
If you haven't yet, check out Sylph - very useful tool (but I don't know how well it would work for your particular analyses): www.nature.com/articles/s41...
27.01.2026 18:52 — 👍 0 🔁 0 💬 1 📌 0
If you haven't yet, check out Sylph - very useful tool (but I don't know how well it would work for your particular analyses): www.nature.com/articles/s41...
27.01.2026 18:52 — 👍 0 🔁 0 💬 1 📌 0Looking for a tool to estimate abundances of aerobes versus anaerobes directly from metagenomes? Here you go: www.biorxiv.org/content/10.6...
26.01.2026 22:05 — 👍 33 🔁 13 💬 1 📌 0Please do send along any questions/comments that emerge from the discussion. You could also invite the lead author to join
14.01.2026 18:14 — 👍 2 🔁 0 💬 1 📌 0
New preprint up on airborne fungi across the US with a focus on fungal allergens - would love feedback.
www.biorxiv.org/content/10.6...
Feel free to shoot the lead author (Hoffert) an email - he can share the hi-res version. Hopefully the final version for publication will be higher resolution.....
08.01.2026 19:20 — 👍 1 🔁 0 💬 0 📌 0
New paper up - inspired by the periodic table of the elements, we attempted to organize bacterial diversity in genome-inferred trait space academic.oup.com/ismej/advanc...
06.01.2026 19:31 — 👍 54 🔁 21 💬 3 📌 1New paper out looking at mycobacterial diversity across soils, premise plumbing, and surface waters to try to figure out the source of environmentally-acquired mycobacterial infections: journals.asm.org/doi/epdf/10....
08.12.2025 19:09 — 👍 7 🔁 1 💬 0 📌 0
New paper out demonstrating how we can use strain-level analyses of soil metagenomes to investigate the biogeographical patterns exhibited by a dominant group of soil bacteria (Bradyrhizobium)
academic.oup.com/ismej/advanc...

New preprint led by Matt Gebert focused on the environmental reservoirs of nontuberculous mycobacterial (NTM) pathogens. NTM disease is environmentally acquired, but routes of exposure unclear. Our work implicates premise plumbing as the main culprit! www.biorxiv.org/content/10.1...
06.08.2025 19:59 — 👍 6 🔁 2 💬 0 📌 0After 15 years, the authors of the infamous 'arsenic life' paper are still trying to argue that their findings are legit and not just a product of 'major errors'? Unbelievable. The authors' response to the retraction is infuriating. Time to let it go. www.science.org/doi/10.1126/...
28.07.2025 18:32 — 👍 3 🔁 0 💬 2 📌 0We hope the approach and our draft 'periodic table' are useful and encourage future efforts to systematically characterize trait variance across the bacterial tree of life.(6/6)
18.07.2025 16:44 — 👍 1 🔁 1 💬 0 📌 0Perhaps more importantly, the workflow for generating the 'periodic table' is adaptable so it can ultimately be expanded to incorporate additional traits/taxa (5/6).
18.07.2025 16:44 — 👍 2 🔁 0 💬 1 📌 0Our draft 'periodic table' of bacterial diversity, like the periodic table of the elements, provides a interpretable & concise way to visualize and explore bacterial diversity in trait space (see Figs 4-5) - linking phylogeny to trait distributions in a quantifiable manner (4/6)
18.07.2025 16:44 — 👍 2 🔁 1 💬 1 📌 0Here we focused on 6 traits - inferring those traits (including autotrophy, O2 tolerance, phototrophy, etc.) using genome-based models across >50,000 bacterial 'species' (thanks @ace-gtdb.bsky.social!) (3/6)
18.07.2025 16:44 — 👍 1 🔁 0 💬 1 📌 0Work was motivated by the challenges associated with linking taxonomic/phylogenetic information to knowledge of what we know (or think we know) about bacterial traits (2/6)
18.07.2025 16:44 — 👍 2 🔁 0 💬 1 📌 0
New pre-print from my group - project led by PhD student Michael Hoffert. We set out on a daunting mission to generate a 'periodic table' of bacterial diversity (1/6) www.biorxiv.org/content/10.1...
18.07.2025 16:44 — 👍 104 🔁 43 💬 4 📌 8I agree with both of these papers. Given that they were published >10 years ago, why is the 'hologenome' concept still a thing?
14.07.2025 17:37 — 👍 4 🔁 0 💬 2 📌 0
See Table 1 of this paper: bmcbiol.biomedcentral.com/articles/10....
25.06.2025 20:15 — 👍 1 🔁 0 💬 1 📌 0Correct - if you filtered a few liters of your saliva for DNA extraction, you would undoubtedly have far far higher DNA yields than a few liters of household water
24.06.2025 13:46 — 👍 2 🔁 0 💬 2 📌 0
Hope this is useful - consensus statement "Guidelines for preventing and reporting contamination in low-biomass microbiome studies" rdcu.be/er3Io
21.06.2025 19:59 — 👍 106 🔁 48 💬 2 📌 2
New study out - we developed a pipeline for targeted assembly of genomes from (complex) soil metagenomes to examine the strain-level diversity and ecology of a dominant soil bacterial genus (Bradyrhizobium). Feedback appreciated. www.biorxiv.org/content/10.1...
30.05.2025 17:59 — 👍 12 🔁 3 💬 0 📌 0
This is cool - bacteria wrap their flagella to propel themselves through narrow passages www.biorxiv.org/content/10.1...
21.05.2025 20:36 — 👍 23 🔁 6 💬 1 📌 2Just your regular reminder that 'sterile' doesn't mean 'DNA-free'! (this rant brought to you by yet another published DNA-based study conducted in a low biomass system where appropriate controls do not appear to have been included)
08.05.2025 20:44 — 👍 68 🔁 15 💬 3 📌 1Excellent work (and very well-written)
07.05.2025 21:09 — 👍 0 🔁 0 💬 0 📌 0Dear fellow olds - recognize that the term 'next generation sequencing' is confusing to those that never had experience with the first generation. Also - what 'generation' are we on anyways?
07.05.2025 20:32 — 👍 50 🔁 8 💬 15 📌 3This looks awesome. Open to PhD students from the US?
01.05.2025 20:24 — 👍 1 🔁 0 💬 1 📌 0👆 Yes. 100% agreed.
29.04.2025 18:38 — 👍 4 🔁 0 💬 0 📌 0
So happy it is still winter here in Colorado
20.04.2025 02:38 — 👍 9 🔁 0 💬 1 📌 0
Might be a useful alternative: www.arb-silva.de/aligner/
14.04.2025 20:02 — 👍 5 🔁 0 💬 3 📌 0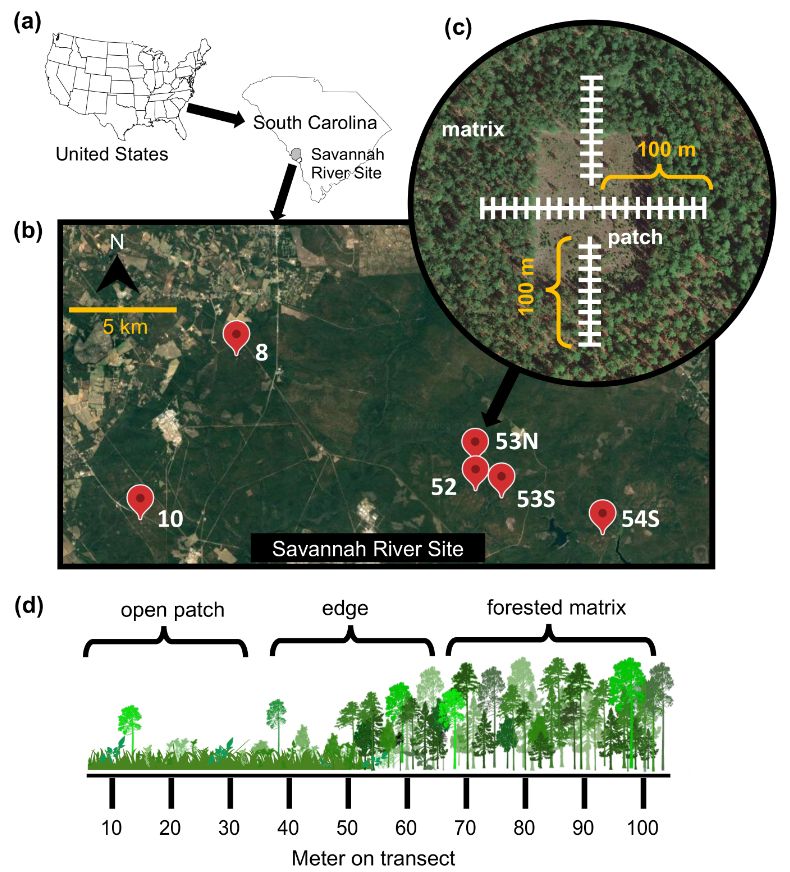
The figure shows study sites at the Savannah River Site in SC, USA, and the edge transects used in the study.
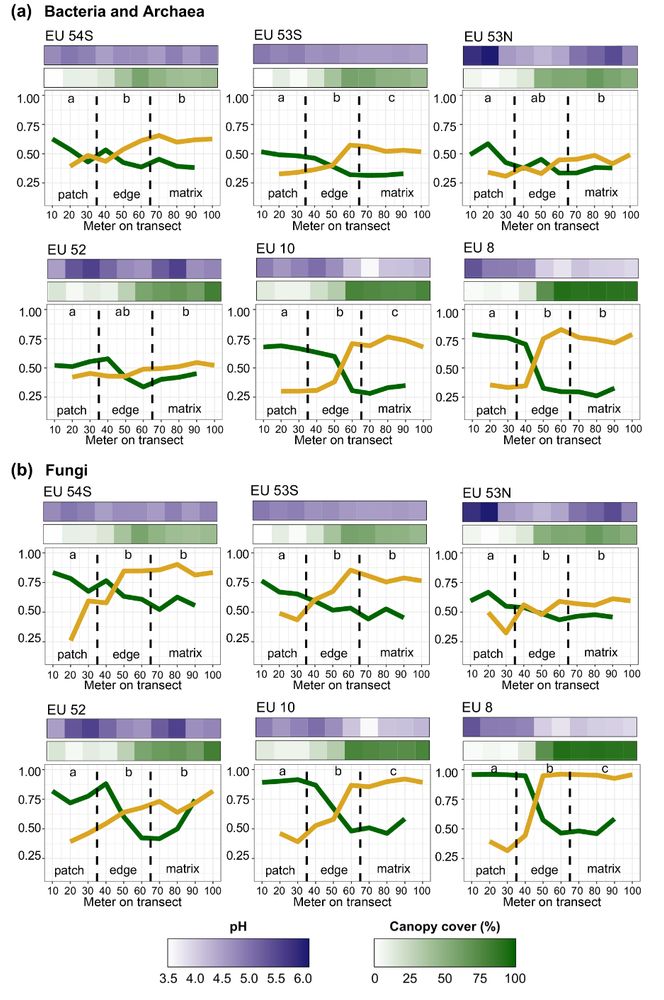
This is a multipanel figure showing bacteria/archaea and fungi turnover across edge transects in different blocks with pH and canopy cover.
Excited to share PhD student Claire Winfrey’s (w/ @noahfierer.bsky.social & me) new paper in @esajournals.bsky.social Ecology! She examines how edge effects in habitat fragments shape soil microbial communities. Check it out 🦠 🧪 🌎 esajournals.onlinelibrary.wiley.com/doi/full/10....
03.04.2025 11:41 — 👍 29 🔁 7 💬 1 📌 1