
Gene-scale in vitro reconstitution reveals histone acetylation directly controls chromatin architecture
Reconstituting 20-kb chromatin shows that tuning acetylation alone reshapes its folding, dynamics, and contact domain formation.
Yohsuke T. Fukai, Kyogo Kawaguchi, et al. reconstituted gene-scale #chromatin arrays with controlled #histone modification patterns in vitro and showed that heterogeneous #acetylation alone can drive pattern-dependent changes in chromatin architecture and dynamics.
doi.org/10.1126/scia...
28.11.2025 08:36 —
👍 2
🔁 1
💬 0
📌 0

We just released a new major version of TrackMate (v8), the cell and organelle tracking plugin of Fiji.
It ships many new features, detailed below, but that are articulated around the following:
20.11.2025 11:42 —
👍 182
🔁 76
💬 4
📌 7

Gene-scale in vitro reconstitution reveals histone acetylation directly controls chromatin architecture
Reconstituting 20-kb chromatin shows that tuning acetylation alone reshapes its folding, dynamics, and contact domain formation.
To probe gene-scale chromatin physics, we built 96-mer (20 kb) arrays with defined histone marks. Combining single-molecule tracking, AFM imaging, and developing in vitro Hi-C, we saw how specific modifications dictate chromatin structure and dynamics. www.science.org/doi/10.1126/...
20.11.2025 07:46 —
👍 60
🔁 21
💬 5
📌 1
Published in Science Advances! Looking forward to discussing the implications of our results and their applications with many researchers.
www.science.org/doi/10.1126/...
20.11.2025 02:24 —
👍 9
🔁 5
💬 1
📌 0
I truly appreciate the organizers, core contributors, and all attendees for this wonderful opportunity! I enjoyed learning and working on napari in a welcoming environment.
Those interested in talking about and contributing to napari, let’s join the regular community meeting together!
31.10.2025 03:16 —
👍 10
🔁 6
💬 2
📌 0

napari core team members coding with community members at the GloBIAS 2025 hackathon at RIKEN BDR in Kobe, Japan.

napari core team members and community members in a discussion at the GloBIAS 2025 hackathon at RIKEN BDR in Kobe, Japan.

napari core team members coding with community members at the GloBIAS 2025 hackathon at RIKEN BDR in Kobe, Japan.

napari core team members and community members posing together for a group photo on a staircase at the GloBIAS 2025 hackathon at RIKEN BDR in Kobe, Japan.
Longer wrap-up to come, but in the meantime: we had a fantastic and productive time at the napari hackathon at @globias.bsky.social 2025 conference! Thanks to all the participants and to the GloBIAS team for hosting us!
30.10.2025 06:53 —
👍 22
🔁 6
💬 0
📌 2
Last call for Les Houches! Apply now or never! ;) (ok, in 2 years again then) and share with your friends.
22.10.2025 19:00 —
👍 3
🔁 6
💬 0
📌 0

FY 2025 Call for RIKEN ECL Team / Unit Leaders
Call for RIKEN ECL Team / Unit Leaders. In my (biased) opinion, this is probably the best option if you are seeking an independent position in Japan. I will be happy to chat/answer questions via email. www.riken.jp/en/careers/p...
23.05.2025 10:40 —
👍 4
🔁 1
💬 0
📌 0
Excited to share our preprint on the molecular architecture of heterochromatin in human cells 🧬🔬w/ @jpkreysing.bsky.social, @johannesbetz.bsky.social,
@marinalusic.bsky.social, Turoňová lab, @hummerlab.bsky.social @becklab.bsky.social @mpibp.bsky.social
🔗 Preprint here tinyurl.com/3a74uanv
11.04.2025 08:35 —
👍 359
🔁 141
💬 12
📌 20
I feel like this post is becoming even more relevant for our early-career researcher friends in the US—unless it’s already too late. It doesn’t have to be our lab or RIKEN; if you’re interested in coming to Japan, I’m always happy to chat!
14.02.2025 09:17 —
👍 1
🔁 2
💬 0
📌 0
Starting in April 2025, our RIKEN Hakubi Lab will transition to a Chief Scientist Lab, and we plan to move to Wako in 2026. My affiliation with the IPI, Department of Physics at The University of Tokyo, will continue as before.
04.02.2025 02:30 —
👍 2
🔁 1
💬 1
📌 0

GitHub - yfukai/laptrack: Particle tracking by solving linear assignment problem.
Particle tracking by solving linear assignment problem. - yfukai/laptrack
Re-posting from my twitter:
LapTrack ( github.com/yfukai/laptr... ) provides you the linear-assignment based robust tracking algorithm, aware of splitting and merging particles, in a simple, Pythonic and flexible API. Installable with “pip install laptrack”.
05.01.2025 09:17 —
👍 2
🔁 0
💬 0
📌 0
We’re entering an exciting era of gene-scale chromatin reconstitution with precise modification patterns🚀 We hope this work will stimulate further studies to understand how different histone modifications and solution environment control chromatin architecture and function. 5/5
05.01.2025 09:11 —
👍 0
🔁 0
💬 0
📌 0

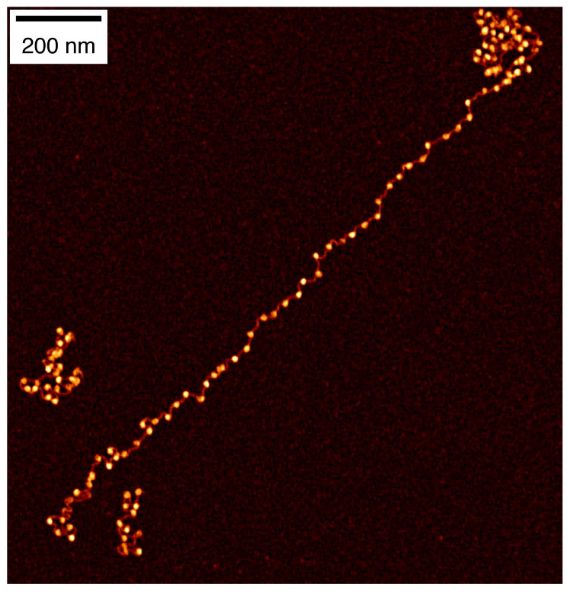
How do histone modifications affect large-scale chromatin structure? We tackled this question in vitro by reconstituting gene-scale chromatin arrays with precise histone modification patterns and analyzing them using single-molecule fluorescence imaging and in vitro Hi-C. 2/5
05.01.2025 09:11 —
👍 0
🔁 0
💬 1
📌 0
Thrilled to share our latest preprint on bioRxiv! 🎉 This collaboration with @kyogok.bsky.social and lab members,
Kurumizaka lab(Kujirai-san in particular!) and Umehara lab sheds new light on how histone modification shapes chromatin architecture. 1/5
biorxiv.org/content/10.1...
05.01.2025 09:11 —
👍 9
🔁 1
💬 1
📌 1


























