

New to me today: the downy mildew Peronospora valerianellae on Valerianella
#WildPlantDisease #FungiFriends


New to me today: the downy mildew Peronospora valerianellae on Valerianella
#WildPlantDisease #FungiFriends

How could a simple self-replicating system emerge at the origins of life? RNA polymerase ribozymes can replicate RNA, but existing ones are so large that their self-replication seems impossible. Could they be smaller?
Excited to share our latest work in @science.org on a new small polymerase.
1/n

Fig. 2 (shortened, full legend in paper): Structurally characterized M. oryzae effectors in complex with the HMA or HMA-like domains of host target proteins and ID-NLRs. Crystal structures of (A) AVR-PikF with the HMA-like domain of OsHIPP19 from rice (7B1I) (Maidment et al., 2021); (B) APikL2A with the HMA-like domain of sHMA25 from foxtail millet (7NLJ) (Bentham et al., 2021); (C) Pwl2 with the HMA domain of OsHIPP43 from rice (8R7D) (Zdrzałek et al., 2024); (D) AVR-PikD with the HMA-like ID of Pikp-1 from rice (5A6W) (Maqbool et al., 2015); (E) AVR-Pia with the HMA-like ID of Pikp-1 (6Q76) (Varden et al., 2019); (F) AVR1-CO39 with the HMA-like ID of RGA5 from rice (5ZNG) (Guo et al., 2018). Complexes are displayed such that the HMA/HMA-like domains are in equivalent orientations. Domains from HPP/HIPP host targets are coloured light green. Domains from ID-NLRs are coloured dark green.
⚙️🦠 REVIEW 🦠⚙️
Turley & Faulkner explore the function of plant heavy metal-associated domain-containing proteins and speculate about their functions at plasmodesmata by drawing from plant–pathogen interaction studies.
🔗 doi.org/10.1093/jxb/...
#PlantScience 🧪 Christine Faulkner

1/12 I'm ecstatic to share my preprint on legume NLR tissue expression! We investigated the NLRomes of 28 legumes + 4 outgroups, examining tissue expression across 7 legume species. Paper thread below 🧵👇
@thesainsburylab.bsky.social @itqbnova.bsky.social
🔗https://doi.org/10.64898/2026.01.25.701577

Are you interested in learning more about identifying plant pathogens in the wild? Come along to my webinar at 7pm on the 27th of January!
bsbi.org/take-part/ev...
#FungiFriends #WildPlantDisease

Pik pairs as resources for determining optimal ‘chassis’ for engineering recognition.
Mismatch screening in Nicotiana benthamiana to explore Pik-1/Pik-2 paired NLR platforms for receptor engineering
Yuxuan Xi (席玉轩) and Mark J. Banfield
nph.onlinelibrary.wiley.com/doi/10.1111/...
#PlantScience
I’m excited to share a new preprint from Carella group #CarellaCapybaras! biorxiv.org/content/10.6...
In this study, we show molecular co-evolution of two popular nonhost resistance genes in plants, RAR1 and SGT1, in ferns. For a quick read before Christmas, here’s the thread:
Stepwise and lineage-specific divergence of a major immune co-chaperone complex in leptosporangiate ferns https://www.biorxiv.org/content/10.64898/2025.12.17.694876v1
18.12.2025 04:02 — 👍 4 🔁 3 💬 0 📌 0Super excited to see this pre-print out from the lab! Happy to have helped to establish & carry out the protoplast cell death assays for this work. Congrats to @danielsyu.bsky.social and all other authors 🥳🌱
16.12.2025 19:52 — 👍 10 🔁 1 💬 0 📌 0Evolution of HMA-integrated tandem kinases accompanied by expansion of target pathogens https://www.biorxiv.org/content/10.64898/2025.12.15.692859v1
16.12.2025 18:01 — 👍 5 🔁 6 💬 0 📌 1
Super-happy sharing my Lab's new work on engineering plant HMA-containing tandem kinase immune receptors. Congrats to all authors (@danielsyu.bsky.social and @njw33.bsky.social on here). Work only possible in collaboration with Soichiro Asuke. See linked paper to follow!
bsky.app/profile/bior...

Excited to share our #plantimmunity preprint on engineering cereal TKPs
www.biorxiv.org/content/10.6...
A short thread below 👇
Engineering plant tandem kinase immune receptors expands effector recognition profiles https://www.biorxiv.org/content/10.64898/2025.12.15.694194v1
16.12.2025 16:02 — 👍 11 🔁 10 💬 0 📌 1Benchling darkmode when
06.12.2025 21:35 — 👍 17 🔁 1 💬 0 📌 0
CC-NLRs exhibit diverse conformations in the resting state. Figure depicts different NLR resting and activated states. While ZAR1 exists as a monomer bound to its guardees in its resting state, several forms have been observed for the NRC2 helper, including dimers, tetramers, and filaments. While filament-like structures were observed via confocal microscopy for NRC2 upon overexpression in planta, these structures were not detected for its paralog NRC4, suggesting a potentially different resting state configuration for NRC4. Our work suggests that the Pik CC-NLR pair forms a hetero-complex bound to the plasma membrane in its resting state. How effector binding to the integrated domain (ID) of the Pik-1 sensor leads to the activation of the complex remains to be determined.
Excited to share our latest work on plant immune receptor biochemistry! @kamounlab.bsky.social @hsuanpai.bsky.social @mpcontreras.bsky.social Jose Salguero Linares @danielluedke.bsky.social @adnroide.bsky.social @jiorgoskourelis.bsky.social
www.biorxiv.org/content/10.6...

Interactions between plant pathogens (left) and plants across five quadrats in a restored species-rich grassland. The basic premise of my PhD is to see if these interaction networks differ between restored and ancient grasslands.
#FungiFriends #NetworkEcology
New pre-print from the team!
The manuscript is @emma-raven.bsky.social's PhD work showing that whether a leaf is a carbon sink or a carbon source influences how they execute immune responses.
Have a read!
#PlantScience
@johninnescentre.bsky.social
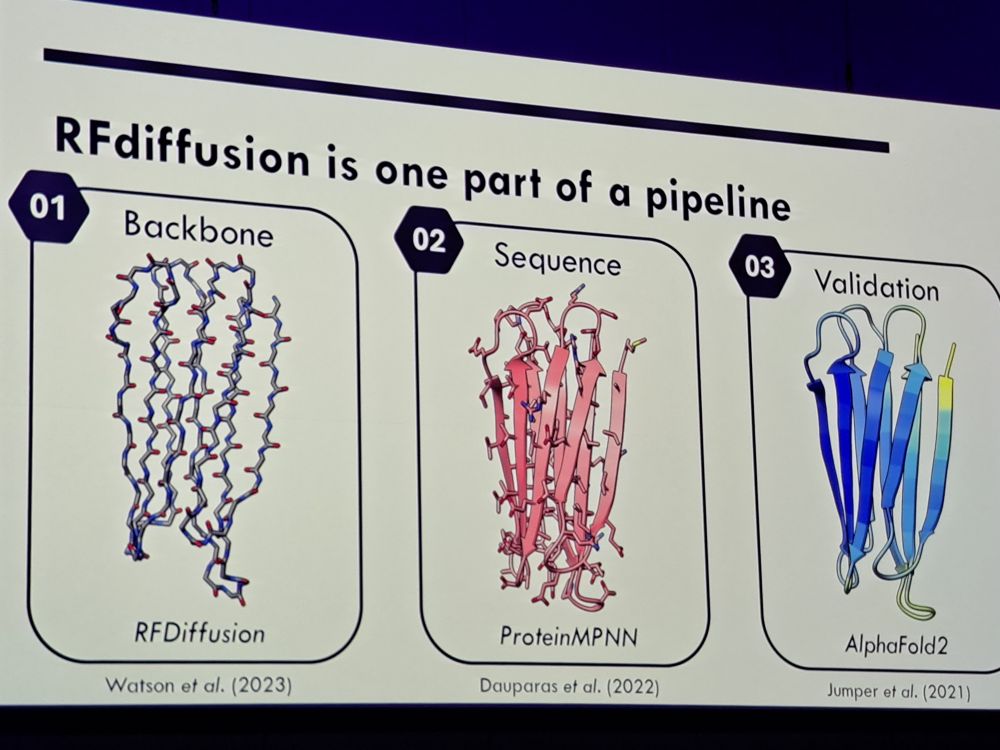
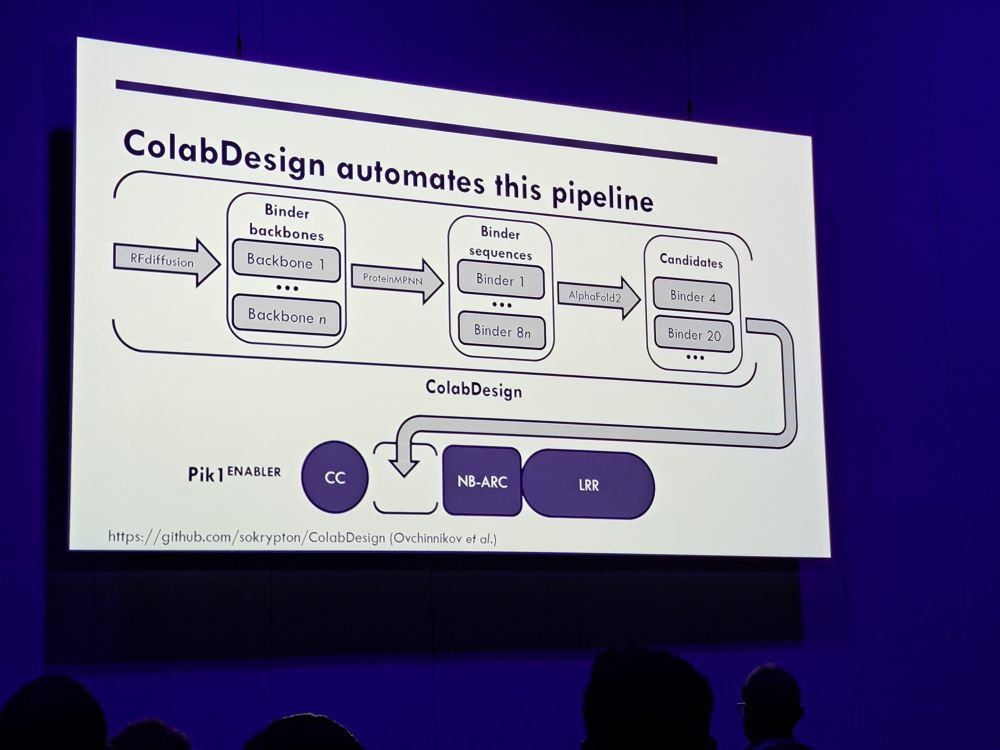

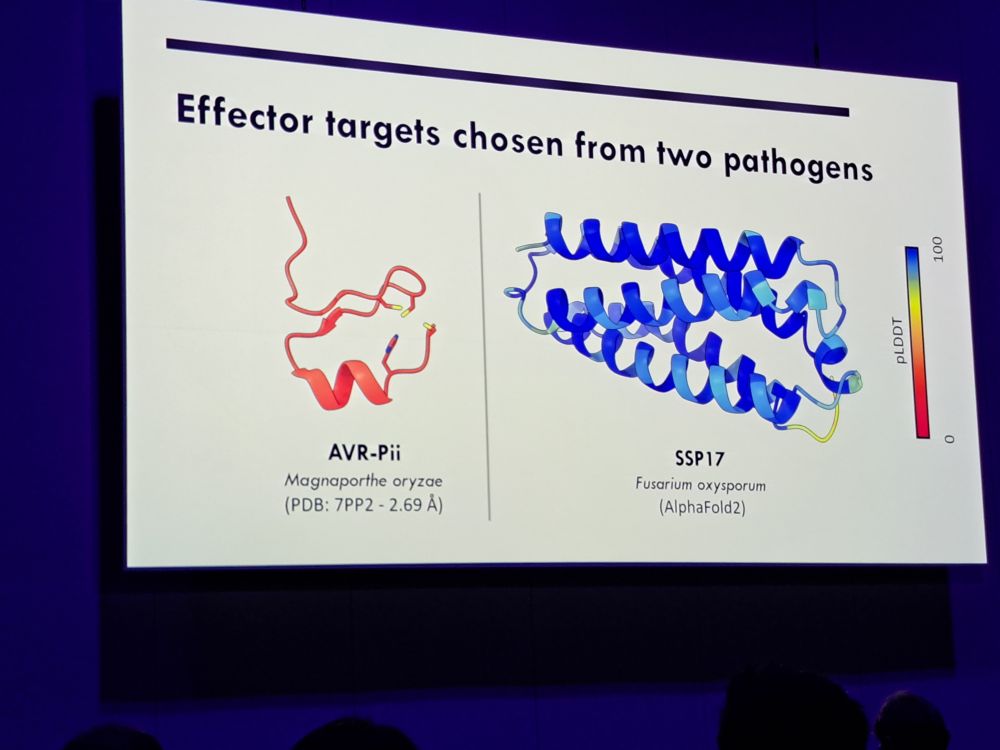
#2025ISMPMI Agnus Bucknell of the Talbot lab gives an exciting presentation to engineer fungal effectors recognizing immune sensors based on Pik1. Great and clear talk.
17.07.2025 08:58 — 👍 14 🔁 5 💬 0 📌 2
Super excited to be giving a talk at #2025ISMPMI today in the Synthetic biology concurrent session from 15:15 - 15:35! Come along if you’re interested in engineering Tandem Kinase Proteins
17.07.2025 05:51 — 👍 10 🔁 3 💬 0 📌 0
Big thanks to everyone who stopped by my poster today at #2025ISMPMI!
For those that missed it, you can check it out on Zenodo!
zenodo.org/records/1588...
Excited to attend my 1st ISMPMI conference, and present our work on a paired wheat CNL/MLKL receptor mechanism! 🌾
Come chat to me at my poster on Tuesday, P-050! 📜
If you miss my poster session, I will be talking in: Crop resistance genetics and genomics on Wednesday - 16:25-16:35!

NoCass 4th to 5th September 2025 - registration now open!
#NoCass2025 registration now open!
If you're a student or post-doc at Norwich or Cambridge, join us at the Norwich-Cambridge Science Symposium on 4-5 September - a great chance to share research & form collaborations.
Find out more & register: lnkd.in/eH3We78t
@nocassofficial.bsky.social
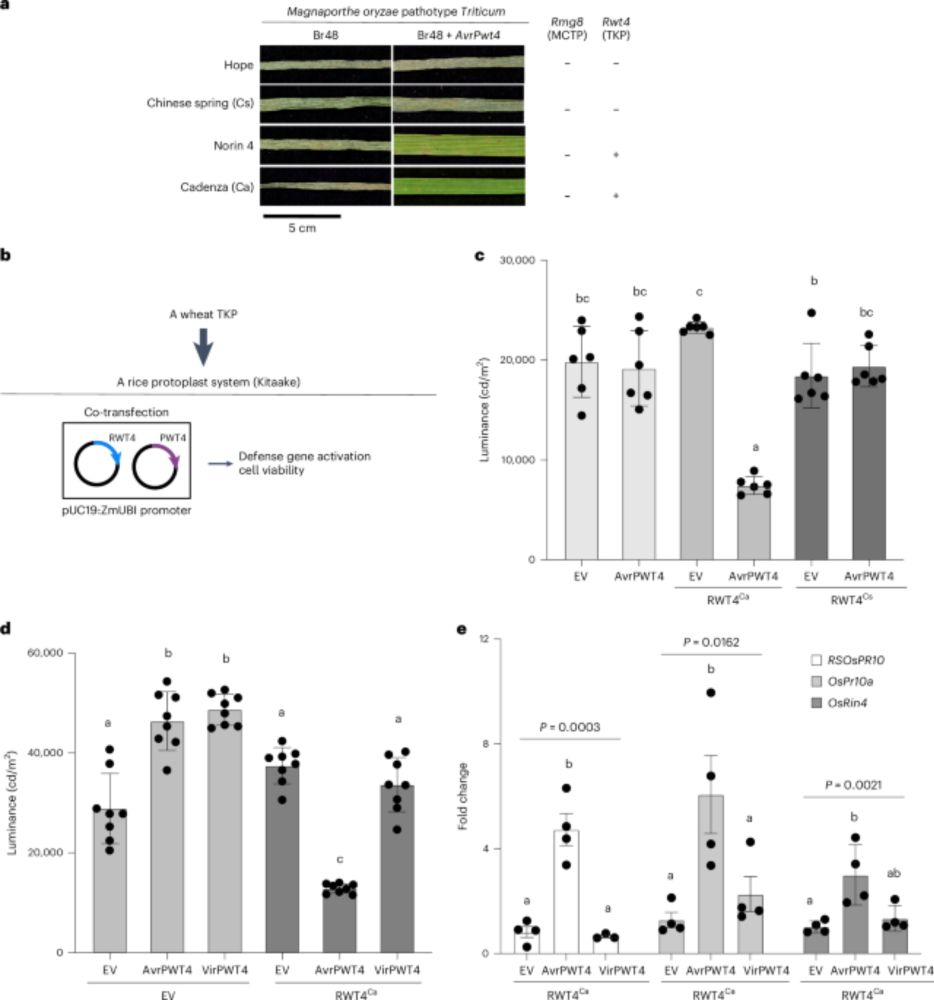
I'm very excited our most recent work on plant tandem kinases (TKPs) has finally been published! TKPs are a fascinating protein family conferring disease resistance to fungi. Originally in bioRxiv. Many thanks to @yichangsung.bsky.social for leading this work. 🧵
www.nature.com/articles/s41...
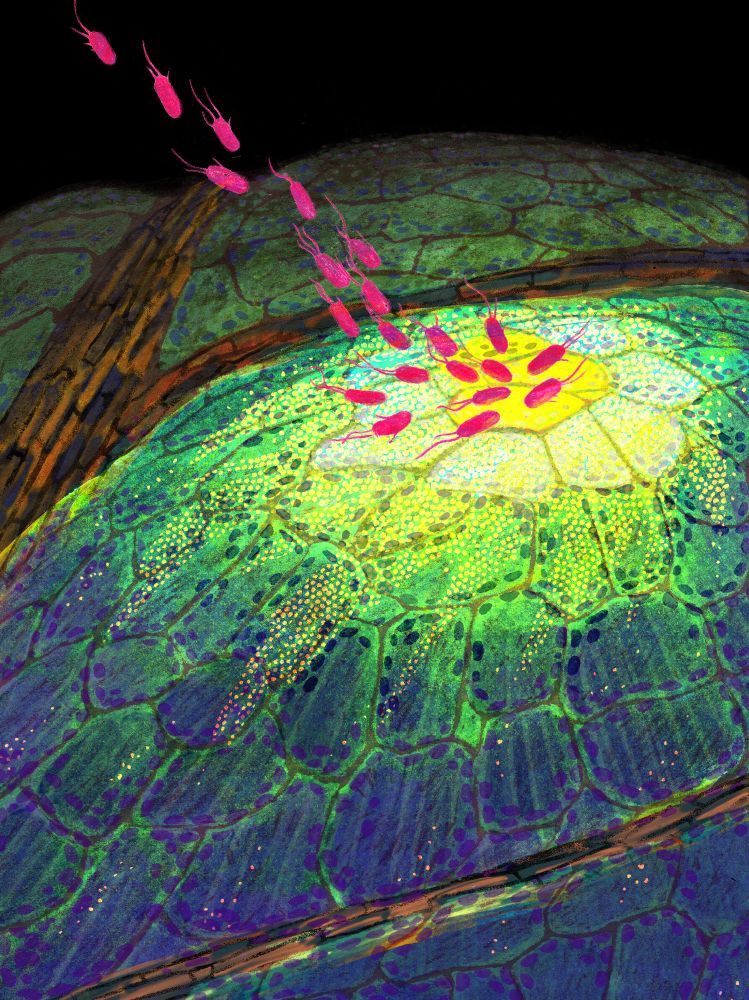
New year, new paper! Now published in @nature.com. We identified and characterised diverse immune cell states in plants under pathogen attack. My postdoc work in the Ecker lab at @salkinstitute.bsky.social. A thread (0/n)
#PlantScience
www.nature.com/articles/s41...