23/Z
New preprint from the lab - the first paper made (almost) entirely here @qmul.bsky.social
We report how human outer kinetochore complexes Ndc80 and Ska form cooperative oligomers, that together stabilise microtubule ends against shortening.
www.biorxiv.org/content/10.1...
Key results below: (1/7)
07.07.2025 09:39 —
👍 26
🔁 8
💬 1
📌 1
Latest preprint from the lab, many years in the making!
By combining #cryoEM with #AlphaFold3 modelling, we propose that norovirus NS3 forms a transmembrane RNA translocase.
This could have big implications for our understanding of viral replication & assembly (🧵)
www.biorxiv.org/content/10.1...
18.06.2025 06:32 —
👍 179
🔁 54
💬 9
📌 3
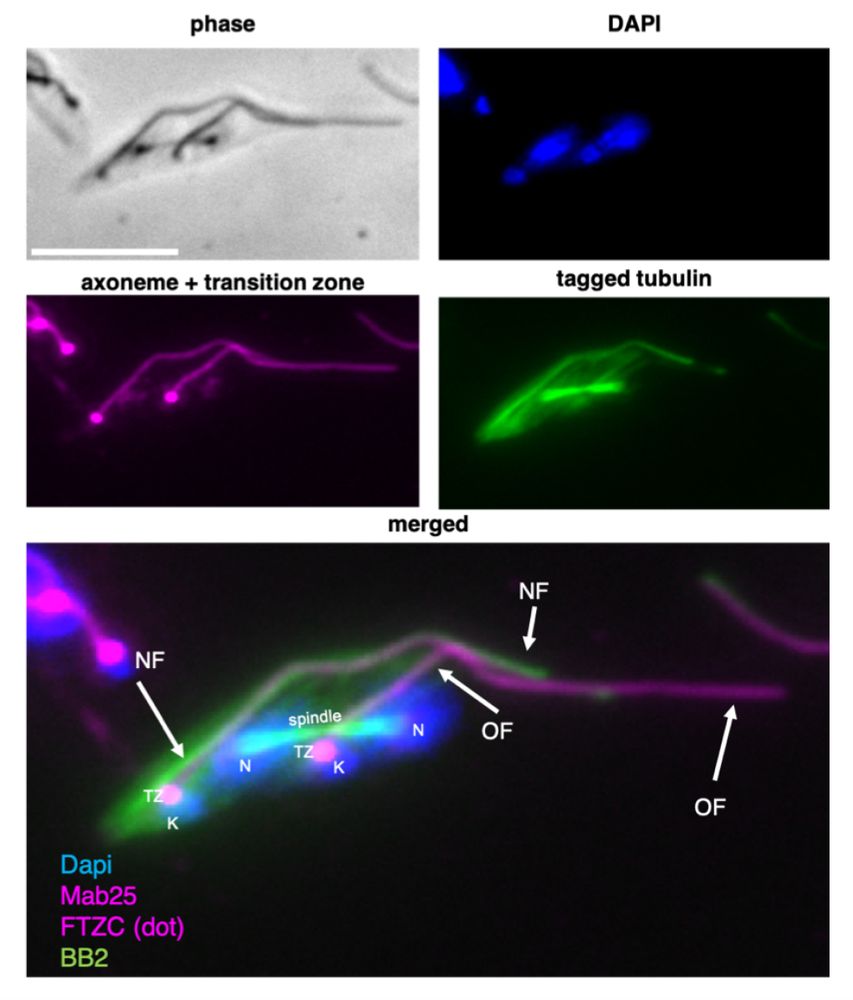
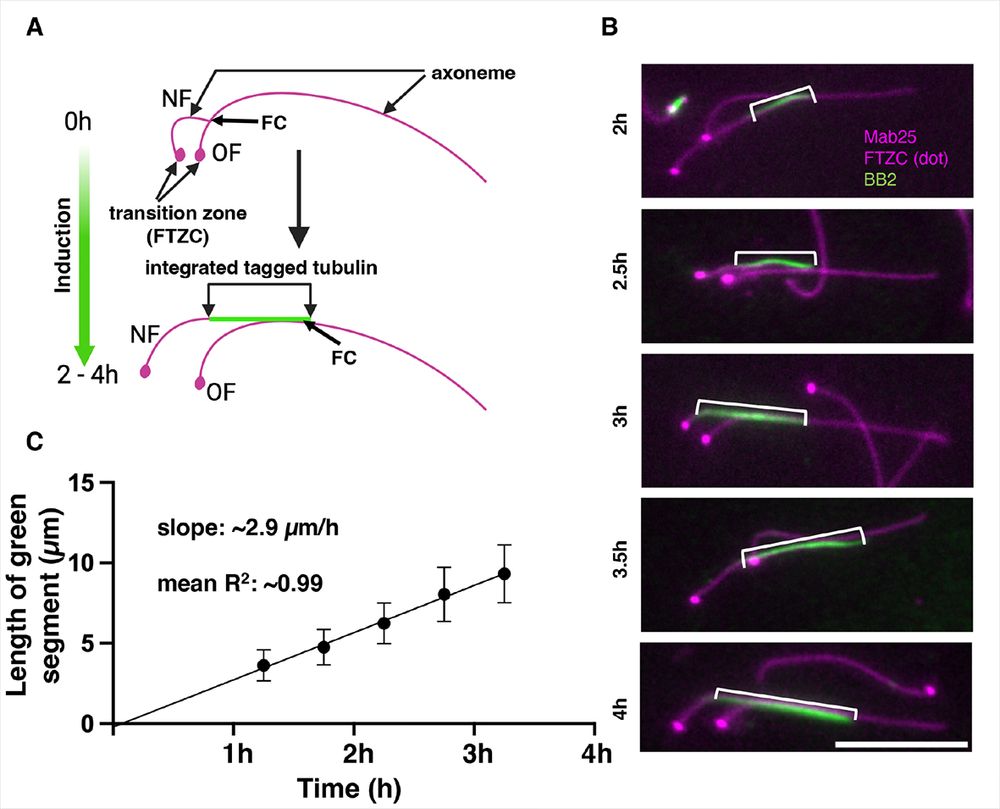
Daniel's paper is now published in @jcellsci.bsky.social, great experience with @reviewcommons.org 😀
Want to know everything tubulin assembly in trypanosome flagella and other microtubules? Here is the link:
journals.biologists.com/jcs/article/...
12.06.2025 08:49 —
👍 43
🔁 20
💬 1
📌 0

Unveiling the structural spectrum of SARS-CoV-2 fusion by in situ cryo-ET | Nature Communications
SARS-CoV-2 entry into host cells is mediated by the spike protein, which drives membrane fusion. While cryo-EM reveals stable prefusion and postfusion conformations of the spike, the transient fusion intermediate states during the fusion process remain poorly understood. Here, we design a near-native viral fusion system that recapitulates SARS-CoV-2 entry and use cryo-electron tomography (cryo-ET) to capture fusion intermediates leading to complete fusion. The spike protein undergoes extensive structural rearrangements, progressing through extended, partially folded, and fully folded intermediates prior to fusion-pore formation, a process that depends on protease cleavage and is inhibited by the WS6 S2 antibody. Upon interaction with ACE2 receptor dimer, spikes cluster at membrane interfaces and following S2’ cleavage concurrently transition to postfusion conformations encircling the hemifusion and initial fusion pores in a distinct conical arrangement. S2’ cleavage is indispensable fo
SARS-CoV-2 entry captured: Cryo-ET reveals fusion intermediates of spike protein during membrane fusion, bridging stable pre- and postfusion states. PMID:40461447, Nat Commun 2025, @NatureComms https://doi.org/10.1038/s41467-025-60406-z #Medsky #Pharmsky #RNA #ASHG #ESHG 🧪
04.06.2025 10:10 —
👍 14
🔁 4
💬 0
📌 0
2026 Canada Excellence Research Chair Opportunities | Research & Innovation
🇨🇦🚨Job Alert🚨🇨🇦
Canada Excellence Research Chair (C$5 or C$10M, 8 yr budget)
U Toronto, Dept of Biochem/LabMedPath
*Was just told they're looking for a structural biologist using cryoEM* (related to infectious disease?)
Are you an established PI ready to move to 🇨🇦?
research.utoronto.ca/funding-oppo...
31.05.2025 12:46 —
👍 37
🔁 45
💬 1
📌 0