
🔍 About the Phagefall game
In Phagefall, you control a bacteriophage inside a petri dish. The objective is to infect and lyse bacteria before they overrun the environment. (2/n)

🔍 About the Phagefall game
In Phagefall, you control a bacteriophage inside a petri dish. The objective is to infect and lyse bacteria before they overrun the environment. (2/n)
Phold's manuscript is now available @narjournal.bsky.social thanks to @susiegriggo.bsky.social @npbhavya.bsky.social @vijinim.bsky.social @linsalrob.bsky.social @martinsteinegger.bsky.social @milot.bsky.social @eunbelivable.bsky.social & others not on bsky #phagesky academic.oup.com/nar/article/...
14.01.2026 05:10 — 👍 83 🔁 44 💬 1 📌 1
⭐ The code is fully open-source here: github.com/metagentools...
(stars help others discover it). All feedback welcome! (4/n)
🐍 Built using Pyodide + igraph + matplotlib — so Python + plotting logic all inside browser.
💡 Great for researchers working on metagenomic binning with assembly graphs — gives a quick, visual way to inspect improvements. (3/n)
🔗 Try it live: metagentools.github.io/graphbin-vis...
🧬 Upload your binning results + assembly graph files and instantly see before/after plots — your data never leaves your computer. (2/n)
#bioinformatics #metagenomics #binning #webapp #pyiodide #webassembly

🚀 Just launched: GraphBin Visualise (WASM) a browser-based visualisation tool for comparing initial metagenomic binning results vs refined results from GraphBin - all running in your browser, no backend required. (1/n)
#bioinformatics #metagenomics #binning #webapp #pyiodide #webassembly
This package to decompose weighted graphs into weighted paths by @alextomescu.bsky.social is going to be very useful. Can't wait to try it out in my viral metagenomic tools. 🤩🧬🖥️
#bioinformatics #graphs #graph-algorithms #flow-decomposition #integer-linear-programming
github.com/algbio/flowp...

Long read Metagenomics, #phage and #prophage in the gut by Ami Bhatt's group. Beautiful data showing changes in phages over two years
#phagesky
www.nature.com/articles/s41...
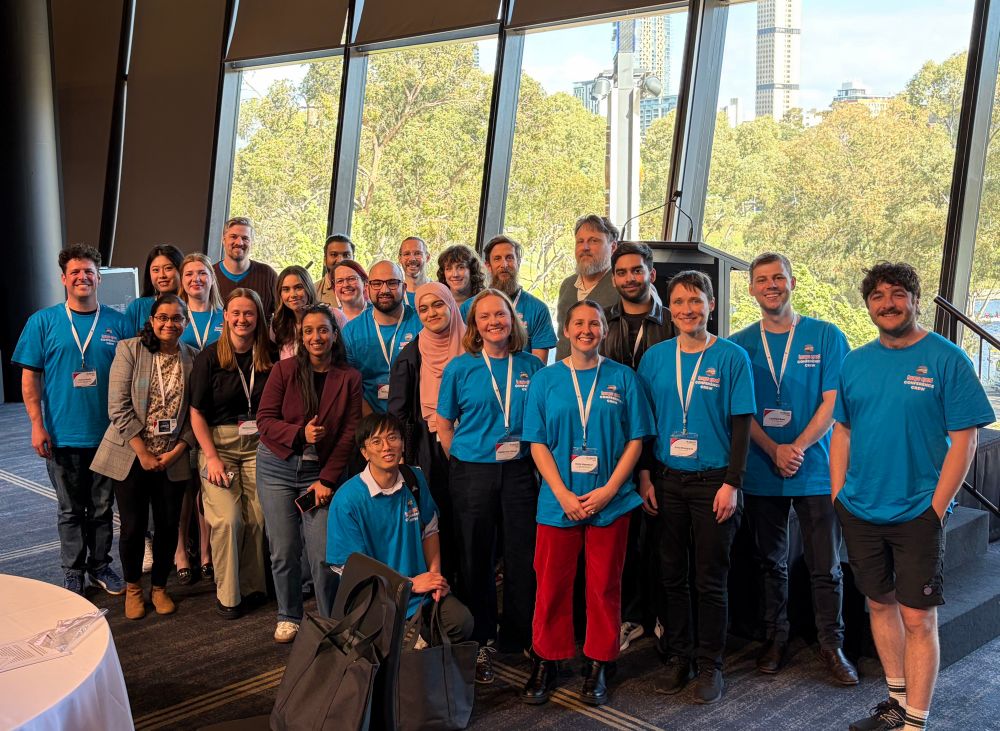

Thanks to the amazing Adelaide Bioinformatics community for making #ABACBS2025 an amazing event. We couldn’t have done it without you. A few more days of workshops to go before the festival finishes for another year 😀
26.11.2025 10:02 — 👍 30 🔁 7 💬 1 📌 3
Congratulations Hiruna Samarakoon (yet to be on bluesky) for winning the #abacbs2025 “Torsten Seemann” Outstanding Bioinformatics Software Developer Award!!! 🎉🎉🎉
26.11.2025 05:01 — 👍 19 🔁 7 💬 0 📌 0
You can check out the agtools preprint on bioRxiv for more details and analysis.
doi.org/10.1101/2025...

It was great to present TRECA at ABACBS 2025 as a lightening talk.
Come find my poster #21.
#ABACBS2025 @abacbs.bsky.social

iplotx by Fabio Zanini supports visualising any network or tree analysis library. Can’t wait to visualise assembly graphs with iplotx! 🤩💻 #ABACBS2025 @abacbs.bsky.social
26.11.2025 01:09 — 👍 7 🔁 3 💬 0 📌 0
@rrwick.bsky.social solving all of our problems in long-read bacterial genome assembly with Autocycler. 😃🧬🦠 #ABACBS2025
26.11.2025 00:52 — 👍 8 🔁 1 💬 1 📌 0Thanks for the capture! 😀
26.11.2025 00:48 — 👍 1 🔁 0 💬 0 📌 0
🧬 Come check out my poster on agtools, an open-source Python framework for analysing and manipulating assembly graphs at #ABACBS2025 Poster #106
•
💻 Github: github.com/Vini2/agtools
📄 Preprint: biorxiv.org/content/10.110…

A gymnastic start to day 2 with a keynote from Ami Bhatt #ABACBS2025
25.11.2025 22:43 — 👍 8 🔁 4 💬 0 📌 0
Thanks mate
Off the press just couple of hours before the talk
www.nature.com/articles/s41...

Great opportunity! #ABACBS2025
25.11.2025 07:26 — 👍 6 🔁 7 💬 0 📌 0
What and incredible speaker! @zaminiqbal.bsky.social I really enjoyed learning all about bacterial plasmids and their evolution. PSA they are also recruiting in Bath UK - go talk to him for the deets #ABACBS2025
25.11.2025 07:45 — 👍 8 🔁 3 💬 0 📌 1
Excited to be at ABACBS 2025. Learning, connecting and growing. 😄🧬💻 #ABACBS2025
24.11.2025 22:46 — 👍 4 🔁 0 💬 0 📌 0
MMseqs2-GPU sets new standards in single query search speed, allows near instant search of big databases, scales to multiple GPUs and is fast beyond VRAM. It enables ColabFold MSA generation in seconds and sub-second Foldseek search against AFDB50. 1/n
📄 www.nature.com/articles/s41...
💿 mmseqs.com
🌟 Exciting news! We’re launching three fully-funded postdoc positions for "New Horizons for Synthetic Phages”
Join us in tackling antimicrobial resistance with cutting-edge synthetic biology + AI bioinformatics. Based at Flinders Uni in vibrant Adelaide.
👇 Read on for details!
#Phage
A big shoutout to my amazing co-authors @rrwick.bsky.social @gbouras13.bsky.social @susiegriggo.bsky.social @npbhavya.bsky.social and @linsalrob.bsky.social. (6/n)
17.09.2025 07:03 — 👍 5 🔁 0 💬 0 📌 0
agtools is open-source and ready to drop into your own pipelines — feedback, issues, and contributions very welcome.
Github: github.com/Vini2/agtools
All you need to know about agtools can be found in the online documentation with lots of examples at agtools.readthedocs.io (5/n)
agtools exposes a Python package interface to load, query and analyse assembly graphs from popular genome and metagenome assemblers. Now you don't have to write custom code to load assembly graphs into your analyses. (4/n)
17.09.2025 07:00 — 👍 2 🔁 0 💬 1 📌 0agtools provides a command-line interface for tasks such as assembly graph format conversion, segment filtering, segment renaming, graph concatenation and component extraction. (3/n)
17.09.2025 07:00 — 👍 2 🔁 0 💬 1 📌 0Assembly graphs power assemblers and have become increasingly important in downstream applications like metagenomic binning, plasmid & viral resolution, and haplotype phasing — but tools providing programmatic access to assembly graphs are scarce. agtools gives you a CLI + Python API for this. (2/n)
17.09.2025 06:59 — 👍 2 🔁 0 💬 1 📌 0
Excited to share our latest preprint on agtools, an open-source Python framework for analysing and manipulating assembly graphs. (1/n)
www.biorxiv.org/content/10.1...
#Bioinformatics #genomics #assembly #assemblygraphs #software

I also put together an extra ColabFold formatted database of nearly 130M phage proteins that can be used to make better viral protein structure predictions github.com/gbouras13/co...
08.08.2025 07:14 — 👍 14 🔁 6 💬 1 📌 0