
In this Molecular Cell Voices piece, @genophoria.bsky.social comments on how he thinks about bridging bench and computational work: www.cell.com/molecular-ce...
11.02.2026 19:55 —
👍 3
🔁 2
💬 1
📌 0
Amazing collaboration with the Jain lab! The impact of systemic hypoxia on cancer!
10.02.2026 21:02 —
👍 3
🔁 2
💬 0
📌 0

Arc Core Investigator @genophoria.bsky.social will speak on a free, online panel with @tkaraletsos.bsky.social, @emmalundberg.bsky.social, and @ronalfa.bsky.social on Wed., Oct. 29, as part of GENbio's “The State of AI in Drug Discovery in 2025.” Register at: webinars.liebertpub.com/e/the-state-...
24.10.2025 17:32 —
👍 6
🔁 3
💬 0
📌 0

Hear how Arc, Ultima Genomics, and @10xgenomics.bsky.social are partnering to generate perturbation data at the scale needed to train virtual cell models on The Bio Report podcast with guests @genophoria.bsky.social, Gilad Almogy, and
Serge Saxonov: thebioreport.podbean.com/e/transformi...
09.10.2025 16:51 —
👍 7
🔁 2
💬 0
📌 0

#CELLFIE for CAR T screening—a new mRNA-based platform for screening primary cells. CAR + gRNA library are delivered by lentivirus, CRISPR modifiers as electroporated mRNA. That’s more flexible and effective than existing T cell screening methods. (1/7)
24.09.2025 15:47 —
👍 3
🔁 1
💬 1
📌 0

Tumor cell-adipocyte gap junctions activate lipolysis and contribute to breast tumorigenesis
Nature Communications - Breast cancer cells interact with neighbouring adipocytes, but the mechanisms are not fully understood. Here, the authors show that triple-negative breast cancer (TNBC)...
New work from stellar scientists Jeremy Williams and Roman Camarda. Many contributors including the amazing Zena Werb and Atul Butte
Work explores how breast cancer cells interact directly with adipocytes via gap junctions, leading to lipid release.
@ctbatucsf.bsky.social @ucsfcancer.bsky.social
20.08.2025 10:19 —
👍 16
🔁 11
💬 0
📌 1
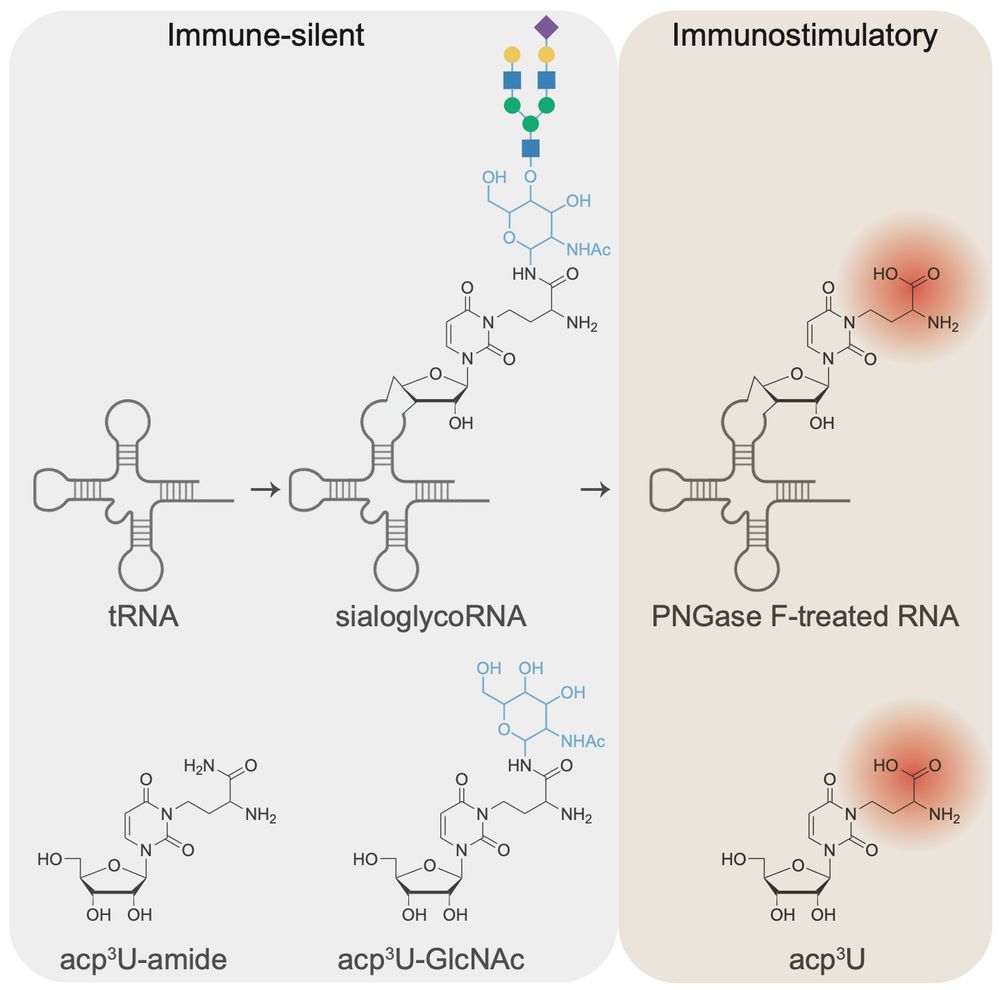
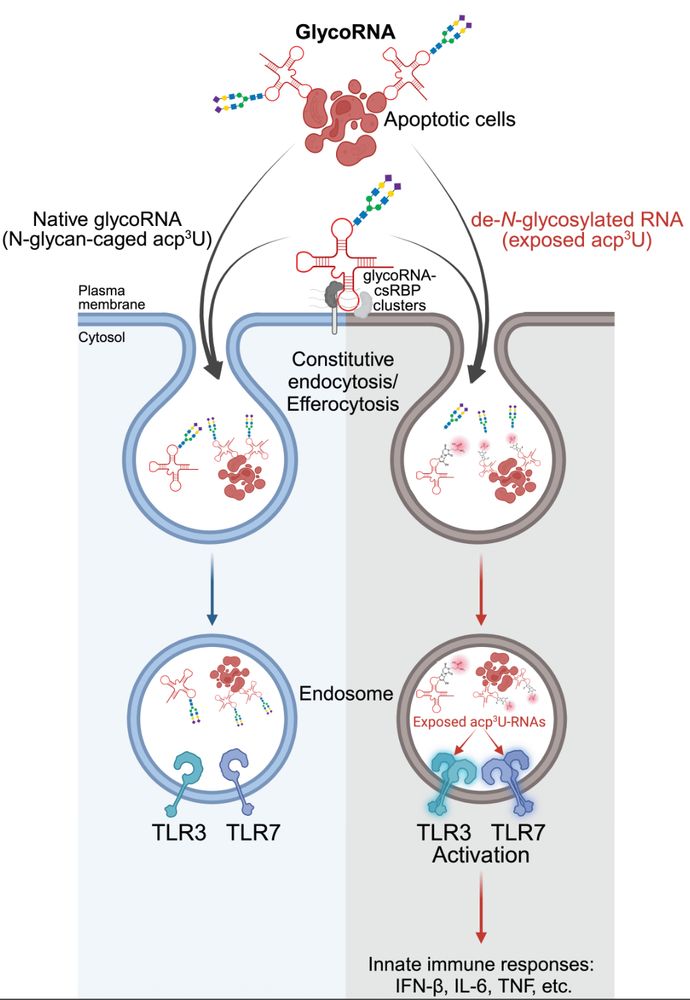
RNA N-glycosylation enables immune evasion and homeostatic efferocytosis by chemically caging acp3U. Excited to report this work lead by Vinnie @vinnieviruses.bsky.social and in collaboration with @vijayrathinam.bsky.social in @nature.com www.nature.com/articles/s41...
06.08.2025 15:36 —
👍 108
🔁 46
💬 7
📌 2
Go @exai.bio! Looking forward to working with Mike as our incoming CEO and our amazing board members.
24.07.2025 17:01 —
👍 1
🔁 0
💬 0
📌 0

Are you currently in the ERE related field & want to find new collaborative opportunities? Join our Chairs @chiappinellilab.bsky.social, @shenhui1986.bsky.social, Ting & Tao at #EREHD25 this Nov!
🎙️Talk Submission extended to 03 Sept
💰Final few $500 grants remaining
Don't Miss out! bit.ly/4eRq9Mp
17.07.2025 15:25 —
👍 11
🔁 8
💬 1
📌 1
Congratulations... amazing!
26.06.2025 06:03 —
👍 0
🔁 0
💬 1
📌 0
Our ENS is vital to gut-brain health. When it breaks down, so does health. Treatments? Almost none.
Access to functional human enteric neurons at scale powers disease modeling and therapeutic discovery. Grateful to everyone who made this possible. www.nature.com/articles/s41...
25.06.2025 21:06 —
👍 6
🔁 2
💬 4
📌 0
Oh no! I am devastated!
14.06.2025 19:00 —
👍 0
🔁 0
💬 0
📌 0
Check out this amazing work from @genophoria.bsky.social’s lab, revealing a novel post-transcriptional tumour suppressive programme in breast cancer 👇👇
09.06.2025 13:38 —
👍 4
🔁 2
💬 0
📌 0
We would especially like to thank all of our colleagues who collaborated with us on this, including @balynzaro.bsky.social and Albertas Navickas, for their contributions to this work. And our funders, including @arcinstitute.org.
09.06.2025 13:29 —
👍 0
🔁 0
💬 0
📌 0

Finally, we used double-knockdown for in vivo lung colonization assays to confirm the expected epistatic interaction between RBMS3 and TXNIP.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

We confirmed that TXNIP is correlated with RBMS3, and similarly its expression is associated with better outcomes. Then we looked at TXNIP expression in a cohort of 96 breast cancer samples in-house, stratified by stage, that confirmed reduced expression as disease progresses.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

To figure out which, if any, of the targets of RBMS3 that we identified are responsible for this phenotype, we performed an in vivo CRISPR screen. Among the targets tested, we found that silencing TXNIP increases metastatic lung colonization.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

To turn this clinical association into causation, we took advantage of metastasis assays in xenografted mice. We showed that silencing and over-expressing RBMS3 modulates metastasis in vivo in two independent models.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

Interestingly, this association was most notable in the basal and Claudin-low subtypes of breast cancer, which are known to be aggressive subtypes.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

We consistently observed that reduced expression of RBMS3 is associated with poor clinical outcomes in breast cancer patients, in multiple datasets.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

To gain insight into the functional consequences and phenotypic effects of this regulon, we looked at pathways and complexes that may be enriched among the RBMS3 target. We saw a significant enrichment for TGFb and VEGF signaling that are associated with metastatic progression in breast cancer.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

Finally, we demonstrated the requirement and sufficiency of RBMS3 binding sites for this RBSM3-mediated transcript stabilization. We cloned binding sites from 13 high confidence mRNA targets into a reporter assay, which showed RBMS3-dependent increase in reporter expression.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

Replacing AUA-carrying mRNAs with these empirical RBMS3 CLIP targets confirmed that RBMS3 direct binding drives transcript stabilization.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

To show that this effect results from a direct interaction between RBMS3 and its target mRNAs, we performed CLIP-seq (UV crosslinking followed by immuno-precipitation and sequencing). We observed pervasive 3’UTR binding, and also a strong enrichment for the AUA element we had identified.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

Silencing RBMS3, as expected, resulted in a significant decrease in the stability and expression of the mRNAs that contain AUA-elements in their 3’UTRs.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

Among these, we chose RBMS3 for further characterization, simply because it was the highest ranking RBP for correlated expression with its putative regulon—identified based on the AUA cis-regulatory element, which was determined by analysis of GreyHound’s sequence importance scores.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0

Applying GreyHound to our breast cancer mRNA stability dataset, we uncovered a network of RBPs associated, and their cognate cis-regulatory elements, as a major determinant of variation in the RNA dynamics of breast cancer.
09.06.2025 13:29 —
👍 0
🔁 0
💬 1
📌 0






















