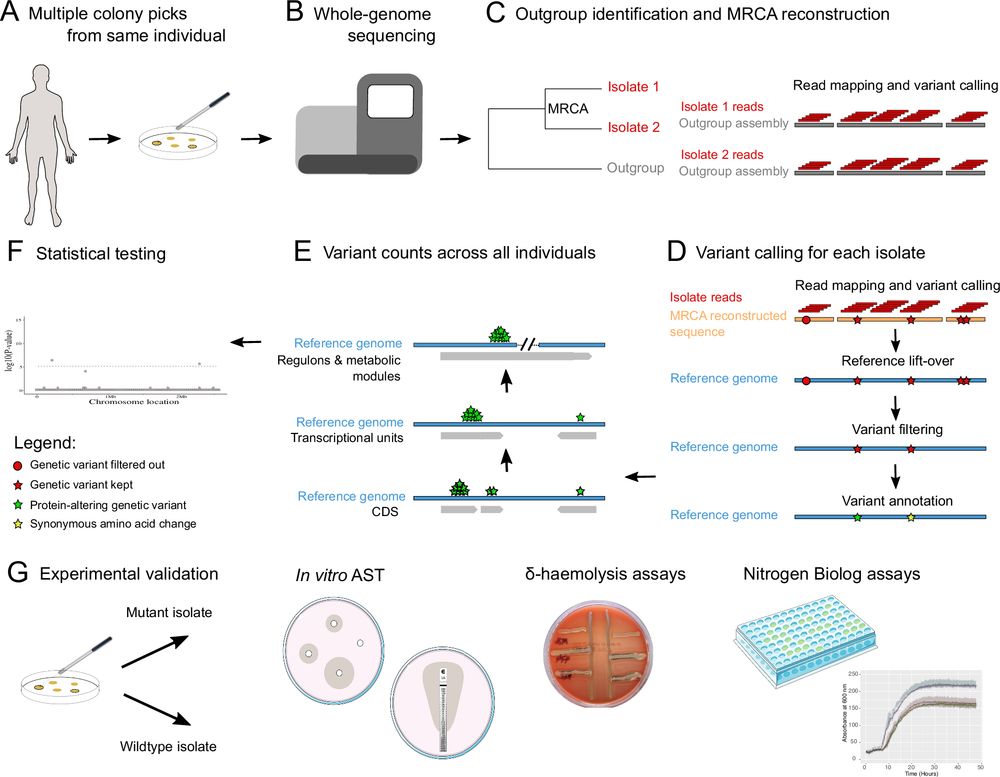
Bacterial metabolic remodeling by convergent evolution unlocks nutrient availability after a host switch
Staphylococcus aureus has undergone metabolic remodeling to adapt to the dairy niche.
Insights into how #Staph aureus adapts to the bovine host by unlocking nutrients from the dairy niche. Great work by Amy Pickering, Jamie Gorzynski and others in the group. Bacterial metabolic remodeling by convergent evolution after a host switch.|Science Advances www.science.org/doi/10.1126/...
18.02.2026 09:40 —
👍 29
🔁 15
💬 0
📌 0
Hi Nick, I sent you an email invite- not sure if you check them though 🙂 ?
20.11.2025 11:10 —
👍 0
🔁 0
💬 0
📌 0
PhD opportunity in bacterial pathogenesis- deadline tomorrow Feb 13 !
12.02.2025 13:12 —
👍 4
🔁 3
💬 1
📌 0

Fantastic to see this growing family of Fleming Fund fellows, mentors and supporters joining to meet the challenges of #AMR 🧪🧫 @edinburgh-uni.bsky.social 🇰🇪🏴🇲🇼🏴🇺🇬
@bachmanntill.bsky.social @edinuni-irr.bsky.social @roslininstitute.bsky.social
31.01.2025 11:44 —
👍 3
🔁 2
💬 0
📌 0

Applications are open for the 2025 GRS/GRC on Staphylococcal Diseases! Make sure to get your abstracts submitted for the potential to present your work in beautiful Barcelona! www.grc.org/staphylococc...
14.01.2025 16:52 —
👍 29
🔁 23
💬 0
📌 1
Good stuff. Yes- keen on that idea
04.12.2024 15:28 —
👍 0
🔁 0
💬 0
📌 0

Project
Project details
NERC Fully-funded PhD available with me & Jo Stevens on Legionella ecological adaptation to man-made water systems.
e4-dtp.ed.ac.uk/e5-dtp/super...
04.12.2024 13:59 —
👍 1
🔁 5
💬 0
📌 0
Yes and now big rush to recruit before start date...
04.12.2024 10:47 —
👍 0
🔁 0
💬 1
📌 0
EID is one of the biggest Infectious Disease networks in Europe
20.11.2024 15:35 —
👍 3
🔁 0
💬 1
📌 0